

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147179.9 + phase: 1 /partial
(375 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
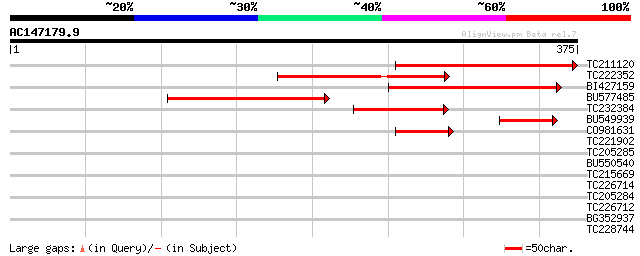
Score E
Sequences producing significant alignments: (bits) Value
TC211120 homologue to UP|Q9C5B2 (Q9C5B2) Translation releasing f... 223 1e-58
TC222352 similar to UP|Q74AS3 (Q74AS3) Peptide chain release fac... 107 1e-23
BI427159 107 1e-23
BU577485 100 2e-21
TC232384 similar to PIR|T02185|T02185 probale translation releas... 56 2e-08
BU549939 52 3e-07
CO981631 42 6e-04
TC221902 homologue to UP|Q9SI66 (Q9SI66) F23N19.20, partial (53%) 37 0.019
TC205285 35 0.072
BU550540 30 1.4
TC215669 similar to UP|Q9FMU9 (Q9FMU9) Similarity to ATFP3, part... 30 2.3
TC226714 29 3.0
TC205284 28 6.7
TC226712 28 8.8
BG352937 28 8.8
TC228744 similar to UP|Q94A21 (Q94A21) AT5g15330/F8M21_220, part... 28 8.8
>TC211120 homologue to UP|Q9C5B2 (Q9C5B2) Translation releasing factor2,
partial (24%)
Length = 571
Score = 223 bits (567), Expect = 1e-58
Identities = 109/120 (90%), Positives = 117/120 (96%)
Frame = +3
Query: 256 KVETAVRITHIPTGVTLRCTEERSQLANKIKALSRLKAKLLVIAEEQRATEFKQIRGDVV 315
+VETAVRITHIPTGVT+RCTEERSQLANKI+ALSRLKAKLLVIAEEQRA+E KQIRGD V
Sbjct: 6 RVETAVRITHIPTGVTVRCTEERSQLANKIRALSRLKAKLLVIAEEQRASEIKQIRGDAV 185
Query: 316 KAEWGQQIRNYVFHPYKLVKDVRTGHETPDITSVIDGELDPFIKSYLKHKYSMTMSTSGV 375
KAEWGQQIRNYVFHPYKLVKDVRTGHET DITSV+DGELDPFIKSYLKHKY+M++STSGV
Sbjct: 186 KAEWGQQIRNYVFHPYKLVKDVRTGHETTDITSVMDGELDPFIKSYLKHKYNMSLSTSGV 365
>TC222352 similar to UP|Q74AS3 (Q74AS3) Peptide chain release factor 2,
partial (27%)
Length = 694
Score = 107 bits (266), Expect = 1e-23
Identities = 58/114 (50%), Positives = 74/114 (64%)
Frame = +1
Query: 178 ATIEVEGRYAYGYLSGEKGTHRIVRQSPFNSKGLRQTSFSGVEVMPLLPEESLNVEIPEE 237
ATI+V+G +A+GY E G HR+V SPF+S R TSF+ V V+P+ + S +V+I E
Sbjct: 211 ATIKVDGEFAFGYAKAEIGVHRLVNISPFDSNKHRHTSFAAVAVIPIPGDGSTHVQINES 390
Query: 238 DLEISFSRAGGKGGQNVNKVETAVRITHIPTGVTLRCTEERSQLANKIKALSRL 291
DL I RA GGQ+VN E+AVRI HIPTGVT C ER Q NK A++ L
Sbjct: 391 DLRIERFRA---GGQHVNTTESAVRIVHIPTGVTATCQNERPQHQNKASAMAVL 543
>BI427159
Length = 410
Score = 107 bits (266), Expect = 1e-23
Identities = 51/115 (44%), Positives = 76/115 (65%)
Frame = -2
Query: 251 GQNVNKVETAVRITHIPTGVTLRCTEERSQLANKIKALSRLKAKLLVIAEEQRATEFKQI 310
G+N + + V I HIPTG++++ ERS ANK+KAL+RLKAKLLV +EQ K I
Sbjct: 391 GKNKRQTDHTVCIQHIPTGISVQSYGERSHFANKMKALNRLKAKLLVTTKEQGVASIKSI 212
Query: 311 RGDVVKAEWGQQIRNYVFHPYKLVKDVRTGHETPDITSVIDGELDPFIKSYLKHK 365
+ + + W ++IR YV HP+KLV DV+TG E PD+ V++G + P I +++ +
Sbjct: 211 QKENIVNLWQEEIRRYVSHPHKLVHDVKTGVEVPDLNYVLEGNIGPLIAAHINSR 47
>BU577485
Length = 409
Score = 99.8 bits (247), Expect = 2e-21
Identities = 47/107 (43%), Positives = 71/107 (65%)
Frame = +1
Query: 105 LNKSIDRFELTQLLSGPYDKEGAVISITAGAGGTDAQDWADMLLRMYVRWGEKQRYKTRV 164
+++ +D++E+++LL GP+D GA + I AG G + WA+ +L MY+RW +++ Y+ R+
Sbjct: 1 VSEIVDQYEISKLLKGPFDMAGACLVIKAGPKGIYPKLWAEQILSMYLRWAKRRGYEGRI 180
Query: 165 VEKSLGEEAGIKSATIEVEGRYAYGYLSGEKGTHRIVRQSPFNSKGL 211
V+K + GI SA IE E AYGYLSGEKG H ++R SP S L
Sbjct: 181 VDKCPFKNGGINSAIIEFEFECAYGYLSGEKGVHYLIRGSPNESSQL 321
Score = 38.9 bits (89), Expect = 0.004
Identities = 19/26 (73%), Positives = 22/26 (84%)
Frame = +3
Query: 277 ERSQLANKIKALSRLKAKLLVIAEEQ 302
ERS ANK+KAL+RLKAKLLV +EQ
Sbjct: 327 ERSHFANKMKALNRLKAKLLVTTKEQ 404
>TC232384 similar to PIR|T02185|T02185 probale translation releasing factor
RF-1 F14M4.15 - Arabidopsis thaliana {Arabidopsis
thaliana;} , partial (22%)
Length = 884
Score = 56.2 bits (134), Expect = 2e-08
Identities = 28/64 (43%), Positives = 42/64 (64%), Gaps = 1/64 (1%)
Frame = +1
Query: 228 ESLNVEIPEEDLEISFSRAGGKGGQNVNKVETAVRITHIPTGVTLRCTEERSQ-LANKIK 286
+ ++V++ EDL I R+GG GGQ+ N +AVR+THIPTG + +E SQ +A+ K
Sbjct: 55 DEVDVQLKHEDLRIDTYRSGGSGGQHANTTNSAVRVTHIPTGTMITIQDECSQHMASVSK 234
Query: 287 ALSR 290
L+R
Sbjct: 235 YLAR 246
>BU549939
Length = 670
Score = 52.4 bits (124), Expect = 3e-07
Identities = 22/38 (57%), Positives = 30/38 (78%)
Frame = -2
Query: 325 NYVFHPYKLVKDVRTGHETPDITSVIDGELDPFIKSYL 362
+YV HPY++VKD+RT +E D SV++G+LD FI SYL
Sbjct: 669 SYVLHPYRMVKDLRTNYEVSDPDSVLEGDLDSFILSYL 556
>CO981631
Length = 697
Score = 41.6 bits (96), Expect = 6e-04
Identities = 17/38 (44%), Positives = 29/38 (75%)
Frame = -2
Query: 256 KVETAVRITHIPTGVTLRCTEERSQLANKIKALSRLKA 293
K E+AVR+ H+PTG+ + +E+RSQ N+ A++RL++
Sbjct: 696 KRESAVRLKHLPTGIIAQASEDRSQHKNRASAINRLRS 583
>TC221902 homologue to UP|Q9SI66 (Q9SI66) F23N19.20, partial (53%)
Length = 621
Score = 36.6 bits (83), Expect = 0.019
Identities = 16/29 (55%), Positives = 22/29 (75%)
Frame = +3
Query: 233 EIPEEDLEISFSRAGGKGGQNVNKVETAV 261
+I + + +SF+R+GG GGQNVNKV T V
Sbjct: 288 KITLDHVTVSFARSGGPGGQNVNKVNTKV 374
>TC205285
Length = 1040
Score = 34.7 bits (78), Expect = 0.072
Identities = 23/106 (21%), Positives = 55/106 (51%)
Frame = +1
Query: 1 FYSLRKDVEIASQRVKEIRESSGLQLLEQELAKLEEEASCSSFWDDRAKAQQTLSTLADV 60
F +++ ++ +Q + + ++SG L+K +EE S S+ RAK ++ +V
Sbjct: 160 FVCVKERMQSENQNNQLVVQNSGSLSFSSHLSKEDEEISRSALSTFRAKEEEIERKKMEV 339
Query: 61 KEKIKLLNDYKTQVEDAETIVMLTEEMESVDKGLYEEASSLIKELN 106
+EK++L E+ + + + EE+E++ + +E + + K ++
Sbjct: 340 REKVQL--QLGRVEEETKRLATIREELEALADPMRKEVALVRKRID 471
>BU550540
Length = 672
Score = 30.4 bits (67), Expect = 1.4
Identities = 35/120 (29%), Positives = 56/120 (46%), Gaps = 12/120 (10%)
Frame = -2
Query: 6 KDVEIASQRVKEIRESSGLQLLEQELAKLEEEAS-----CSSFWDDRAKAQQTLSTLADV 60
KD+ Q KE RE L++ L + EE+ DR K+ +T L D
Sbjct: 638 KDLLTLEQFKKENRE------LKERLQRKEEDXXGEVLHALQHEKDRCKSLET--QLIDA 483
Query: 61 KEKIKLLNDYKTQVEDAETIV-MLTEEMESVD------KGLYEEASSLIKELNKSIDRFE 113
++K++ LN+ + ET++ + EE + D + EEAS+ IKEL + I + E
Sbjct: 482 EKKLEELNN------EQETLIDVFAEERDRRDAEEKKLRNKLEEASNTIKELLEKIRKLE 321
>TC215669 similar to UP|Q9FMU9 (Q9FMU9) Similarity to ATFP3, partial (54%)
Length = 1305
Score = 29.6 bits (65), Expect = 2.3
Identities = 16/61 (26%), Positives = 32/61 (52%)
Frame = +2
Query: 226 PEESLNVEIPEEDLEISFSRAGGKGGQNVNKVETAVRITHIPTGVTLRCTEERSQLANKI 285
PE+ VE +E+ + ++GG+G +N K E ++ T T TE+ +++ ++
Sbjct: 806 PEKKEKVEEAKEEKKEEEKKSGGEGEENKEKKEEEAKVEEATTPAT---TEDTNKVVPEV 976
Query: 286 K 286
K
Sbjct: 977 K 979
>TC226714
Length = 1721
Score = 29.3 bits (64), Expect = 3.0
Identities = 26/81 (32%), Positives = 36/81 (44%), Gaps = 2/81 (2%)
Frame = +3
Query: 6 KDVEIASQRVKEIRE--SSGLQLLEQELAKLEEEASCSSFWDDRAKAQQTLSTLADVKEK 63
K V SQ +KE E +L + A +EE S S R ++ L ++E
Sbjct: 345 KAVITRSQYIKEATELGKKARELKKAAEALHQEERSGSKGRKFRKNVKEVEKELFQLEED 524
Query: 64 IKLLNDYKTQVEDAETIVMLT 84
+KLL + Q E AET LT
Sbjct: 525 VKLLEEMYPQGEKAETTWALT 587
>TC205284
Length = 436
Score = 28.1 bits (61), Expect = 6.7
Identities = 20/77 (25%), Positives = 43/77 (54%), Gaps = 1/77 (1%)
Frame = +3
Query: 31 LAKLEEEASCSSFWDDRAKAQQTLSTLADVKEKIKL-LNDYKTQVEDAETIVMLTEEMES 89
L+K +EE S S+ RAK ++ +V+EK++L L+ + + + TI +E+E+
Sbjct: 15 LSKEDEEMSRSALSTFRAKEEEIERKKMEVREKVQLQLSRVEEETKRLATIHEEYQELEA 194
Query: 90 VDKGLYEEASSLIKELN 106
+ + +E + + K ++
Sbjct: 195 LADPMRKEVALVRKRID 245
>TC226712
Length = 1402
Score = 27.7 bits (60), Expect = 8.8
Identities = 24/76 (31%), Positives = 35/76 (45%), Gaps = 5/76 (6%)
Frame = +1
Query: 14 RVKEIRESSGLQLLEQELAKL-----EEEASCSSFWDDRAKAQQTLSTLADVKEKIKLLN 68
R + I+E++ L +EL K +EE S S R + L ++E +KLL
Sbjct: 253 RSQYIKEATELGKKAKELKKAAESLHQEERSGSKGRKFRKNVKSVEKELFQLEEDVKLLE 432
Query: 69 DYKTQVEDAETIVMLT 84
+ Q E AET LT
Sbjct: 433 EMYPQGEKAETTWALT 480
>BG352937
Length = 486
Score = 27.7 bits (60), Expect = 8.8
Identities = 13/29 (44%), Positives = 19/29 (64%)
Frame = +3
Query: 235 PEEDLEISFSRAGGKGGQNVNKVETAVRI 263
P+ LEI+ + GKGG N+N E++ RI
Sbjct: 156 PKPSLEIASALKEGKGGLNINIPESSSRI 242
>TC228744 similar to UP|Q94A21 (Q94A21) AT5g15330/F8M21_220, partial (69%)
Length = 1196
Score = 27.7 bits (60), Expect = 8.8
Identities = 26/109 (23%), Positives = 49/109 (44%), Gaps = 2/109 (1%)
Frame = +1
Query: 18 IRESSGLQLLEQELAKL--EEEASCSSFWDDRAKAQQTLSTLADVKEKIKLLNDYKTQVE 75
+++ S QLL+ ++ EE + F+ D K ++ + ++KE+I+ L + +Q E
Sbjct: 229 LQQPSSPQLLQAWFVRILNEELEKFNDFYVD--KEEEFVIRFQELKERIECLKEKSSQGE 402
Query: 76 DAETIVMLTEEMESVDKGLYEEASSLIKELNKSIDRFELTQLLSGPYDK 124
+ +EEM + K L ++ N S F + YDK
Sbjct: 403 VYTSDCEFSEEMMDIRKDLVTIHGEMVLLKNYSSLNFAGLVKILKKYDK 549
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.313 0.131 0.357
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,995,832
Number of Sequences: 63676
Number of extensions: 124469
Number of successful extensions: 469
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 468
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 468
length of query: 375
length of database: 12,639,632
effective HSP length: 99
effective length of query: 276
effective length of database: 6,335,708
effective search space: 1748655408
effective search space used: 1748655408
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 60 (27.7 bits)
Medicago: description of AC147179.9


