

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
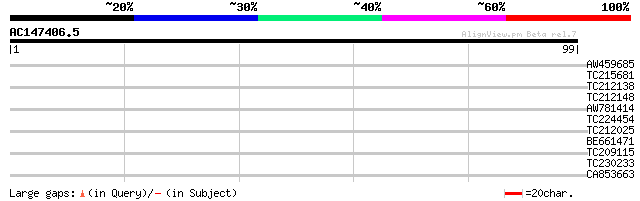
Query= AC147406.5 - phase: 0 /pseudo
(99 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AW459685 similar to GP|13506749|gb| receptor-like protein kinase... 27 1.2
TC215681 similar to UP|Q9LPR4 (Q9LPR4) F15H18.3, partial (79%) 27 1.2
TC212138 27 1.6
TC212148 27 2.1
AW781414 26 2.8
TC224454 25 4.7
TC212025 similar to UP|Q8H934 (Q8H934) Phototropin, partial (27%) 25 6.1
BE661471 weakly similar to PIR|A86318|A86 protein F15H18.11 [imp... 25 6.1
TC209115 25 6.1
TC230233 homologue to UP|Q39811 (Q39811) Alpha galactosidase , p... 25 6.1
CA853663 weakly similar to PIR|A86318|A86 protein F15H18.11 [imp... 25 8.0
>AW459685 similar to GP|13506749|gb| receptor-like protein kinase 6
{Arabidopsis thaliana}, partial (5%)
Length = 408
Score = 27.3 bits (59), Expect = 1.2
Identities = 16/44 (36%), Positives = 23/44 (51%)
Frame = +2
Query: 30 VVKLLGKNLGYHAMRERLQKTWRTQGGFEIMDNDNGFYMVKFDE 73
VV L+ Y R R QK ++++GG +DND +K DE
Sbjct: 197 VVLLVAFTYNYLGARRRRQKPFQSEGGEGELDND-----IKIDE 313
>TC215681 similar to UP|Q9LPR4 (Q9LPR4) F15H18.3, partial (79%)
Length = 2427
Score = 27.3 bits (59), Expect = 1.2
Identities = 16/50 (32%), Positives = 28/50 (56%)
Frame = +3
Query: 33 LLGKNLGYHAMRERLQKTWRTQGGFEIMDNDNGFYMVKFDEAADKEKVIT 82
+LGK G HA+R+RL++ G+E+ D+ F A+++K +T
Sbjct: 1389 VLGKLSGRHALRKRLEEL-----GYELNDDQVQTLFWCFKAVAEQKKRVT 1523
>TC212138
Length = 1006
Score = 26.9 bits (58), Expect = 1.6
Identities = 17/47 (36%), Positives = 22/47 (46%), Gaps = 3/47 (6%)
Frame = -3
Query: 7 FYPRFISNNQSFNNYVRHGRMQFVVKLLGK---NLGYHAMRERLQKT 50
F PRFI N+ FNN+ H + K K N Y A +L +T
Sbjct: 959 FCPRFILTNEKFNNFPLHLFYLLITKKQFKT*NNNAYPARNIKLTQT 819
>TC212148
Length = 700
Score = 26.6 bits (57), Expect = 2.1
Identities = 9/18 (50%), Positives = 13/18 (72%)
Frame = -2
Query: 7 FYPRFISNNQSFNNYVRH 24
F P+F++ N SFNN + H
Sbjct: 297 FSPKFLTTNTSFNNLLHH 244
>AW781414
Length = 241
Score = 26.2 bits (56), Expect = 2.8
Identities = 10/20 (50%), Positives = 14/20 (70%)
Frame = +1
Query: 77 KEKVITGGPWLIFDHCLAVT 96
++KV+ W+IFDH L VT
Sbjct: 1 RKKVMNRRSWMIFDHYLVVT 60
>TC224454
Length = 660
Score = 25.4 bits (54), Expect = 4.7
Identities = 12/34 (35%), Positives = 14/34 (40%)
Frame = +2
Query: 65 GFYMVKFDEAADKEKVITGGPWLIFDHCLAVTHW 98
GFY F D KV G W + L +T W
Sbjct: 413 GFYKFNFASLEDLHKVWIAGSWKLQLGILRLTQW 514
>TC212025 similar to UP|Q8H934 (Q8H934) Phototropin, partial (27%)
Length = 978
Score = 25.0 bits (53), Expect = 6.1
Identities = 10/24 (41%), Positives = 17/24 (70%)
Frame = +2
Query: 32 KLLGKNLGYHAMRERLQKTWRTQG 55
KLLG+ + ++ +R R+ KTW+ G
Sbjct: 362 KLLGEIVVFYRVRTRIPKTWQR*G 433
>BE661471 weakly similar to PIR|A86318|A86 protein F15H18.11 [imported] -
Arabidopsis thaliana, partial (4%)
Length = 662
Score = 25.0 bits (53), Expect = 6.1
Identities = 12/32 (37%), Positives = 18/32 (55%)
Frame = +2
Query: 35 GKNLGYHAMRERLQKTWRTQGGFEIMDNDNGF 66
G+N+ Y + QK + Q GFE+ + NGF
Sbjct: 125 GQNISYPFYIQGKQKPFCGQPGFELTCSHNGF 220
>TC209115
Length = 1284
Score = 25.0 bits (53), Expect = 6.1
Identities = 13/47 (27%), Positives = 20/47 (41%), Gaps = 9/47 (19%)
Frame = +3
Query: 51 WRTQGGFEIMDNDNGFYMVKFDE---------AADKEKVITGGPWLI 88
W+ F+ + D G Y+ + + D K+ TG PWLI
Sbjct: 384 WKESDRFQPIVVDPGLYLARKSQIFQATQKRPTPDAFKLFTGSPWLI 524
>TC230233 homologue to UP|Q39811 (Q39811) Alpha galactosidase , partial (82%)
Length = 1276
Score = 25.0 bits (53), Expect = 6.1
Identities = 15/44 (34%), Positives = 21/44 (47%)
Frame = -2
Query: 4 EIVFYPRFISNNQSFNNYVRHGRMQFVVKLLGKNLGYHAMRERL 47
EI +YP ++ F+NY H F KLL + H R +L
Sbjct: 537 EISYYPTYLECLLLFSNYFPH----FWPKLLDLLIPIHTKRRKL 418
>CA853663 weakly similar to PIR|A86318|A86 protein F15H18.11 [imported] -
Arabidopsis thaliana, partial (4%)
Length = 435
Score = 24.6 bits (52), Expect = 8.0
Identities = 12/32 (37%), Positives = 17/32 (52%)
Frame = +2
Query: 35 GKNLGYHAMRERLQKTWRTQGGFEIMDNDNGF 66
G+N+ Y + QK + Q GFE+ NGF
Sbjct: 143 GQNISYPFYIQGKQKPFCGQPGFELTCGHNGF 238
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.327 0.141 0.460
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,490,204
Number of Sequences: 63676
Number of extensions: 45680
Number of successful extensions: 291
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 291
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 291
length of query: 99
length of database: 12,639,632
effective HSP length: 75
effective length of query: 24
effective length of database: 7,863,932
effective search space: 188734368
effective search space used: 188734368
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 51 (24.3 bits)
Medicago: description of AC147406.5


