

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147434.11 - phase: 0
(150 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
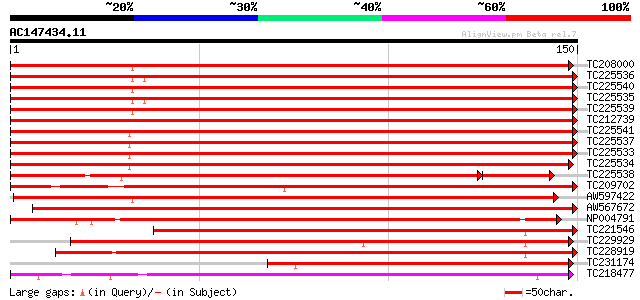
Score E
Sequences producing significant alignments: (bits) Value
TC208000 similar to UP|HS12_MEDSA (P27880) 18.2 kDa class I heat... 251 1e-67
TC225536 UP|HS16_SOYBN (P05478) 18.5 kDa class I heat shock prot... 236 2e-63
TC225540 UP|HS11_SOYBN (P02519) 17.3 kDa class I heat shock prot... 233 3e-62
TC225535 homologue to UP|HS16_SOYBN (P05478) 18.5 kDa class I he... 231 1e-61
TC225539 homologue to UP|HS11_SOYBN (P02519) 17.3 kDa class I he... 231 1e-61
TC212739 similar to UP|HS16_SOYBN (P05478) 18.5 kDa class I heat... 231 1e-61
TC225541 homologue to UP|HS13_SOYBN (P04793) 17.5 kDa class I he... 230 2e-61
TC225537 UP|HS15_SOYBN (P04795) 17.6 kDa class I heat shock prot... 229 3e-61
TC225533 homologue to UP|HS14_SOYBN (P04794) 17.5 kDa class I he... 227 1e-60
TC225534 homologue to UP|HS13_SOYBN (P04793) 17.5 kDa class I he... 224 1e-59
TC225538 homologue to UP|HS16_SOYBN (P05478) 18.5 kDa class I he... 174 1e-46
TC209702 similar to UP|Q9SYV0 (Q9SYV0) 17.6 kD class I small hea... 176 3e-45
AW597422 similar to SP|P02519|HS11 17.3 kDa class I heat shock p... 164 1e-41
AW567672 weakly similar to SP|P05478|HS16 18.5 kDa class I heat ... 154 1e-38
NP004791 seed maturation protein PM31 147 1e-36
TC221546 homologue to UP|HS41_SOYBN (P30236) 22.0 kDa class IV h... 138 1e-33
TC229929 UP|Q39819 (Q39819) Hsp22.3, complete 134 1e-32
TC228919 UP|Q39820 (Q39820) Hsp22.5, complete 129 5e-31
TC231174 similar to UP|Q41028 (Q41028) Pisum sativum 17.9 kDa he... 122 6e-29
TC218477 weakly similar to UP|HS11_MEDSA (P27879) 18.1 kDa class... 117 2e-27
>TC208000 similar to UP|HS12_MEDSA (P27880) 18.2 kDa class I heat shock
protein, partial (92%)
Length = 666
Score = 251 bits (640), Expect = 1e-67
Identities = 120/156 (76%), Positives = 138/156 (87%), Gaps = 7/156 (4%)
Frame = +3
Query: 1 MSLIPSFFGGRQNNVFDPFSMDIWDPLQGFP-------SSARETTALANTRVDWKETQEA 53
MS+IP+ FGGR++NVFDPFS+D+WDP +GFP SS E++A+ANTRVDWKET A
Sbjct: 39 MSIIPNLFGGRRSNVFDPFSLDVWDPFEGFPFSTGHVPSSGGESSAIANTRVDWKETPAA 218
Query: 54 HVFSVDLPGLKKEEVKVEIEDGNVLQISGERNKEQEEKDDKWHRVERSSGKFMRRFRLPE 113
HVF+VDLPGLKKEEVKVE+EDG VLQISGER KEQE+KDD+WHRVERS+GKFMRRFRLPE
Sbjct: 219 HVFNVDLPGLKKEEVKVEVEDGRVLQISGERTKEQEQKDDRWHRVERSTGKFMRRFRLPE 398
Query: 114 NVKMDQVKAGMENGVLTVTVPKEEEKKSEVKSIEIS 149
N KMDQVKA MENGVLTVTVPKEE+KK +VKSI+IS
Sbjct: 399 NAKMDQVKAAMENGVLTVTVPKEEDKKPQVKSIQIS 506
>TC225536 UP|HS16_SOYBN (P05478) 18.5 kDa class I heat shock protein (HSP
18.5), complete
Length = 874
Score = 236 bits (603), Expect = 2e-63
Identities = 118/161 (73%), Positives = 134/161 (82%), Gaps = 11/161 (6%)
Frame = +3
Query: 1 MSLIPSFFGGRQNNVFDPFSMDIWDPLQGFP-----SSA------RETTALANTRVDWKE 49
MSLIP+FFGGR+NNVFDPFS+D+WDP + FP SSA RE +A +TRVDWKE
Sbjct: 75 MSLIPNFFGGRRNNVFDPFSLDVWDPFKDFPFPNTLSSASFPEFSRENSAFVSTRVDWKE 254
Query: 50 TQEAHVFSVDLPGLKKEEVKVEIEDGNVLQISGERNKEQEEKDDKWHRVERSSGKFMRRF 109
T EAHVF D+PGLKKEEVKV+IED VLQISGERN E+E+K+D WHRVERSSGKFMRRF
Sbjct: 255 TPEAHVFKADIPGLKKEEVKVQIEDDKVLQISGERNVEKEDKNDTWHRVERSSGKFMRRF 434
Query: 110 RLPENVKMDQVKAGMENGVLTVTVPKEEEKKSEVKSIEISG 150
RLPEN K++QVKA MENGVLTVTVPKEE KK +VK+IEISG
Sbjct: 435 RLPENAKVEQVKASMENGVLTVTVPKEEVKKPDVKAIEISG 557
>TC225540 UP|HS11_SOYBN (P02519) 17.3 kDa class I heat shock protein (HSP
17.3), complete
Length = 792
Score = 233 bits (594), Expect = 3e-62
Identities = 112/153 (73%), Positives = 132/153 (86%), Gaps = 3/153 (1%)
Frame = +3
Query: 1 MSLIPSFFGGRQNNVFDPFSMDIWDPLQGFP---SSARETTALANTRVDWKETQEAHVFS 57
MSLIPSFFGGR+++VFDPFS+D+WDP + FP S + E +A +TRVDWKET EAHVF
Sbjct: 135 MSLIPSFFGGRRSSVFDPFSLDVWDPFKDFPFPSSLSAENSAFVSTRVDWKETPEAHVFK 314
Query: 58 VDLPGLKKEEVKVEIEDGNVLQISGERNKEQEEKDDKWHRVERSSGKFMRRFRLPENVKM 117
D+PGLKKEEVK+EI+DG VLQISGERN E+E+K+D WHRVERSSGK +RRFRLPEN K+
Sbjct: 315 ADIPGLKKEEVKLEIQDGRVLQISGERNVEKEDKNDTWHRVERSSGKLVRRFRLPENAKV 494
Query: 118 DQVKAGMENGVLTVTVPKEEEKKSEVKSIEISG 150
DQVKA MENGVLTVTVPKEE KK +VK+I+ISG
Sbjct: 495 DQVKASMENGVLTVTVPKEEIKKPDVKAIDISG 593
>TC225535 homologue to UP|HS16_SOYBN (P05478) 18.5 kDa class I heat shock
protein (HSP 18.5), complete
Length = 791
Score = 231 bits (589), Expect = 1e-61
Identities = 114/161 (70%), Positives = 135/161 (83%), Gaps = 11/161 (6%)
Frame = +3
Query: 1 MSLIPSFFGGRQNNVFDPFSMDIWDPLQGFP-----SSA------RETTALANTRVDWKE 49
MSLIP+FFGGR++NVFDPFS+D+WDP + FP SSA RE +A +TRVDWKE
Sbjct: 84 MSLIPNFFGGRRSNVFDPFSLDVWDPFKDFPFPNTLSSASFPEFSRENSAFVSTRVDWKE 263
Query: 50 TQEAHVFSVDLPGLKKEEVKVEIEDGNVLQISGERNKEQEEKDDKWHRVERSSGKFMRRF 109
T EAHVF D+PGLKKEEVKV+IED VLQISGERN E+E++++ WHRVERSSGKFMRRF
Sbjct: 264 TPEAHVFKADIPGLKKEEVKVQIEDDKVLQISGERNVEKEDRNNTWHRVERSSGKFMRRF 443
Query: 110 RLPENVKMDQVKAGMENGVLTVTVPKEEEKKSEVKSIEISG 150
RLPEN K+D+VKA MENGVLTVTVPKEE KK++VK+I+ISG
Sbjct: 444 RLPENAKVDKVKASMENGVLTVTVPKEEVKKADVKNIQISG 566
>TC225539 homologue to UP|HS11_SOYBN (P02519) 17.3 kDa class I heat shock
protein (HSP 17.3), complete
Length = 714
Score = 231 bits (589), Expect = 1e-61
Identities = 111/153 (72%), Positives = 130/153 (84%), Gaps = 3/153 (1%)
Frame = +1
Query: 1 MSLIPSFFGGRQNNVFDPFSMDIWDPLQGFP---SSARETTALANTRVDWKETQEAHVFS 57
MSLIPSFFGGR++NV DPFS+D+WDP + FP S + E +A +TRVDWKET EAHVF
Sbjct: 58 MSLIPSFFGGRRSNVLDPFSLDVWDPFKDFPFPTSLSAENSAFVSTRVDWKETPEAHVFK 237
Query: 58 VDLPGLKKEEVKVEIEDGNVLQISGERNKEQEEKDDKWHRVERSSGKFMRRFRLPENVKM 117
D+PGLKKEEVK+EI+D +LQISGERN E+E+K+D WHRVERSSGKFMR FRLP+N K+
Sbjct: 238 ADIPGLKKEEVKLEIQDDRILQISGERNVEKEDKNDTWHRVERSSGKFMRSFRLPDNAKV 417
Query: 118 DQVKAGMENGVLTVTVPKEEEKKSEVKSIEISG 150
DQVKA MENGVLTVTVPKEE KK +VK+IEISG
Sbjct: 418 DQVKASMENGVLTVTVPKEEIKKPDVKAIEISG 516
>TC212739 similar to UP|HS16_SOYBN (P05478) 18.5 kDa class I heat shock
protein (HSP 18.5), partial (93%)
Length = 543
Score = 231 bits (588), Expect = 1e-61
Identities = 108/150 (72%), Positives = 129/150 (86%)
Frame = +1
Query: 1 MSLIPSFFGGRQNNVFDPFSMDIWDPLQGFPSSARETTALANTRVDWKETQEAHVFSVDL 60
MSLIPSFFGGR++NVFDPFS+D+WDP + + E +A TRVDWKET EAHVF D+
Sbjct: 94 MSLIPSFFGGRRSNVFDPFSLDVWDPFKDLSFPSAEDSAFLKTRVDWKETPEAHVFKADI 273
Query: 61 PGLKKEEVKVEIEDGNVLQISGERNKEQEEKDDKWHRVERSSGKFMRRFRLPENVKMDQV 120
PGLKKE+VKVEIED VLQISGER+ E+E+K+DKWHRVERSSGKF+R+FRLPEN K+DQV
Sbjct: 274 PGLKKEQVKVEIEDDKVLQISGERSVEKEDKNDKWHRVERSSGKFLRKFRLPENAKVDQV 453
Query: 121 KAGMENGVLTVTVPKEEEKKSEVKSIEISG 150
KA +ENGVLTVTVPKEE KK +VK+++ISG
Sbjct: 454 KASIENGVLTVTVPKEEVKKPDVKAVQISG 543
>TC225541 homologue to UP|HS13_SOYBN (P04793) 17.5 kDa class I heat shock
protein (HSP 17.5-M), complete
Length = 849
Score = 230 bits (586), Expect = 2e-61
Identities = 112/153 (73%), Positives = 129/153 (84%), Gaps = 3/153 (1%)
Frame = +1
Query: 1 MSLIPSFFGGRQNNVFDPFSMDIWDPLQGF---PSSARETTALANTRVDWKETQEAHVFS 57
MSLIPS FGGR++NVFDPFS+D+WDP + F S + E +A NTRVDWKET EAHVF
Sbjct: 1 MSLIPSIFGGRRSNVFDPFSLDVWDPFKDFHFPTSLSAENSASVNTRVDWKETPEAHVFK 180
Query: 58 VDLPGLKKEEVKVEIEDGNVLQISGERNKEQEEKDDKWHRVERSSGKFMRRFRLPENVKM 117
D+PGLKKEEVKV+IED VLQISGERN E+E+K+D WHR+ERSSG FMRRFRLPEN K+
Sbjct: 181 ADIPGLKKEEVKVQIEDDRVLQISGERNLEKEDKNDTWHRLERSSGNFMRRFRLPENAKV 360
Query: 118 DQVKAGMENGVLTVTVPKEEEKKSEVKSIEISG 150
+QVKA MENGVLTVTVPKEE KK +VK+IEISG
Sbjct: 361 EQVKASMENGVLTVTVPKEEVKKPDVKAIEISG 459
>TC225537 UP|HS15_SOYBN (P04795) 17.6 kDa class I heat shock protein (HSP
17.6-L), complete
Length = 617
Score = 229 bits (585), Expect = 3e-61
Identities = 113/154 (73%), Positives = 129/154 (83%), Gaps = 4/154 (2%)
Frame = +2
Query: 1 MSLIPSFFGGRQNNVFDPFSMDIWDPLQGF----PSSARETTALANTRVDWKETQEAHVF 56
MSLIPS FGG ++NVFDPFS+D+WDP + F S + E +A NTRVDWKETQEAHV
Sbjct: 83 MSLIPSIFGGPRSNVFDPFSLDMWDPFKDFHVPTSSVSAENSAFVNTRVDWKETQEAHVL 262
Query: 57 SVDLPGLKKEEVKVEIEDGNVLQISGERNKEQEEKDDKWHRVERSSGKFMRRFRLPENVK 116
D+PGLKKEEVKV+IED VLQISGERN E+E+K+D WHRVERSSGKFMRRFRLPEN K
Sbjct: 263 KADIPGLKKEEVKVQIEDDRVLQISGERNVEKEDKNDTWHRVERSSGKFMRRFRLPENAK 442
Query: 117 MDQVKAGMENGVLTVTVPKEEEKKSEVKSIEISG 150
++QVKA MENGVLTVT+PKEE KKS+VK IEISG
Sbjct: 443 VEQVKACMENGVLTVTIPKEEVKKSDVKPIEISG 544
>TC225533 homologue to UP|HS14_SOYBN (P04794) 17.5 kDa class I heat shock
protein (HSP 17.5-E), complete
Length = 757
Score = 227 bits (579), Expect = 1e-60
Identities = 111/154 (72%), Positives = 129/154 (83%), Gaps = 4/154 (2%)
Frame = +3
Query: 1 MSLIPSFFGGRQNNVFDPFSMDIWDPLQGF----PSSARETTALANTRVDWKETQEAHVF 56
MSLIP FFG R++NVFDPFS+DIWDP + F S + E +A +TRVDWKET EAHVF
Sbjct: 78 MSLIPGFFGARRSNVFDPFSLDIWDPFKDFHVPTSSVSAENSAFVSTRVDWKETPEAHVF 257
Query: 57 SVDLPGLKKEEVKVEIEDGNVLQISGERNKEQEEKDDKWHRVERSSGKFMRRFRLPENVK 116
D+PGLKKEEVKV+IED VLQISGERN E+E+K+D WHRVERSSGKF+RRFRLPEN K
Sbjct: 258 KADIPGLKKEEVKVQIEDDRVLQISGERNVEKEDKNDTWHRVERSSGKFVRRFRLPENAK 437
Query: 117 MDQVKAGMENGVLTVTVPKEEEKKSEVKSIEISG 150
+++VKA MENGVLTVTVPKEE KK +VK+IEISG
Sbjct: 438 VNEVKASMENGVLTVTVPKEEVKKPDVKAIEISG 539
>TC225534 homologue to UP|HS13_SOYBN (P04793) 17.5 kDa class I heat shock
protein (HSP 17.5-M), complete
Length = 594
Score = 224 bits (571), Expect = 1e-59
Identities = 110/152 (72%), Positives = 127/152 (83%), Gaps = 3/152 (1%)
Frame = +2
Query: 1 MSLIPSFFGGRQNNVFDPFSMDIWDPLQGF---PSSARETTALANTRVDWKETQEAHVFS 57
MSLIPSFF G ++NVFDPFS+D+WDP + F S + E +A +TRVDWKET EAHV
Sbjct: 92 MSLIPSFFSGPRSNVFDPFSLDVWDPFKDFHFPTSVSAENSAFVSTRVDWKETPEAHVLK 271
Query: 58 VDLPGLKKEEVKVEIEDGNVLQISGERNKEQEEKDDKWHRVERSSGKFMRRFRLPENVKM 117
D+PGLKKEEVKV+IED VLQISGERN E+E+K+D WHRVERSSGKFMRRFRLPEN K+
Sbjct: 272 ADIPGLKKEEVKVQIEDDRVLQISGERNLEKEDKNDTWHRVERSSGKFMRRFRLPENAKV 451
Query: 118 DQVKAGMENGVLTVTVPKEEEKKSEVKSIEIS 149
+QVKA MENGVLTVTVPKEE KK +VK+IEIS
Sbjct: 452 EQVKASMENGVLTVTVPKEEIKKPDVKAIEIS 547
>TC225538 homologue to UP|HS16_SOYBN (P05478) 18.5 kDa class I heat shock
protein (HSP 18.5), partial (89%)
Length = 704
Score = 174 bits (440), Expect(2) = 1e-46
Identities = 87/128 (67%), Positives = 101/128 (77%), Gaps = 3/128 (2%)
Frame = +3
Query: 1 MSLIPSFFGGRQNNVFDPFSMDIWDPLQ---GFPSSARETTALANTRVDWKETQEAHVFS 57
MSLIP+ FGGR++NVFDPF D P FP +RE +A +TRVDWKET EAHVF
Sbjct: 60 MSLIPNIFGGRRSNVFDPFK-DFPFPNSVSTSFPEFSRENSAFVSTRVDWKETPEAHVFK 236
Query: 58 VDLPGLKKEEVKVEIEDGNVLQISGERNKEQEEKDDKWHRVERSSGKFMRRFRLPENVKM 117
D+PGLKKEEVKV+IED VLQISGERN E E+K+D WHRVERSSGKFMRRFRLPEN K+
Sbjct: 237 ADIPGLKKEEVKVQIEDDKVLQISGERNVENEDKNDTWHRVERSSGKFMRRFRLPENAKV 416
Query: 118 DQVKAGME 125
++VKA M+
Sbjct: 417 NEVKASMD 440
Score = 28.5 bits (62), Expect(2) = 1e-46
Identities = 12/19 (63%), Positives = 15/19 (78%)
Frame = +1
Query: 126 NGVLTVTVPKEEEKKSEVK 144
NG+LTVTVPK+E K + K
Sbjct: 442 NGLLTVTVPKKEVKNHDAK 498
>TC209702 similar to UP|Q9SYV0 (Q9SYV0) 17.6 kD class I small heat shock
protein (Type I small heat shock protein 17.6 kDa
isoform), partial (72%)
Length = 701
Score = 176 bits (447), Expect = 3e-45
Identities = 93/151 (61%), Positives = 110/151 (72%), Gaps = 1/151 (0%)
Frame = +2
Query: 1 MSLIPSFFGGRQNNVFDPFSMDIWDPLQGFPSSARETTALANTRVDWKETQEAHVFSVDL 60
MSLIPS R N+ DPFS +IW P S E +A N RVDWKET E+HVF DL
Sbjct: 68 MSLIPSLLSNR--NIMDPFSTNIWAP----SDSDSEVSAFVNARVDWKETPESHVFKADL 229
Query: 61 PGLKKEEVKVE-IEDGNVLQISGERNKEQEEKDDKWHRVERSSGKFMRRFRLPENVKMDQ 119
PGLKKEEVK ++G VL ISGER+ E+E+K++KWHRVER GKF R+F LPE+ K+D+
Sbjct: 230 PGLKKEEVKGGGWKEGRVLNISGERSVEKEDKNEKWHRVERGRGKFQRKFWLPEDAKVDE 409
Query: 120 VKAGMENGVLTVTVPKEEEKKSEVKSIEISG 150
VKA MENGVLTV VPK +KK EVK+IEISG
Sbjct: 410 VKASMENGVLTVIVPKVPDKKPEVKTIEISG 502
>AW597422 similar to SP|P02519|HS11 17.3 kDa class I heat shock protein (HSP
17.3). [Soybean] {Glycine max}, partial (96%)
Length = 442
Score = 164 bits (416), Expect = 1e-41
Identities = 81/147 (55%), Positives = 111/147 (75%), Gaps = 3/147 (2%)
Frame = +2
Query: 2 SLIPSFFGGRQNNVFDPFSMDIWDPLQGFP---SSARETTALANTRVDWKETQEAHVFSV 58
S+IPSFFGG++N VFDPFS+D+WDPL+ FP S + E +A + +VD K T EAHVF
Sbjct: 2 SVIPSFFGGQRNRVFDPFSLDVWDPLKDFPFPXSLSAENSAFGSAQVD*KXTPEAHVFKA 181
Query: 59 DLPGLKKEEVKVEIEDGNVLQISGERNKEQEEKDDKWHRVERSSGKFMRRFRLPENVKMD 118
++ GLKKEEVK+EI+D +LQIS ++N E+E+K++ HRVER SG M+++R P N ++D
Sbjct: 182 NIAGLKKEEVKLEIQDSRILQISRQKNVEKEDKNNT*HRVERXSGXLMKKWR*PXNTEVD 361
Query: 119 QVKAGMENGVLTVTVPKEEEKKSEVKS 145
QVKA EN +LT+TVPKEE + VK+
Sbjct: 362 QVKACDENRILTMTVPKEEIRSRMVKA 442
>AW567672 weakly similar to SP|P05478|HS16 18.5 kDa class I heat shock
protein (HSP 18.5). [Soybean] {Glycine max}, partial
(89%)
Length = 433
Score = 154 bits (390), Expect = 1e-38
Identities = 74/144 (51%), Positives = 101/144 (69%)
Frame = +2
Query: 7 FFGGRQNNVFDPFSMDIWDPLQGFPSSARETTALANTRVDWKETQEAHVFSVDLPGLKKE 66
FFGGR+NNV DPF +D+WDP + + E +AL T +DWKET +AHVF D+PGLK E
Sbjct: 2 FFGGRRNNVSDPF*LDVWDPFKDLSFPSAEDSALLKTPMDWKETPQAHVFKADIPGLKNE 181
Query: 67 EVKVEIEDGNVLQISGERNKEQEEKDDKWHRVERSSGKFMRRFRLPENVKMDQVKAGMEN 126
+VK+EIED VL+I+ +R+ E+++K+D WH V R+ GK LPEN D+V A +EN
Sbjct: 182 QVKLEIEDDMVLEINVDRSVEKQDKNDNWHLVYRTIGKISMNCTLPENS*FDRVNASIEN 361
Query: 127 GVLTVTVPKEEEKKSEVKSIEISG 150
GVLTV V E+ K+ +V ++ ISG
Sbjct: 362 GVLTVIVRHEQVKEPDV*AVHISG 433
>NP004791 seed maturation protein PM31
Length = 462
Score = 147 bits (372), Expect = 1e-36
Identities = 75/149 (50%), Positives = 108/149 (72%), Gaps = 3/149 (2%)
Frame = +1
Query: 1 MSLIPSFFGGRQNNVF-DPFS--MDIWDPLQGFPSSARETTALANTRVDWKETQEAHVFS 57
M I ++ GG+++ + DP S D+WDP + + T++LA+ VDW+ET +AH+F
Sbjct: 1 MDWIGAYRGGQRSRDWCDPSSPFTDLWDPRR-VGDADDITSSLAHAHVDWRETDKAHIFR 177
Query: 58 VDLPGLKKEEVKVEIEDGNVLQISGERNKEQEEKDDKWHRVERSSGKFMRRFRLPENVKM 117
DLPG+KKE++KV++E+ +LQISGER KE+E+++DKWHRVER G F+RRFRLPE+
Sbjct: 178 ADLPGVKKEDLKVQVEENKILQISGERVKEKEDQNDKWHRVERQCGSFLRRFRLPEDANP 357
Query: 118 DQVKAGMENGVLTVTVPKEEEKKSEVKSI 146
+Q+ +ENGVL VTVPK EKK E K++
Sbjct: 358 NQISCTLENGVLNVTVPK-VEKKPENKNV 441
>TC221546 homologue to UP|HS41_SOYBN (P30236) 22.0 kDa class IV heat shock
protein precursor, complete
Length = 811
Score = 138 bits (347), Expect = 1e-33
Identities = 65/113 (57%), Positives = 87/113 (76%), Gaps = 1/113 (0%)
Frame = +1
Query: 39 ALANTRVDWKETQEAHVFSVDLPGLKKEEVKVEIEDGNVLQISGERNKEQEEKDDKWHRV 98
A++ RVDWKET E HV +D+PGLK+EE+KVE+E+ VL++SGER KE+E+K D WHRV
Sbjct: 187 AMSPARVDWKETPEGHVIMLDVPGLKREEIKVEVEENRVLRVSGERKKEEEKKGDHWHRV 366
Query: 99 ERSSGKFMRRFRLPENVKMDQVKAGMENGVLTVTVPK-EEEKKSEVKSIEISG 150
ERS GKF R+FRLP+NV +D VKA +ENGVLT+T+ K +K + + I+G
Sbjct: 367 ERSYGKFWRQFRLPQNVDLDSVKAKLENGVLTLTLDKLSPDKIKGPRVVSIAG 525
>TC229929 UP|Q39819 (Q39819) Hsp22.3, complete
Length = 925
Score = 134 bits (338), Expect = 1e-32
Identities = 68/139 (48%), Positives = 95/139 (67%), Gaps = 6/139 (4%)
Frame = +3
Query: 17 DPFSMDIWDPLQGFPSSARETTALANTRVDWKETQEAHVFSVDLPGLKKEEVKVEIEDGN 76
DPF + +P P+ LA R DWKET AHV +DLPG+KK++VK+E+E+
Sbjct: 168 DPFGILEQNPFNNIPNIRGGAETLALARADWKETPSAHVIVLDLPGMKKKDVKIEVEESR 347
Query: 77 VLQISGERNKEQEEKD-----DKWHRVERSSGKFMRRFRLPENVKMDQVKAGMENGVLTV 131
VL+ISGER E+EE++ +KWHR ER++GKFMR+FRLP N +++V A +ENGVL +
Sbjct: 348 VLRISGERKGEEEEEEEEVEGEKWHRAERTNGKFMRQFRLPVNADLEKVTARLENGVLRI 527
Query: 132 TVPK-EEEKKSEVKSIEIS 149
TV K E+KK + K I+I+
Sbjct: 528 TVGKFGEDKKRQPKVIDIA 584
>TC228919 UP|Q39820 (Q39820) Hsp22.5, complete
Length = 814
Score = 129 bits (324), Expect = 5e-31
Identities = 65/139 (46%), Positives = 93/139 (66%), Gaps = 1/139 (0%)
Frame = +2
Query: 13 NNVFDPFSMDIWDPLQGFPSSARETTALANTRVDWKETQEAHVFSVDLPGLKKEEVKVEI 72
N+ DPF + P G T ++ RVDWKET E HV +D+PGLK++E+K+E+
Sbjct: 257 NHFPDPFRVLEQIPF-GVDKDETFTALSSHARVDWKETPEGHVIMLDVPGLKRDEIKIEV 433
Query: 73 EDGNVLQISGERNKEQEEKDDKWHRVERSSGKFMRRFRLPENVKMDQVKAGMENGVLTVT 132
E VL++SGER +E+E++ D WHRVERS GK R+F++P+NV +D VKA MENGVLT+T
Sbjct: 434 EGNRVLRVSGERKREEEKEGDHWHRVERSYGKSWRQFKVPDNVDLDSVKAKMENGVLTLT 613
Query: 133 VPK-EEEKKSEVKSIEISG 150
+ K +K + + I+G
Sbjct: 614 MNKLSPDKVKGPRLVSIAG 670
>TC231174 similar to UP|Q41028 (Q41028) Pisum sativum 17.9 kDa heat shock
protein (hsp17.9) (Fragment), partial (52%)
Length = 427
Score = 122 bits (306), Expect = 6e-29
Identities = 59/82 (71%), Positives = 72/82 (86%), Gaps = 1/82 (1%)
Frame = +2
Query: 69 KVEIED-GNVLQISGERNKEQEEKDDKWHRVERSSGKFMRRFRLPENVKMDQVKAGMENG 127
KVEIE+ G VLQISG+R KE+E+K+D WHR+ERSSG F+RRFRLPEN K+DQVKAGMENG
Sbjct: 2 KVEIEEEGRVLQISGQRTKEKEDKNDTWHRLERSSGSFLRRFRLPENAKLDQVKAGMENG 181
Query: 128 VLTVTVPKEEEKKSEVKSIEIS 149
VLTVTVPK + KK +VK ++I+
Sbjct: 182 VLTVTVPKVDVKKPDVKPVQIT 247
>TC218477 weakly similar to UP|HS11_MEDSA (P27879) 18.1 kDa class I heat
shock protein (Fragment), partial (64%)
Length = 887
Score = 117 bits (293), Expect = 2e-27
Identities = 71/172 (41%), Positives = 98/172 (56%), Gaps = 23/172 (13%)
Frame = +1
Query: 1 MSLIPS--FFGGRQNNVFDPFSMDIWD--------------------PLQGFPSSARETT 38
MSL+ S FFG R+N+ P WD P FPS + +
Sbjct: 73 MSLLSSGGFFGRRRND--PPPHQPTWDHYQAQDHHHPLGVSQPHHPPPFMSFPSDS--SP 240
Query: 39 ALANTRVDWKETQEAHVFSVDLPGLKKEEVKVEIEDGNVLQISGERNKEQEEKDDKWHRV 98
L ++WKET EAHV++ LPG K+ +V+VE++D VL I ++ E+EE+ WHRV
Sbjct: 241 VLNTALIEWKETPEAHVYNAHLPGYKRNDVRVEVDDDRVLCIVCGKSVEKEEQRGGWHRV 420
Query: 99 ERSSGKFMRRFRLPENVKMDQVKAGMENGVLTVTVPKEEE-KKSEVKSIEIS 149
E SSG+F++R LPEN +D VKA M+NGVLT+TVPK + V++I IS
Sbjct: 421 ELSSGQFVQRLTLPENSMVDHVKAYMDNGVLTITVPKHHRGVNNRVRNINIS 576
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.312 0.131 0.372
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,204,278
Number of Sequences: 63676
Number of extensions: 54711
Number of successful extensions: 369
Number of sequences better than 10.0: 99
Number of HSP's better than 10.0 without gapping: 333
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 337
length of query: 150
length of database: 12,639,632
effective HSP length: 89
effective length of query: 61
effective length of database: 6,972,468
effective search space: 425320548
effective search space used: 425320548
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 54 (25.4 bits)
Medicago: description of AC147434.11


