

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147429.11 - phase: 0
(149 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
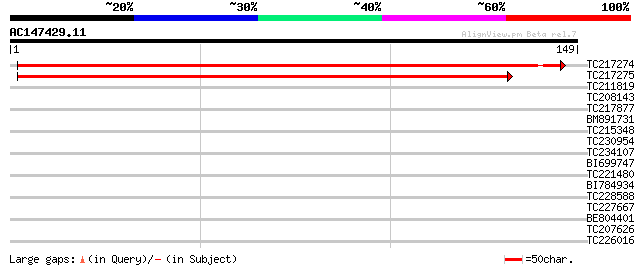
Score E
Sequences producing significant alignments: (bits) Value
TC217274 similar to UP|Q8HQU1 (Q8HQU1) Maturase, partial (3%) 232 6e-62
TC217275 193 3e-50
TC211819 similar to UP|Q8S8Z7 (Q8S8Z7) Syringolide-induced prote... 37 0.003
TC208143 UP|Q8S8Z7 (Q8S8Z7) Syringolide-induced protein B15-3-5,... 37 0.003
TC217877 similar to PRF|1908418A.0|445113|1908418A acid phosphat... 37 0.003
BM891731 32 0.11
TC215348 similar to PIR|T00795|T00795 26S proteasome regulatory ... 28 2.1
TC230954 similar to UP|SRR1_ARATH (Q8GWZ6) Protein SENSITIVITY T... 27 3.6
TC234107 similar to UP|G2O2_PEA (Q9XHM5) Gibberellin 2-beta-diox... 27 3.6
BI699747 27 4.7
TC221480 26 6.1
BI784934 26 6.1
TC228588 similar to UP|Q6NLV4 (Q6NLV4) At5g13480, partial (29%) 26 6.1
TC227667 weakly similar to UP|Q6Z3T8 (Q6Z3T8) Auxin-regulated pr... 26 6.1
BE804401 26 8.0
TC207626 26 8.0
TC226016 similar to UP|Q6ISX8 (Q6ISX8) Pathogen-inducible trypsi... 26 8.0
>TC217274 similar to UP|Q8HQU1 (Q8HQU1) Maturase, partial (3%)
Length = 1322
Score = 232 bits (591), Expect = 6e-62
Identities = 112/144 (77%), Positives = 132/144 (90%)
Frame = +3
Query: 3 SSFVLESGFYITSFSATIFIAGFAALGLLLVTLLVSMAMMLQSCQNSNGGIIELRNINND 62
SSFVLESGFYITSF+ATIF+A AA GLLL+TLLVS+AMMLQSCQ+S+ GII+L+NIN++
Sbjct: 267 SSFVLESGFYITSFAATIFVAALAATGLLLITLLVSLAMMLQSCQSSHAGIIQLQNINDE 446
Query: 63 YSYCKIHSLHAKLNNLEEHNVPNICKDLALQYIKGGQYARDLDSTKSVIEDYFNGVKPSE 122
Y+YCK++SLHAKLNNLE HN P++CKDLA++YIKGGQYARDLDSTKSVIEDYFN V+PS+
Sbjct: 447 YNYCKLYSLHAKLNNLERHNFPSLCKDLAMKYIKGGQYARDLDSTKSVIEDYFNSVRPSD 626
Query: 123 DGFDVVLIDIDSLFQWNPPHSSNL 146
DG DVVLIDID +F N PHSSNL
Sbjct: 627 DGLDVVLIDIDGIFPPN-PHSSNL 695
>TC217275
Length = 721
Score = 193 bits (490), Expect = 3e-50
Identities = 94/130 (72%), Positives = 113/130 (86%)
Frame = +2
Query: 3 SSFVLESGFYITSFSATIFIAGFAALGLLLVTLLVSMAMMLQSCQNSNGGIIELRNINND 62
SSFVLESGFYITSF ATIF+A AA GLLL+T LVS+AMMLQSCQ+S+ GIIEL+NIN+
Sbjct: 332 SSFVLESGFYITSFVATIFVAALAAAGLLLIT*LVSLAMMLQSCQSSHAGIIELQNINDQ 511
Query: 63 YSYCKIHSLHAKLNNLEEHNVPNICKDLALQYIKGGQYARDLDSTKSVIEDYFNGVKPSE 122
Y+YCK++SLH LNNLE HN P++CKDLA+++IKG Q+AR+L STKSV E YFN V+P +
Sbjct: 512 YNYCKVYSLHV*LNNLEGHNFPSLCKDLAMKFIKGSQFARELGSTKSVNEGYFNRVRPLD 691
Query: 123 DGFDVVLIDI 132
DGFDVVLIDI
Sbjct: 692 DGFDVVLIDI 721
>TC211819 similar to UP|Q8S8Z7 (Q8S8Z7) Syringolide-induced protein B15-3-5,
partial (39%)
Length = 607
Score = 37.4 bits (85), Expect = 0.003
Identities = 18/51 (35%), Positives = 26/51 (50%)
Frame = -3
Query: 83 VPNICKDLALQYIKGGQYARDLDSTKSVIEDYFNGVKPSEDGFDVVLIDID 133
VP C + YI GGQY DL+ I Y + + + DG D ++D+D
Sbjct: 206 VPEKCYNHVQNYISGGQYHNDLEVIVEHILSYASKIPLAGDGMDAWILDVD 54
>TC208143 UP|Q8S8Z7 (Q8S8Z7) Syringolide-induced protein B15-3-5, complete
Length = 962
Score = 37.4 bits (85), Expect = 0.003
Identities = 22/71 (30%), Positives = 34/71 (46%), Gaps = 1/71 (1%)
Frame = +3
Query: 64 SYCKIHSLHAKLNNLEEHN-VPNICKDLALQYIKGGQYARDLDSTKSVIEDYFNGVKPSE 122
SY + L + NN VP+ C + Y+ GGQY DL+ I Y + + +
Sbjct: 159 SYGRSWRLTVEANNARPWRIVPDNCYNHLQNYMSGGQYQLDLNLVVQHILSYAHEIPLAA 338
Query: 123 DGFDVVLIDID 133
DG D ++D+D
Sbjct: 339 DGMDAWILDVD 371
>TC217877 similar to PRF|1908418A.0|445113|1908418A acid phosphatase 1.
{Lycopersicon esculentum;} , partial (67%)
Length = 963
Score = 37.0 bits (84), Expect = 0.003
Identities = 19/60 (31%), Positives = 26/60 (42%)
Frame = +3
Query: 83 VPNICKDLALQYIKGGQYARDLDSTKSVIEDYFNGVKPSEDGFDVVLIDIDSLFQWNPPH 142
+P C + Y+ G YA DL+ E+Y V DG D + DID N P+
Sbjct: 231 IPEECAEYVKDYMSGKGYALDLEMVSKEAEEYARTVPLGYDGKDAWVFDIDETLLSNLPY 410
>BM891731
Length = 335
Score = 32.0 bits (71), Expect = 0.11
Identities = 29/98 (29%), Positives = 44/98 (44%), Gaps = 8/98 (8%)
Frame = +3
Query: 21 FIAGFAALG-----LLLVTLLVSMAMMLQSCQNSNGGIIELRNINNDYS--YCKIHSLHA 73
FI F A+ L LV + VS + + S I+ L + + + S YC L
Sbjct: 21 FIITFPAMDSGAWLLFLVVVAVSTSGHIHS-----EAILRLPSESEEISRDYCDSWMLAX 185
Query: 74 KLNNLEEHN-VPNICKDLALQYIKGGQYARDLDSTKSV 110
++ N N VP C D +YI G +Y RD D +++
Sbjct: 186 EIINAGTWNRVPASCVDFVAEYITGDRYRRDCDXIRNL 299
>TC215348 similar to PIR|T00795|T00795 26S proteasome regulatory subunit
[imported] - Arabidopsis thaliana {Arabidopsis
thaliana;} , partial (34%)
Length = 1564
Score = 27.7 bits (60), Expect = 2.1
Identities = 13/46 (28%), Positives = 26/46 (56%)
Frame = -3
Query: 33 VTLLVSMAMMLQSCQNSNGGIIELRNINNDYSYCKIHSLHAKLNNL 78
+ +LV + S + SN + ++N+D+S+C +L K+NN+
Sbjct: 290 ILMLVQNNFLQLSPKCSNA*VTSFTDLNHDHSHCYKGTLTNKINNI 153
>TC230954 similar to UP|SRR1_ARATH (Q8GWZ6) Protein SENSITIVITY TO RED LIGHT
REDUCED 1, partial (52%)
Length = 964
Score = 26.9 bits (58), Expect = 3.6
Identities = 13/37 (35%), Positives = 20/37 (53%)
Frame = +2
Query: 105 DSTKSVIEDYFNGVKPSEDGFDVVLIDIDSLFQWNPP 141
D ++ + DYFN V SE +V+ I S+ + PP
Sbjct: 317 DQIQTSVVDYFNRVLGSEIKMQMVIYGIGSIKLYEPP 427
>TC234107 similar to UP|G2O2_PEA (Q9XHM5) Gibberellin 2-beta-dioxygenase 2
(Gibberellin 2-beta-hydroxylase 2) (Gibberellin
2-oxidase 2) (GA 2-oxidase 2) , partial (40%)
Length = 687
Score = 26.9 bits (58), Expect = 3.6
Identities = 11/32 (34%), Positives = 18/32 (55%)
Frame = -2
Query: 66 CKIHSLHAKLNNLEEHNVPNICKDLALQYIKG 97
CK+H H+ L + +H V + CK Q ++G
Sbjct: 284 CKVHH*HSGLCCVRQHPVSHACKPSICQNLQG 189
>BI699747
Length = 413
Score = 26.6 bits (57), Expect = 4.7
Identities = 10/23 (43%), Positives = 17/23 (73%)
Frame = -1
Query: 51 GGIIELRNINNDYSYCKIHSLHA 73
G I+ LR ++N ++ CK H+LH+
Sbjct: 152 GNILRLRQLSN-FNICKFHNLHS 87
>TC221480
Length = 763
Score = 26.2 bits (56), Expect = 6.1
Identities = 15/45 (33%), Positives = 24/45 (53%)
Frame = +3
Query: 9 SGFYITSFSATIFIAGFAALGLLLVTLLVSMAMMLQSCQNSNGGI 53
S F++ S SAT+F A+ LL + S + +C +NGG+
Sbjct: 51 SDFFVFSASATVFHHSRDAVVQLLRSCFASTLGLGSACIYNNGGV 185
>BI784934
Length = 421
Score = 26.2 bits (56), Expect = 6.1
Identities = 18/65 (27%), Positives = 33/65 (50%), Gaps = 2/65 (3%)
Frame = +1
Query: 61 NDYSYCKIHSLHAKLNNLEEHNVPNICKDLALQYIKGGQYARDLDSTKSVIEDYF--NGV 118
N Y Y K+ HA +++EEH+ +D A +Y+ ++D + I +++ N
Sbjct: 85 NWYKYLKLQKGHAGGSDVEEHS-----RDSAAKYV----ILEEMDEQEDGI*NHYAPNSS 237
Query: 119 KPSED 123
KP+ D
Sbjct: 238 KPNND 252
>TC228588 similar to UP|Q6NLV4 (Q6NLV4) At5g13480, partial (29%)
Length = 1178
Score = 26.2 bits (56), Expect = 6.1
Identities = 13/29 (44%), Positives = 15/29 (50%)
Frame = -3
Query: 119 KPSEDGFDVVLIDIDSLFQWNPPHSSNLL 147
KP EDG V D +W P SSN+L
Sbjct: 228 KPMEDGTIVTPTDKVLFMEWMPCQSSNIL 142
>TC227667 weakly similar to UP|Q6Z3T8 (Q6Z3T8) Auxin-regulated protein-like,
partial (33%)
Length = 1445
Score = 26.2 bits (56), Expect = 6.1
Identities = 18/61 (29%), Positives = 31/61 (50%), Gaps = 3/61 (4%)
Frame = +3
Query: 21 FIAG-FAALGLLLVTLLVSMAMMLQSCQNSNGGIIELRNINNDYSYCKIHSLHAKL--NN 77
F+ G F ++ + LVT ++ +Q Q++ GGI + ++D H+ H K NN
Sbjct: 1119 FLRGHFGSVSVNLVTFCNCSSVAIQFTQDAAGGIFVVCLFSSDVKMSLYHNYHRKKIDNN 1298
Query: 78 L 78
L
Sbjct: 1299 L 1301
>BE804401
Length = 405
Score = 25.8 bits (55), Expect = 8.0
Identities = 17/60 (28%), Positives = 31/60 (51%)
Frame = +2
Query: 27 ALGLLLVTLLVSMAMMLQSCQNSNGGIIELRNINNDYSYCKIHSLHAKLNNLEEHNVPNI 86
+L LLL ++L+S+ + + S GIIE + +LH +++LE H P++
Sbjct: 17 SLSLLLFSILLSLTGLRNNELRSQPGIIERAFL--------*FTLHGHIHSLEHHQRPSL 172
>TC207626
Length = 1824
Score = 25.8 bits (55), Expect = 8.0
Identities = 8/21 (38%), Positives = 14/21 (66%)
Frame = -2
Query: 66 CKIHSLHAKLNNLEEHNVPNI 86
C++ +H + LE HN+PN+
Sbjct: 1814 CRMIGVHRRSTRLEIHNIPNL 1752
>TC226016 similar to UP|Q6ISX8 (Q6ISX8) Pathogen-inducible
trypsin-inhibitor-like protein, partial (85%)
Length = 853
Score = 25.8 bits (55), Expect = 8.0
Identities = 11/30 (36%), Positives = 17/30 (56%)
Frame = +2
Query: 65 YCKIHSLHAKLNNLEEHNVPNICKDLALQY 94
Y + L + N EEHN +IC L+L++
Sbjct: 5 YSNTN*LTKSIQNNEEHNAASICPCLSLEF 94
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.321 0.138 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,772,025
Number of Sequences: 63676
Number of extensions: 85178
Number of successful extensions: 431
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 431
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 431
length of query: 149
length of database: 12,639,632
effective HSP length: 89
effective length of query: 60
effective length of database: 6,972,468
effective search space: 418348080
effective search space used: 418348080
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 54 (25.4 bits)
Medicago: description of AC147429.11


