

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147404.2 + phase: 0
(599 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
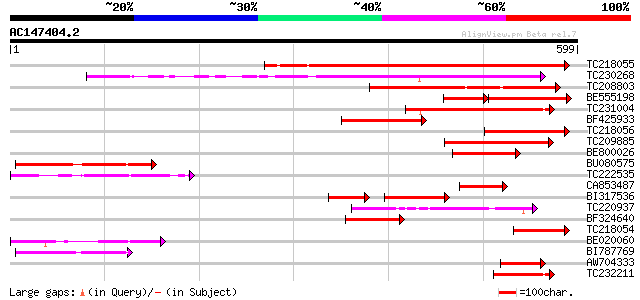
Sequences producing significant alignments: (bits) Value
TC218055 similar to UP|Q9SNF1 (Q9SNF1) Inositol-1,4,5-trisphosph... 476 e-134
TC230268 weakly similar to UP|I5P2_HUMAN (P32019) Type II inosit... 260 1e-69
TC208803 similar to UP|Q8H0Z6 (Q8H0Z6) At1g71710/F14O23_9, parti... 228 6e-60
BE555198 similar to GP|10444263|gb inositol polyphosphate 5-phos... 139 7e-54
TC231004 similar to UP|Q9LR47 (Q9LR47) T25N20.12, partial (30%) 159 4e-39
BF425933 similar to GP|10444263|gb| inositol polyphosphate 5-pho... 147 2e-35
TC218056 similar to UP|Q9FUR2 (Q9FUR2) Inositol polyphosphate 5-... 140 1e-33
TC209885 similar to GB|AAO64848.1|29028938|BT005913 At3g63240 {A... 129 3e-30
BE800026 similar to GP|10444263|gb| inositol polyphosphate 5-pho... 128 8e-30
BU080575 123 2e-28
TC222535 similar to UP|OCRL_HUMAN (Q01968) Inositol polyphosphat... 97 3e-20
CA853487 similar to GP|10444263|gb| inositol polyphosphate 5-pho... 96 5e-20
BI317536 61 8e-20
TC220937 weakly similar to GB|AAB60525.1|1166575|U45479 145 kDa ... 89 4e-18
BF324640 similar to GP|29028938|gb| At3g63240 {Arabidopsis thali... 88 1e-17
TC218054 similar to UP|Q9FUR2 (Q9FUR2) Inositol polyphosphate 5-... 88 1e-17
BE020060 86 4e-17
BI787769 weakly similar to GP|15081807|gb| At1g71710/F14O23_9 {A... 79 7e-15
AW704333 75 6e-14
TC232211 homologue to UP|Q9LR47 (Q9LR47) T25N20.12, partial (12%) 70 3e-12
>TC218055 similar to UP|Q9SNF1 (Q9SNF1) Inositol-1,4,5-trisphosphate
5-Phosphatase , partial (39%)
Length = 1457
Score = 476 bits (1225), Expect = e-134
Identities = 236/325 (72%), Positives = 275/325 (84%), Gaps = 3/325 (0%)
Frame = +1
Query: 270 KSLHQSSGNFSLLWSEKQQEIVPQVFDSHLDVSDMLSDEDNDTFSELANNEDANGIISVK 329
K H+SSGN LLW +QQ+++P+V DS DVSD+LS E DTF + N+ED + + +
Sbjct: 10 KRSHRSSGNLGLLW--QQQQVIPEVVDSLEDVSDVLSAEGGDTFI-VPNDEDEDEFGTTE 180
Query: 330 SHP--KYVRIVSKQMVGIYVSVWVQRKLRRHVHHLKVSQVGVGLMGYMGNKGSVSVSMSV 387
S P +YVRIVSKQMVGIYVSVWVQR+LRRH+++LKVS VGVGLMGYMGNKGSVS+SMS+
Sbjct: 181 SCPSTRYVRIVSKQMVGIYVSVWVQRRLRRHINNLKVSPVGVGLMGYMGNKGSVSISMSL 360
Query: 388 FQSRMCFVCSHLASGQKDGAEQRRNSDVHEILQRTRFSS-VFDTDQPQKIPSHDKIFWFG 446
FQSRMCFVCSHL SGQK+GAE RRNSDVHEIL+RT FSS VFD DQPQ IPSHD+IFWFG
Sbjct: 361 FQSRMCFVCSHLTSGQKEGAEHRRNSDVHEILRRTCFSSSVFDADQPQTIPSHDQIFWFG 540
Query: 447 DLNYRINMSDGEIRKLVDLKKWNELMKFDQLSNELCKGHVFEGWKEGLINFPPTYKYEFN 506
DLNYRINM D E+RKLV L+KW+EL +DQLS EL GHVF+GWKEGLINFPPTYKYEFN
Sbjct: 541 DLNYRINMLDAEVRKLVALRKWDELKNYDQLSKELRMGHVFDGWKEGLINFPPTYKYEFN 720
Query: 507 SDKHVGGNTQEGEKRRAPAWCDRILWLGKGIKQLKYQSAENQLSDHRPVSSIFLVDVEVI 566
SD++VG + +EGEKRR+PAWCDRILWLGKGIKQL+Y AE +LSDHRPVSS FLV+VEV
Sbjct: 721 SDRYVGESPKEGEKRRSPAWCDRILWLGKGIKQLQYGRAEIKLSDHRPVSSAFLVEVEVF 900
Query: 567 DHRKLERAIYFASAVVHPDVFLKED 591
DHRKL+RA+ F A VHP++FL ED
Sbjct: 901 DHRKLKRALNFTRAAVHPEIFLDED 975
>TC230268 weakly similar to UP|I5P2_HUMAN (P32019) Type II
inositol-1,4,5-trisphosphate 5-phosphatase precursor
(Phosphoinositide 5-phosphatase) (5PTase) (Fragment) ,
partial (8%)
Length = 1744
Score = 260 bits (665), Expect = 1e-69
Identities = 170/491 (34%), Positives = 255/491 (51%), Gaps = 6/491 (1%)
Frame = +2
Query: 82 RKSETLRAQYINTKEVRVTIGTWNVAGKHPCNDLEIEGWLCTEEEPSDIYIIGFQEVVPL 141
R SE + + RV TWNV G+ P +L++ +L EP D+Y
Sbjct: 26 RGSEPSMSSSEAIQNFRVFAATWNVGGQCPTGNLDLNDFLQVRNEP-DMY---------- 172
Query: 142 NAGNVFGAEDSKPIPKWDALIRRTLNKSSEPGTKKKSNSAPPSPIRRISSFNTQINPLDS 201
V G ++ P+ + L+ + +EP K + ++
Sbjct: 173 ----VLGFQEIVPLNAGNVLVL----EDNEPAAK-------------------WLALINQ 271
Query: 202 ALDKKEEIKTIISIEKNLQLSKIYDIDLQTILDWPELRLDPIHHVDSSPKMRRVQSTSDS 261
+L+ ++ + K L+L+ + L P +++++ T
Sbjct: 272 SLNGSSDLAS-----KGLKLTASFGGPL----------------FSQKPSLKKIKKTFKK 388
Query: 262 ASLYGFEMKSLHQSSGNFSLLWSEKQ--QEIVPQVFDSHLDVSDMLSDEDNDTFSELANN 319
L G +KS + +L E++ ++ + +S+ + D ++E+++ F+
Sbjct: 389 --LNGKRLKSCN------CVLEMERKAAKDFCFRCQESNFNSDDSSTEEEDENFTI---- 532
Query: 320 EDANGIISVKSHPKYVRIVSKQMVGIYVSVWVQRKLRRHVHHLKVSQVGVGLMGYMGNKG 379
+ S KY + KQMVGI+VSVW++R+L ++V HL++ G+MG +GNKG
Sbjct: 533 ----PVALATSQMKYSLVACKQMVGIFVSVWMRRELVQYVGHLRICCTSRGIMGCLGNKG 700
Query: 380 SVSVSMSVFQSRMCFVCSHLASGQKDGAEQRRNSDVHEILQRTRFSSVFDTD---QPQKI 436
+SVSMS +Q+ CF+CSHLASG+K+G E RRN DV EIL+ T+F + T P KI
Sbjct: 701 CISVSMSFYQTSFCFICSHLASGEKEGDELRRNLDVIEILKNTQFPRICKTPHSRMPDKI 880
Query: 437 PSHDKIFWFGDLNYRINMSDGEIRKLVDLKKWNELMKFDQLSNELCKGHVFEGWKEGLIN 496
HD+I WFGDLNYRI++S + ++LV+ + W L DQL E G VF+GWKEG I
Sbjct: 881 LDHDRIIWFGDLNYRISLSHDDAKRLVEKRDWPALFNKDQLKMEREAGRVFKGWKEGKIY 1060
Query: 497 FPPTYKYEFNSDK-HVGGNTQEGEKRRAPAWCDRILWLGKGIKQLKYQSAENQLSDHRPV 555
F PTYKY FNSD +V G KRR PAWCDRILW G+GI+QL Y E + SDHRPV
Sbjct: 1061FAPTYKYAFNSDTYYVEGVKVSKNKRRTPAWCDRILWHGRGIQQLSYVRREFKFSDHRPV 1240
Query: 556 SSIFLVDVEVI 566
+ F V+VEV+
Sbjct: 1241CATFNVEVEVM 1273
>TC208803 similar to UP|Q8H0Z6 (Q8H0Z6) At1g71710/F14O23_9, partial (29%)
Length = 923
Score = 228 bits (581), Expect = 6e-60
Identities = 108/202 (53%), Positives = 151/202 (74%)
Frame = +1
Query: 381 VSVSMSVFQSRMCFVCSHLASGQKDGAEQRRNSDVHEILQRTRFSSVFDTDQPQKIPSHD 440
VSVSMS+ Q+ CF+C+HL SG+K+G E +RN+DVH+IL+RT F S+ P+KI H+
Sbjct: 1 VSVSMSIHQTLFCFICTHLTSGEKEGDELKRNADVHDILRRTHFHSLSYIGLPKKILDHE 180
Query: 441 KIFWFGDLNYRINMSDGEIRKLVDLKKWNELMKFDQLSNELCKGHVFEGWKEGLINFPPT 500
+I WFGDLNYRIN+S+ + L+ K+W++L++ DQL EL K VF GW EG++NFPPT
Sbjct: 181 RIIWFGDLNYRINLSNVVTKDLISKKQWSKLVEKDQLIREL-KNGVFGGWSEGVLNFPPT 357
Query: 501 YKYEFNSDKHVGGNTQEGEKRRAPAWCDRILWLGKGIKQLKYQSAENQLSDHRPVSSIFL 560
YKYE NSDK+ G + + G +R+PAWCDRIL GKG++ L Y+ AE +LSDHRPV++ ++
Sbjct: 358 YKYEVNSDKYYGEDPKVG--KRSPAWCDRILSYGKGMRLLSYKRAELKLSDHRPVTATYM 531
Query: 561 VDVEVIDHRKLERAIYFASAVV 582
V+VE+ RKL+RA+ F A +
Sbjct: 532 VEVEIFSPRKLQRALTFTDAEI 597
>BE555198 similar to GP|10444263|gb inositol polyphosphate 5-phosphatase II
{Arabidopsis thaliana}, partial (18%)
Length = 421
Score = 139 bits (351), Expect(2) = 7e-54
Identities = 63/87 (72%), Positives = 76/87 (86%)
Frame = +2
Query: 507 SDKHVGGNTQEGEKRRAPAWCDRILWLGKGIKQLKYQSAENQLSDHRPVSSIFLVDVEVI 566
SD ++G N +EGEKRR+PAWCDRILWLGKGIKQL+Y+ +EN+LSDHRPVSSIF VDVEV
Sbjct: 152 SDTYIGENQKEGEKRRSPAWCDRILWLGKGIKQLEYRRSENKLSDHRPVSSIFSVDVEVF 331
Query: 567 DHRKLERAIYFASAVVHPDVFLKEDED 593
DHRKL+RA+ F +A VH ++FLKED D
Sbjct: 332 DHRKLQRALNFTNAAVHHEIFLKEDSD 412
Score = 90.1 bits (222), Expect(2) = 7e-54
Identities = 40/48 (83%), Positives = 43/48 (89%)
Frame = +1
Query: 459 IRKLVDLKKWNELMKFDQLSNELCKGHVFEGWKEGLINFPPTYKYEFN 506
+RKLV LKKW+ELM DQLSNEL GHVF+GWKEGLINFPPTYKYEFN
Sbjct: 7 VRKLVALKKWDELMNCDQLSNELRSGHVFDGWKEGLINFPPTYKYEFN 150
>TC231004 similar to UP|Q9LR47 (Q9LR47) T25N20.12, partial (30%)
Length = 860
Score = 159 bits (401), Expect = 4e-39
Identities = 78/160 (48%), Positives = 101/160 (62%), Gaps = 3/160 (1%)
Frame = +1
Query: 419 LQRTRFSSVFDTDQ---PQKIPSHDKIFWFGDLNYRINMSDGEIRKLVDLKKWNELMKFD 475
L++TRF V D PQ I HD+I W GDLNYRI +S + LV+++ W L++ D
Sbjct: 1 LKKTRFPRVQGVDNEKSPQTILEHDRIIWLGDLNYRIALSYRSAKALVEMQNWRALLEND 180
Query: 476 QLSNELCKGHVFEGWKEGLINFPPTYKYEFNSDKHVGGNTQEGEKRRAPAWCDRILWLGK 535
QL E +G F GW EG I FPPTYKY NSD++ G + EKRR PAWCDRILW G+
Sbjct: 181 QLRIEQKRGRAFVGWNEGKIYFPPTYKYSTNSDRYAGDDMHPKEKRRTPAWCDRILWYGE 360
Query: 536 GIKQLKYQSAENQLSDHRPVSSIFLVDVEVIDHRKLERAI 575
G+ QL Y E++ SDHRPV IF +VE H +L++ +
Sbjct: 361 GLHQLSYVRGESKFSDHRPVYGIFWAEVE-STHGRLKKTM 477
>BF425933 similar to GP|10444263|gb| inositol polyphosphate 5-phosphatase II
{Arabidopsis thaliana}, partial (11%)
Length = 354
Score = 147 bits (370), Expect = 2e-35
Identities = 73/91 (80%), Positives = 81/91 (88%), Gaps = 1/91 (1%)
Frame = +3
Query: 351 VQRKLRRHVHHLKVSQVGVGLMGYMGNKGSVSVSMSVFQSRMCFVCSHLASGQKDGAEQR 410
VQR+LRRH+++LKVS VGVGLMGYMGNKGSVS+SMS+F SRMCFVCSHL SGQK+GA R
Sbjct: 3 VQRRLRRHINNLKVSPVGVGLMGYMGNKGSVSISMSLFHSRMCFVCSHLTSGQKEGA*HR 182
Query: 411 RNSDVHEILQRTRF-SSVFDTDQPQKIPSHD 440
RNSDVHEIL RT F SSVFD DQPQ IPSH+
Sbjct: 183 RNSDVHEILIRTCFSSSVFDADQPQTIPSHE 275
>TC218056 similar to UP|Q9FUR2 (Q9FUR2) Inositol polyphosphate 5-phosphatase
II, partial (11%)
Length = 791
Score = 140 bits (354), Expect = 1e-33
Identities = 65/90 (72%), Positives = 76/90 (84%)
Frame = +1
Query: 502 KYEFNSDKHVGGNTQEGEKRRAPAWCDRILWLGKGIKQLKYQSAENQLSDHRPVSSIFLV 561
KYE NSD++VG +EGEKRR+PAWCDRILWLGKGIKQL+Y AE +LSDHRPVSS FLV
Sbjct: 1 KYEINSDRYVGERPKEGEKRRSPAWCDRILWLGKGIKQLQYGRAEIKLSDHRPVSSAFLV 180
Query: 562 DVEVIDHRKLERAIYFASAVVHPDVFLKED 591
+VEV DHRKL+RA+ F A VHP++FL ED
Sbjct: 181 EVEVFDHRKLKRALNFTRAAVHPEIFLDED 270
>TC209885 similar to GB|AAO64848.1|29028938|BT005913 At3g63240 {Arabidopsis
thaliana;} , partial (24%)
Length = 977
Score = 129 bits (325), Expect = 3e-30
Identities = 57/115 (49%), Positives = 79/115 (68%)
Frame = +2
Query: 460 RKLVDLKKWNELMKFDQLSNELCKGHVFEGWKEGLINFPPTYKYEFNSDKHVGGNTQEGE 519
+ LV++ W L++ DQL E +G VFEGW EG I FPPTYKY NSD++ G + +
Sbjct: 5 KALVEMHDWKTLLENDQLCIEQRQGRVFEGWNEGKIYFPPTYKYSNNSDRYAGDDRHSKQ 184
Query: 520 KRRAPAWCDRILWLGKGIKQLKYQSAENQLSDHRPVSSIFLVDVEVIDHRKLERA 574
KRR PAWCDRILW G+G+ QL Y E++ SDHRPV S+FL +VE + +++++
Sbjct: 185 KRRTPAWCDRILWYGRGLHQLSYVRGESRFSDHRPVYSMFLAEVESVSCNQIKKS 349
>BE800026 similar to GP|10444263|gb| inositol polyphosphate 5-phosphatase II
{Arabidopsis thaliana}, partial (10%)
Length = 214
Score = 128 bits (321), Expect = 8e-30
Identities = 55/71 (77%), Positives = 64/71 (89%)
Frame = +2
Query: 469 NELMKFDQLSNELCKGHVFEGWKEGLINFPPTYKYEFNSDKHVGGNTQEGEKRRAPAWCD 528
+ELM +DQLSNEL GHVF+GWKEGLINFPPTYKYE NSDK++G N EG+K+R+P+WCD
Sbjct: 2 DELMNYDQLSNELRSGHVFDGWKEGLINFPPTYKYEINSDKYIGENP*EGDKKRSPSWCD 181
Query: 529 RILWLGKGIKQ 539
RILWLGKGIKQ
Sbjct: 182 RILWLGKGIKQ 214
>BU080575
Length = 429
Score = 123 bits (309), Expect = 2e-28
Identities = 64/150 (42%), Positives = 94/150 (62%), Gaps = 1/150 (0%)
Frame = +2
Query: 7 EVFWPSTVMKKWLNVKQKVYDFSEDEANTETD-ESEDDVKLIEIETNHTSKPQANTTKPT 65
++FW VM+KWLN+ D+S D + + D ES D + + ++ +A++
Sbjct: 8 QLFWARVVMRKWLNMGSYESDYSADPVDDDDDSESGSDNEEWGRRSRFANEDEASSESTE 187
Query: 66 QLNTSCKDYEMKRHRRRKSETLRAQYINTKEVRVTIGTWNVAGKHPCNDLEIEGWLCTEE 125
L + RR+KS T R+QYIN KE+RV +GTWNV GK P +DL+I+ WL
Sbjct: 188 FL---------PKLRRQKSSTYRSQYINKKELRVCVGTWNVGGKLPPDDLDIDDWLGV-N 337
Query: 126 EPSDIYIIGFQEVVPLNAGNVFGAEDSKPI 155
EP+DIY++G QE+VPLN GN+FGAED++P+
Sbjct: 338 EPADIYVLGLQEIVPLNPGNIFGAEDTRPV 427
>TC222535 similar to UP|OCRL_HUMAN (Q01968) Inositol polyphosphate
5-phosphatase OCRL-1 (Lowe's oculocerebrorenal syndrome
protein) , partial (3%)
Length = 754
Score = 96.7 bits (239), Expect = 3e-20
Identities = 67/195 (34%), Positives = 93/195 (47%)
Frame = +2
Query: 1 MKGKRSEVFWPSTVMKKWLNVKQKVYDFSEDEANTETDESEDDVKLIEIETNHTSKPQAN 60
+K K S+ WP ++KWLN++ F D ++ S P+
Sbjct: 248 LKKKISKSSWPKFNVRKWLNIRSNDDKFHSD-------------------ASYYSLPEGW 370
Query: 61 TTKPTQLNTSCKDYEMKRHRRRKSETLRAQYINTKEVRVTIGTWNVAGKHPCNDLEIEGW 120
T E+K H E I+T +R+ +GTWNV GK P L + W
Sbjct: 371 LMDSTN--------ELK-HSASVMEAPSVIDIDTLNLRMFVGTWNVGGKSPNEGLNLRDW 523
Query: 121 LCTEEEPSDIYIIGFQEVVPLNAGNVFGAEDSKPIPKWDALIRRTLNKSSEPGTKKKSNS 180
L + +DIY+IGFQE++PLNAGNV G EDS P KW LIR+ LN ++ S+S
Sbjct: 524 LMLPSQ-ADIYVIGFQEIIPLNAGNVLGPEDSGPASKWLNLIRQALNSNT-------SSS 679
Query: 181 APPSPIRRISSFNTQ 195
SP SSFN++
Sbjct: 680 GENSP---TSSFNSR 715
>CA853487 similar to GP|10444263|gb| inositol polyphosphate 5-phosphatase II
{Arabidopsis thaliana}, partial (7%)
Length = 401
Score = 95.9 bits (237), Expect = 5e-20
Identities = 41/51 (80%), Positives = 46/51 (89%)
Frame = -1
Query: 476 QLSNELCKGHVFEGWKEGLINFPPTYKYEFNSDKHVGGNTQEGEKRRAPAW 526
QLSNEL GHVF+GWKEGLINFPPTYKYEFNSD ++G N +EGEKRR+PAW
Sbjct: 338 QLSNELRSGHVFDGWKEGLINFPPTYKYEFNSDTYIGENQKEGEKRRSPAW 186
>BI317536
Length = 440
Score = 61.2 bits (147), Expect(2) = 8e-20
Identities = 32/68 (47%), Positives = 42/68 (61%)
Frame = +2
Query: 397 SHLASGQKDGAEQRRNSDVHEILQRTRFSSVFDTDQPQKIPSHDKIFWFGDLNYRINMSD 456
SHLASG ++G E+ RNS+V EI RT F D P+ I HD + GDLNYRI++ +
Sbjct: 233 SHLASGGREGDEKHRNSNVAEIFSRTSFPRGPLLDLPRTILDHDHVILLGDLNYRISLPE 412
Query: 457 GEIRKLVD 464
R LV+
Sbjct: 413 ETTRLLVE 436
Score = 54.3 bits (129), Expect(2) = 8e-20
Identities = 24/44 (54%), Positives = 33/44 (74%)
Frame = +1
Query: 337 IVSKQMVGIYVSVWVQRKLRRHVHHLKVSQVGVGLMGYMGNKGS 380
I+SKQMVG+++SVW++R L + H VS VG G+MG +GNK S
Sbjct: 109 IISKQMVGLFISVWIRRDLCPFIRHPSVSCVGCGIMGCLGNKPS 240
>TC220937 weakly similar to GB|AAB60525.1|1166575|U45479 145 kDa synaptojanin
isoform {Rattus norvegicus;} , partial (4%)
Length = 814
Score = 89.4 bits (220), Expect = 4e-18
Identities = 71/205 (34%), Positives = 102/205 (49%), Gaps = 9/205 (4%)
Frame = +2
Query: 362 LKVSQVGVGLMGYM--GNKGSVSVSMSVFQSRMCFVCSHLASGQKDGAEQRRNSDVHEIL 419
LKV + VG G + KG+V++ ++ RM F+ HL++ ++ E RNS I
Sbjct: 125 LKVDKQSVGGCGGIIGRKKGAVAIRINYKGIRMVFISCHLSAHARNVEE--RNSQCRHI- 295
Query: 420 QRTRFSSVFDTDQPQKIPSHDKIFWFGDLNYRINMSDG-EIRKLVDLKKWNELMKFDQLS 478
+ FS ++ P PSH I W GDLNYR+ D R L++ L DQL
Sbjct: 296 SHSLFSKFWN---PYSRPSHITI-WLGDLNYRLQGIDTYPARSLIEQNLHRRLHGKDQLL 463
Query: 479 NELCKGHVFEGWKEGLINFPPTYKYEFNSDKHVGGNTQEGEKRRAPAWCDRILWLGKGIK 538
E +G +F G+ EG +NF PTYKY S N K R PAW DRIL+ +
Sbjct: 464 QEAGRGQIFNGFCEGTLNFKPTYKYNKGS-----SNYDTSHKIRVPAWTDRILFRIEDEN 628
Query: 539 QLK-----YQSAENQL-SDHRPVSS 557
+++ Y+S + SDH+PV +
Sbjct: 629 KMEATLHSYESMDEIYGSDHKPVKA 703
>BF324640 similar to GP|29028938|gb| At3g63240 {Arabidopsis thaliana},
partial (11%)
Length = 196
Score = 87.8 bits (216), Expect = 1e-17
Identities = 40/63 (63%), Positives = 50/63 (78%)
Frame = +1
Query: 355 LRRHVHHLKVSQVGVGLMGYMGNKGSVSVSMSVFQSRMCFVCSHLASGQKDGAEQRRNSD 414
+R VH++KVS VG GLMGY+GN GS+S++MS+ Q+ CF+CSHL SGQKDG E RR SD
Sbjct: 7 IRDDVHNMKVSCVGRGLMGYLGNNGSISITMSLHQTSFCFICSHLTSGQKDGDELRRTSD 186
Query: 415 VHE 417
V E
Sbjct: 187 VME 195
>TC218054 similar to UP|Q9FUR2 (Q9FUR2) Inositol polyphosphate 5-phosphatase
II, partial (7%)
Length = 537
Score = 87.8 bits (216), Expect = 1e-17
Identities = 42/59 (71%), Positives = 49/59 (82%)
Frame = +1
Query: 533 LGKGIKQLKYQSAENQLSDHRPVSSIFLVDVEVIDHRKLERAIYFASAVVHPDVFLKED 591
LGKGIKQL+Y AE +LSDHRPVSS FLV+VEV DHRKL+RA+ F A VHP++FL ED
Sbjct: 7 LGKGIKQLQYGRAEIKLSDHRPVSSAFLVEVEVFDHRKLKRALNFTRAAVHPEIFLDED 183
>BE020060
Length = 423
Score = 86.3 bits (212), Expect = 4e-17
Identities = 55/170 (32%), Positives = 77/170 (44%), Gaps = 6/170 (3%)
Frame = +2
Query: 1 MKGKRSEVFWPSTVMKKWLNVKQKVYDFSEDEANTE------TDESEDDVKLIEIETNHT 54
+K K S+ WP ++KWLN++ +F D + E T+E + ++E
Sbjct: 23 LKKKISKSSWPKFNVRKWLNIRSNDDNFHSDYSLPEGWLMDSTNELKHSASVMEA----- 187
Query: 55 SKPQANTTKPTQLNTSCKDYEMKRHRRRKSETLRAQYINTKEVRVTIGTWNVAGKHPCND 114
P N T +T +R+ +GTWNV GK P
Sbjct: 188 --PPVNDT------------------------------DTLNLRMFVGTWNVGGKSPNEG 271
Query: 115 LEIEGWLCTEEEPSDIYIIGFQEVVPLNAGNVFGAEDSKPIPKWDALIRR 164
L + WL P+DIY+IGFQE++PLNAGNV G EDS P W LI +
Sbjct: 272 LNLRNWLMLPS-PADIYVIGFQEIIPLNAGNVLGPEDSGPASTWLNLIHQ 418
>BI787769 weakly similar to GP|15081807|gb| At1g71710/F14O23_9 {Arabidopsis
thaliana}, partial (11%)
Length = 421
Score = 78.6 bits (192), Expect = 7e-15
Identities = 45/123 (36%), Positives = 64/123 (51%)
Frame = +1
Query: 7 EVFWPSTVMKKWLNVKQKVYDFSEDEANTETDESEDDVKLIEIETNHTSKPQANTTKPTQ 66
++ W VM+KWLN+ D++ D + ++ E D E P +
Sbjct: 70 QLLWARVVMRKWLNMASNEPDYTADPDDDNEEDPESDSDNEEWGKRTRFGDSREELAPIE 249
Query: 67 LNTSCKDYEMKRHRRRKSETLRAQYINTKEVRVTIGTWNVAGKHPCNDLEIEGWLCTEEE 126
N + R RR+KS T R+QYIN KE+RV +GTWNV GK P +DL+I+ WL E
Sbjct: 250 SNEF-----LPRLRRQKSLTSRSQYINKKELRVCVGTWNVGGKLPPDDLDIDDWLGI-NE 411
Query: 127 PSD 129
P+D
Sbjct: 412 PAD 420
>AW704333
Length = 431
Score = 75.5 bits (184), Expect = 6e-14
Identities = 30/48 (62%), Positives = 42/48 (87%)
Frame = +3
Query: 519 EKRRAPAWCDRILWLGKGIKQLKYQSAENQLSDHRPVSSIFLVDVEVI 566
EKRRAPAWCDRI+W G+G+KQL+Y + E++LSDHRPV ++F+ +V V+
Sbjct: 6 EKRRAPAWCDRIVWCGEGLKQLQYTTIESKLSDHRPVKAMFIAEVRVL 149
>TC232211 homologue to UP|Q9LR47 (Q9LR47) T25N20.12, partial (12%)
Length = 660
Score = 70.1 bits (170), Expect = 3e-12
Identities = 32/64 (50%), Positives = 43/64 (67%)
Frame = +2
Query: 512 GGNTQEGEKRRAPAWCDRILWLGKGIKQLKYQSAENQLSDHRPVSSIFLVDVEVIDHRKL 571
G + EKRR PAWCDRILW G+G+ QL Y E++ SDHRPV IF +VE H +L
Sbjct: 2 GDDMHPKEKRRTPAWCDRILWYGEGLHQLSYVRGESRFSDHRPVYGIFWAEVE-SSHGRL 178
Query: 572 ERAI 575
++++
Sbjct: 179 KKSM 190
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.315 0.132 0.391
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 26,486,817
Number of Sequences: 63676
Number of extensions: 348380
Number of successful extensions: 2119
Number of sequences better than 10.0: 64
Number of HSP's better than 10.0 without gapping: 2085
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2101
length of query: 599
length of database: 12,639,632
effective HSP length: 103
effective length of query: 496
effective length of database: 6,081,004
effective search space: 3016177984
effective search space used: 3016177984
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 62 (28.5 bits)
Medicago: description of AC147404.2


