

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147430.8 + phase: 1 /pseudo
(437 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
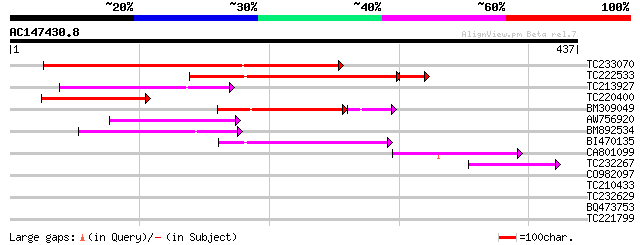
Score E
Sequences producing significant alignments: (bits) Value
TC233070 weakly similar to UP|Q9LRP0 (Q9LRP0) Emb|CAB39405.1, pa... 208 4e-54
TC222533 similar to UP|Q9SKV9 (Q9SKV9) F5J5.11, partial (11%) 159 8e-42
TC213927 98 6e-21
TC220400 92 6e-19
BM309049 77 2e-17
AW756920 similar to GP|7715599|gb|A F20B17.17 {Arabidopsis thali... 78 7e-15
BM892534 weakly similar to GP|24414144|dbj P0005C02.7 {Oryza sat... 70 1e-12
BI470135 weakly similar to GP|7715599|gb| F20B17.17 {Arabidopsis... 61 1e-09
CA801099 58 1e-08
TC232267 similar to GB|AAF68117.1|7715599|AC010793 F20B17.17 {Ar... 44 1e-04
CO982097 32 0.56
TC210433 similar to UP|Q9LRP0 (Q9LRP0) Emb|CAB39405.1, partial (... 32 0.74
TC232629 27 1.00
BQ473753 28 6.2
TC221799 similar to UP|Q8LP44 (Q8LP44) Betaine/proline transport... 28 8.1
>TC233070 weakly similar to UP|Q9LRP0 (Q9LRP0) Emb|CAB39405.1, partial (28%)
Length = 765
Score = 208 bits (529), Expect = 4e-54
Identities = 101/232 (43%), Positives = 156/232 (66%), Gaps = 1/232 (0%)
Frame = +2
Query: 27 FNIMCDMISRHGNGFKPPSYYDIRVKLLNQEVKSTNDALEEHKREWKKTSCTIMTDEWTD 86
F+ M +++ ++G G P+ + + L +E+ S + L E+K W T C+IM D W D
Sbjct: 65 FHKMLEVVGQYGQGLVCPASQLMSGRFLQEEINSIKNYLVEYKASWAITGCSIMADSWID 244
Query: 87 RRRRTILNFLVNSPRGTVFLKSVDASNICKTSEKIFEMIDAVVEEVGEDNVVQVVTDNAA 146
+ RTI+NFLV+ P G F+ SVDA+N+ + + +F+++D +VEEVGE+NVVQV+T+N
Sbjct: 245 TQGRTIINFLVSCPHGVYFVSSVDATNVVEDAPNLFKLLDKIVEEVGEENVVQVITENTP 424
Query: 147 NYKAAGEMLMRKRKKLYWTPCAAHCLDLMLEDLEKKIPIHGETIPKGKKITTFIYARTSL 206
NYKAAG+ML KR+ L+WTPCA +C++ MLED KI E + KG+KIT IY + L
Sbjct: 425 NYKAAGKMLEEKRRNLFWTPCATYCINRMLEDF-TKIRCVEECMEKGQKITKLIYNQIWL 601
Query: 207 ISILHSH-TKNRDLVRPGATRFATSYLTLGCLYENKEALIRMFTSKEWKSGK 257
++++ S T+ ++L++P ATRFA+S+ TL L +++ L RMF S +W S +
Sbjct: 602 LNLMKSEFTEGQELLKPSATRFASSFATLQSLLDHRVGLRRMFLSNKWISSR 757
>TC222533 similar to UP|Q9SKV9 (Q9SKV9) F5J5.11, partial (11%)
Length = 596
Score = 159 bits (403), Expect(2) = 8e-42
Identities = 81/164 (49%), Positives = 108/164 (65%)
Frame = +3
Query: 139 QVVTDNAANYKAAGEMLMRKRKKLYWTPCAAHCLDLMLEDLEKKIPIHGETIPKGKKITT 198
QVVTDN +NY AG++L KRK +YWTPCAAHC+DLMLED+ K +P+ +TI + +
Sbjct: 3 QVVTDNGSNYILAGKLLEEKRKHIYWTPCAAHCIDLMLEDIGK-LPLIRKTIRRAINLVG 179
Query: 199 FIYARTSLISILHSHTKNRDLVRPGATRFATSYLTLGCLYENKEALIRMFTSKEWKSGKC 258
FIY +S +S+L + T R+LVR TRFATSYLTL L++ K + +MFTS EW K
Sbjct: 180 FIYVHSSTLSLLRNFTNKRELVRHAITRFATSYLTLERLHKEKANIRKMFTSDEWTLNKL 359
Query: 259 AKLRDGKALEDIILDKKFWKDIVICLKGATPLMKVLRLADLDKK 302
+K GK ++L FW +V LK PL+KVLRL D ++K
Sbjct: 360 SKEPKGKEAAKVVLMPSFWNSVVYTLKVMAPLVKVLRLVDGERK 491
Score = 29.3 bits (64), Expect(2) = 8e-42
Identities = 13/23 (56%), Positives = 17/23 (73%)
Frame = +2
Query: 301 KKTSHGIHL*SNGSSKGANSKEF 323
K+TSHG++L*SNG K N + F
Sbjct: 485 KETSHGLYL*SNGQGKRNNYQVF 553
>TC213927
Length = 662
Score = 98.2 bits (243), Expect = 6e-21
Identities = 48/135 (35%), Positives = 77/135 (56%)
Frame = +1
Query: 39 NGFKPPSYYDIRVKLLNQEVKSTNDALEEHKREWKKTSCTIMTDEWTDRRRRTILNFLVN 98
NG++P Y +R L E + + L+ K W + +I++D W+D +RR+++NF+V
Sbjct: 247 NGYQPRGYNKLRTTLFQNERRHVENLLQPIKNAWSQKGVSIVSDGWSDPQRRSLINFMVV 426
Query: 99 SPRGTVFLKSVDASNICKTSEKIFEMIDAVVEEVGEDNVVQVVTDNAANYKAAGEMLMRK 158
+ +FLK++D SN K I + + V+ EVG N VQ+VTDN A KA ++ +
Sbjct: 427 TESRPMFLKAIDCSNEIKDKNFIAKHMREVIMEVGHSN-VQIVTDNVAVCKATSLIIKAE 603
Query: 159 RKKLYWTPCAAHCLD 173
+YWTPC H L+
Sbjct: 604 FPSIYWTPCVVHTLN 648
>TC220400
Length = 665
Score = 91.7 bits (226), Expect = 6e-19
Identities = 42/84 (50%), Positives = 58/84 (69%)
Frame = -2
Query: 25 KEFNIMCDMISRHGNGFKPPSYYDIRVKLLNQEVKSTNDALEEHKREWKKTSCTIMTDEW 84
K F M I ++G PSY+DIRV LL +EV+ T + ++ H+ +W K CTIM+D W
Sbjct: 253 KSFENMVAAIGQYGPHLPIPSYHDIRVPLLKKEVEYTENLMKGHREQWVKYGCTIMSDAW 74
Query: 85 TDRRRRTILNFLVNSPRGTVFLKS 108
TDR++R I+NFL+NS GT+FLKS
Sbjct: 73 TDRKQRCIINFLINSQAGTMFLKS 2
>BM309049
Length = 430
Score = 76.6 bits (187), Expect(2) = 2e-17
Identities = 36/100 (36%), Positives = 60/100 (60%)
Frame = +3
Query: 161 KLYWTPCAAHCLDLMLEDLEKKIPIHGETIPKGKKITTFIYARTSLISILHSHTKNRDLV 220
KLYW+PCA C++LML+D+ K + E + KIT +IY + ++ +T +D++
Sbjct: 12 KLYWSPCATQCINLMLDDIMKLEEV-SEIVSLASKITKYIYNHYYSLYLMRKYTGGKDIL 188
Query: 221 RPGATRFATSYLTLGCLYENKEALIRMFTSKEWKSGKCAK 260
P T+ AT+++ L + +K+AL M TS++W S AK
Sbjct: 189 GPAPTQSATNFIALQSILAHKDALRAMVTSRDWTSSAYAK 308
Score = 30.0 bits (66), Expect(2) = 2e-17
Identities = 16/38 (42%), Positives = 22/38 (57%)
Frame = +1
Query: 261 LRDGKALEDIILDKKFWKDIVICLKGATPLMKVLRLAD 298
LR K +E I LD +FWK +K PL+ VL++ D
Sbjct: 313 LRQKKIVEQI-LDSRFWKKCADIVKLTEPLVHVLQIVD 423
>AW756920 similar to GP|7715599|gb|A F20B17.17 {Arabidopsis thaliana},
partial (10%)
Length = 304
Score = 78.2 bits (191), Expect = 7e-15
Identities = 36/101 (35%), Positives = 61/101 (59%)
Frame = +2
Query: 78 TIMTDEWTDRRRRTILNFLVNSPRGTVFLKSVDASNICKTSEKIFEMIDAVVEEVGEDNV 137
TI+ D WTD I+ FLV+SP TVF K VDAS K ++ + ++ D+V++E G DNV
Sbjct: 2 TIIADTWTDYTSEAIIYFLVSSPSRTVFHKCVDASAYFKNTKWLADLFDSVIQEFGSDNV 181
Query: 138 VQVVTDNAANYKAAGEMLMRKRKKLYWTPCAAHCLDLMLED 178
Q++ D NY A +++ ++ ++ + C + L+L ++
Sbjct: 182 GQMIMDGCVNYTAIAHHIVQMKRTIFVSSCGSQWLNLFYDE 304
>BM892534 weakly similar to GP|24414144|dbj P0005C02.7 {Oryza sativa
(japonica cultivar-group)}, partial (6%)
Length = 430
Score = 70.5 bits (171), Expect = 1e-12
Identities = 38/126 (30%), Positives = 68/126 (53%)
Frame = -2
Query: 54 LNQEVKSTNDALEEHKREWKKTSCTIMTDEWTDRRRRTILNFLVNSPRGTVFLKSVDASN 113
L E + ++ + W + TI++D +D +RR ++NF+V + +FLK+V+ S
Sbjct: 429 LTTERSHVENLMQPIRNSWNQKGVTIVSDGRSDPQRRPLINFMVVTESEPMFLKAVNCSG 250
Query: 114 ICKTSEKIFEMIDAVVEEVGEDNVVQVVTDNAANYKAAGEMLMRKRKKLYWTPCAAHCLD 173
K + I + + + +VG NVVQ+VTD A K G ++ + +YWTP H +
Sbjct: 249 DIKDKDFIAQHMKDAIMKVGSSNVVQIVTD-AVLCKTIGMIIENEFPTIYWTPYVVHTSN 73
Query: 174 LMLEDL 179
L L+++
Sbjct: 72 LALKNI 55
>BI470135 weakly similar to GP|7715599|gb| F20B17.17 {Arabidopsis thaliana},
partial (26%)
Length = 421
Score = 60.8 bits (146), Expect = 1e-09
Identities = 37/135 (27%), Positives = 69/135 (50%), Gaps = 1/135 (0%)
Frame = +1
Query: 162 LYWTPCAAHCLDLMLEDLEKKIPIHGETIPKGKKITTFIYARTSLISILHSHTKNRDLVR 221
++ +PC + CL+L+ E+ K + I + + I IY SL+ ++ +T ++L+R
Sbjct: 4 IFVSPCGSQCLNLI*EEFSK-VDWISRCILQAQTI*NLIYNNASLLDLMKKYTGGQELIR 180
Query: 222 PGATRFATSYLTLGCLYENKEALIRMFTSKEWKSGKC-AKLRDGKALEDIILDKKFWKDI 280
G T+ +++L+L + + + L MF S E+ S A + I D FW+ +
Sbjct: 181 TGITKSVSTFLSLQSMLKLRTRLKNMFHSHEYASNTSNANKPQSLSCIAIAEDGDFWRTV 360
Query: 281 VICLKGATPLMKVLR 295
C+ + P +KVLR
Sbjct: 361 EECVAISEPFLKVLR 405
>CA801099
Length = 438
Score = 57.8 bits (138), Expect = 1e-08
Identities = 36/103 (34%), Positives = 54/103 (51%), Gaps = 3/103 (2%)
Frame = +2
Query: 296 LADLDKKTSHGIHL*SNGSSKGANSKEF*WCQKE--PVWNIIDDRWDRMLHRPLHAAAYY 353
L + KK + G + +K A K F + + + V+ IID RW LH PLHAA ++
Sbjct: 2 LLTVRKKPAMGYIYEAMEKAKEAIRKSFEYNESKYKEVFEIIDSRWTCQLHHPLHAAGHF 181
Query: 354 LNPQLHY-RPGFKADIEVRKGLMDCITRMVEDPEEQAKIEVQM 395
LNP L + + D EV GL C+ ++V D E + KI ++
Sbjct: 182 LNPDLFFSNHSMEFDFEVVNGLYVCLEKLVPDQEVRQKILTEL 310
>TC232267 similar to GB|AAF68117.1|7715599|AC010793 F20B17.17 {Arabidopsis
thaliana;} , partial (35%)
Length = 851
Score = 43.9 bits (102), Expect = 1e-04
Identities = 20/71 (28%), Positives = 35/71 (49%)
Frame = +1
Query: 354 LNPQLHYRPGFKADIEVRKGLMDCITRMVEDPEEQAKIEVQMHDFKKQIGYFGTKIAKLT 413
LNP + Y P K +++ + + +++ P+ + I Q++ F K G FG +AK
Sbjct: 1 LNPSIQYNPEIKFISSIKEDFFNVLEKLLPVPDMRRDITNQIYTFTKAHGMFGCSLAKEA 180
Query: 414 LDKSTPADWWE 424
+ P WWE
Sbjct: 181 RNTVAPWLWWE 213
>CO982097
Length = 805
Score = 32.0 bits (71), Expect = 0.56
Identities = 21/65 (32%), Positives = 29/65 (44%), Gaps = 1/65 (1%)
Frame = -2
Query: 374 LMDCITRMVE-DPEEQAKIEVQMHDFKKQIGYFGTKIAKLTLDKSTPADWWESVVFEYLE 432
L+D I R DP+ Q + +M FK FG +A + P +WWES
Sbjct: 804 LLDVIERYAHGDPDLQFNLASEMRIFKNAELDFGRLVATRERNTVMPDEWWESYGCGTPN 625
Query: 433 LQRFA 437
LQ+ A
Sbjct: 624 LQKLA 610
>TC210433 similar to UP|Q9LRP0 (Q9LRP0) Emb|CAB39405.1, partial (10%)
Length = 748
Score = 31.6 bits (70), Expect = 0.74
Identities = 19/66 (28%), Positives = 29/66 (43%)
Frame = +3
Query: 372 KGLMDCITRMVEDPEEQAKIEVQMHDFKKQIGYFGTKIAKLTLDKSTPADWWESVVFEYL 431
+G M+ + D + +Q+ + FGT++A T PA WW+ L
Sbjct: 33 EGSMNAWSGWSPDNMRRISASMQIAHYNAAQDDFGTELAISTRTGLEPAAWWQQHGISCL 212
Query: 432 ELQRFA 437
ELQR A
Sbjct: 213 ELQRIA 230
>TC232629
Length = 514
Score = 27.3 bits (59), Expect(2) = 1.00
Identities = 14/31 (45%), Positives = 17/31 (54%)
Frame = +1
Query: 154 MLMRKRKKLYWTPCAAHCLDLMLEDLEKKIP 184
+L+RK K W C C DL E +EKK P
Sbjct: 10 LLVRKN*KQRWPSC---CSDLNFETMEKKSP 93
Score = 22.3 bits (46), Expect(2) = 1.00
Identities = 7/13 (53%), Positives = 10/13 (76%)
Frame = +2
Query: 211 HSHTKNRDLVRPG 223
HSH +R++ RPG
Sbjct: 113 HSHETHREITRPG 151
>BQ473753
Length = 409
Score = 28.5 bits (62), Expect = 6.2
Identities = 11/20 (55%), Positives = 17/20 (85%)
Frame = -1
Query: 194 KKITTFIYARTSLISILHSH 213
++I +IY+RTSLIS+LH +
Sbjct: 154 RRIMIYIYSRTSLISLLHHY 95
>TC221799 similar to UP|Q8LP44 (Q8LP44) Betaine/proline transporter, partial
(26%)
Length = 398
Score = 28.1 bits (61), Expect = 8.1
Identities = 16/35 (45%), Positives = 18/35 (50%), Gaps = 3/35 (8%)
Frame = +2
Query: 274 KKFWKDIVIC---LKGATPLMKVLRLADLDKKTSH 305
+K W I IC L A + LRL DLD KT H
Sbjct: 239 QKLWHWINICFFALMSAAAAIAALRLIDLDSKTYH 343
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.325 0.138 0.432
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,856,550
Number of Sequences: 63676
Number of extensions: 264019
Number of successful extensions: 1711
Number of sequences better than 10.0: 31
Number of HSP's better than 10.0 without gapping: 1698
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1706
length of query: 437
length of database: 12,639,632
effective HSP length: 100
effective length of query: 337
effective length of database: 6,272,032
effective search space: 2113674784
effective search space used: 2113674784
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 60 (27.7 bits)
Medicago: description of AC147430.8


