

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
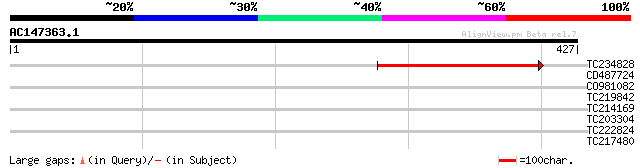
Query= AC147363.1 - phase: 0 /pseudo
(427 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC234828 105 3e-23
CD487724 39 0.004
CO981082 30 2.7
TC219842 similar to UP|Q94BY0 (Q94BY0) AT3g49400/F2K15_260 (Frag... 29 3.5
TC214169 similar to UP|Q8LF25 (Q8LF25) DNA-damage inducible prot... 29 3.5
TC203304 similar to UP|O82529 (O82529) Ribosomal protein L27a, p... 28 6.0
TC222824 weakly similar to GB|AAA98791.1|212879|CHKVITB vitellog... 28 6.0
TC217480 weakly similar to GB|AAM78037.1|21928017|AY125527 AT3g4... 28 6.0
>TC234828
Length = 857
Score = 105 bits (263), Expect = 3e-23
Identities = 55/125 (44%), Positives = 75/125 (60%)
Frame = +3
Query: 278 RVFALTGTQTENEDRLIIGTCYINNTPLVAIIDIGATHCFIAFDCASTLGLDMSNMNGEM 337
RVFA++G++ D LI G C I + L + D GATH FI+ C LGL + + +M
Sbjct: 132 RVFAMSGSEAAASDDLIRGKCLIADKLLDVLYDSGATHSFISHACVERLGLCATELPYDM 311
Query: 338 VVETPAKGSVTTSLVCLKSSLSMFGRDFEMDLVCLPLSGMYVILGMNWFE*NHVHINYFS 397
VV TP VTTS VCLK + + GR F DL+CLPL+ + VILGM+W NH+ ++
Sbjct: 312 VVSTPTSEPVTTSRVCLKCPIIVEGRSFMADLICLPLAHLDVILGMDWLSTNHIFLDCKE 491
Query: 398 KSVYF 402
K + F
Sbjct: 492 KMLVF 506
>CD487724
Length = 676
Score = 38.9 bits (89), Expect = 0.004
Identities = 27/100 (27%), Positives = 49/100 (49%), Gaps = 1/100 (1%)
Frame = +3
Query: 304 PLVAIIDIGATHCFIAFDCASTLGLDMSNMNGEMVVETPAKGSVTTSLVCLKSSLSMFGR 363
PLV ++D G+TH F+ S LGL + + V + + +C +S+
Sbjct: 21 PLVYLVDGGSTHNFVQQPLVSQLGLPCRS-TPPLRVMVGNGHHLKCTTICEAIPISIQNI 197
Query: 364 DFEMDLVCLPLSGMYVILGMNWFE-*NHVHINYFSKSVYF 402
+F + L LP+ G ++LG+ W + + ++Y S S+ F
Sbjct: 198 EFLVHLYVLPIVGANIVLGVQWLKTLGPILVDYNSLSMQF 317
>CO981082
Length = 670
Score = 29.6 bits (65), Expect = 2.7
Identities = 14/37 (37%), Positives = 21/37 (55%), Gaps = 1/37 (2%)
Frame = +1
Query: 336 EMVVETPAKGSV-TTSLVCLKSSLSMFGRDFEMDLVC 371
E+ V +P K T S++CL S +SMF F ++C
Sbjct: 229 ELWVTSPTKAKAFTMSIICLCSCISMFPGHFPFSIIC 339
>TC219842 similar to UP|Q94BY0 (Q94BY0) AT3g49400/F2K15_260 (Fragment),
partial (3%)
Length = 773
Score = 29.3 bits (64), Expect = 3.5
Identities = 23/68 (33%), Positives = 36/68 (52%), Gaps = 3/68 (4%)
Frame = -3
Query: 313 ATHCFIAFDCASTLGL---DMSNMNGEMVVETPAKGSVTTSLVCLKSSLSMFGRDFEMDL 369
A HC I CAS G D+ ++G ++ A+ TSL +SSLS+F R F +
Sbjct: 318 AIHCSI---CASPTGYQQPDLKFVSGLDMLVWKAEKLKPTSLS--RSSLSLFRRSFVHLV 154
Query: 370 VCLPLSGM 377
+C ++G+
Sbjct: 153 ICFSMAGL 130
>TC214169 similar to UP|Q8LF25 (Q8LF25) DNA-damage inducible protein
DDI1-like, partial (90%)
Length = 1604
Score = 29.3 bits (64), Expect = 3.5
Identities = 13/29 (44%), Positives = 16/29 (54%)
Frame = +2
Query: 300 INNTPLVAIIDIGATHCFIAFDCASTLGL 328
+N PL A +D GA I+ CA LGL
Sbjct: 764 VNGVPLKAFVDSGAQSTIISKSCAERLGL 850
>TC203304 similar to UP|O82529 (O82529) Ribosomal protein L27a, partial (95%)
Length = 870
Score = 28.5 bits (62), Expect = 6.0
Identities = 16/40 (40%), Positives = 22/40 (55%)
Frame = +3
Query: 5 IILLLSREDMTPMEPRRGLRRSREFFVLCNALKFRRCDLV 44
I LLL R + P PRR LR+ F + L+F+ C L+
Sbjct: 372 ISLLLLRPSLFPRSPRRRLRKPVALFF--SPLRFKNCFLI 485
>TC222824 weakly similar to GB|AAA98791.1|212879|CHKVITB vitellogenin {Gallus
gallus;} , partial (17%)
Length = 703
Score = 28.5 bits (62), Expect = 6.0
Identities = 17/41 (41%), Positives = 22/41 (53%)
Frame = +3
Query: 264 GSQCKQPKKAPTIGRVFALTGTQTENEDRLIIGTCYINNTP 304
GS+C Q KA T+G+ A TG Q E+ T Y+N P
Sbjct: 258 GSECFQASKAQTVGQEMA-TGNQEAKEETPGF-TVYVNRRP 374
>TC217480 weakly similar to GB|AAM78037.1|21928017|AY125527
AT3g46450/F18L15_170 {Arabidopsis thaliana;} , partial
(34%)
Length = 950
Score = 28.5 bits (62), Expect = 6.0
Identities = 13/45 (28%), Positives = 27/45 (59%)
Frame = -2
Query: 363 RDFEMDLVCLPLSGMYVILGMNWFE*NHVHINYFSKSVYFSSVEE 407
+ ++++L CL LSG+ I G+ ++ + N K+ Y+SS ++
Sbjct: 784 KGYKLNLTCLALSGLVAITGIKYYS-TKSYKNPKYKNAYYSSSQQ 653
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.343 0.151 0.478
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,231,590
Number of Sequences: 63676
Number of extensions: 251161
Number of successful extensions: 2040
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 2028
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2039
length of query: 427
length of database: 12,639,632
effective HSP length: 100
effective length of query: 327
effective length of database: 6,272,032
effective search space: 2050954464
effective search space used: 2050954464
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.5 bits)
S2: 60 (27.7 bits)
Medicago: description of AC147363.1


