

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147179.5 - phase: 0
(706 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
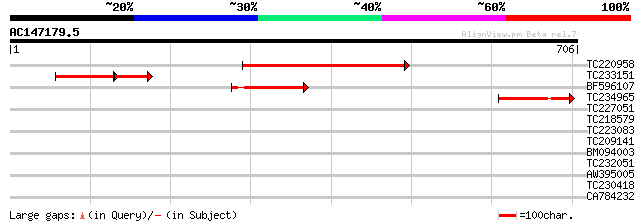
Score E
Sequences producing significant alignments: (bits) Value
TC220958 similar to UP|Q6YU42 (Q6YU42) Calcineurin-related phosp... 352 4e-97
TC233151 weakly similar to UP|Q6YU42 (Q6YU42) Calcineurin-relate... 127 4e-50
BF596107 152 6e-37
TC234965 weakly similar to UP|Q6YU42 (Q6YU42) Calcineurin-relate... 131 1e-30
TC227051 weakly similar to GB|AAM19768.1|20453045|AY094388 AT3g5... 33 0.57
TC218579 similar to UP|Q9LZ26 (Q9LZ26) Aspartyl aminopeptidase, ... 32 0.97
TC223083 31 2.2
TC209141 weakly similar to UP|Q9SW27 (Q9SW27) Geranylgeranylated... 30 2.8
BM094003 30 4.8
TC232051 29 8.2
AW395005 GP|5059126|gb| cytochrome P450 monooxygenaseCYP93D1 {Gl... 29 8.2
TC230418 weakly similar to UP|ALA9_ARATH (Q9SX33) Potential phos... 29 8.2
CA784232 29 8.2
>TC220958 similar to UP|Q6YU42 (Q6YU42) Calcineurin-related
phosphoesterase-like protein, partial (6%)
Length = 629
Score = 352 bits (902), Expect = 4e-97
Identities = 169/209 (80%), Positives = 181/209 (85%), Gaps = 1/209 (0%)
Frame = +2
Query: 290 SLQNFFQFNVHQNSFESTVNCSIGAPP-QEFWEWEIGDWRKSRAIRILAIDRGHVSYVDL 348
SLQ FQFN+HQ+SFEST NCS+GAPP QEFWEWE+GDWRKSRA RILAIDRGHVSYVD
Sbjct: 2 SLQKLFQFNIHQSSFESTANCSMGAPPIQEFWEWEMGDWRKSRAFRILAIDRGHVSYVDT 181
Query: 349 DFKSGAKHAIILPTFPLDSRFMQTSSWHHNYECQSVASSSYETIRALVFSASPVESVVAR 408
DFKS K AIILPTFPLDSRFM TSS HNYECQSVA SSYETIRALVFS SP+ SVVAR
Sbjct: 182 DFKSFTKRAIILPTFPLDSRFMLTSSCRHNYECQSVAPSSYETIRALVFSLSPIVSVVAR 361
Query: 409 VYDSRYGDLVLVVEAHMTKRADENSRGNLYVAPWNYKAFEDTSPDRFWFQIESNDIMGRS 468
VYDSR G+L LV+E M K + +NSRGNLYVAPWNY+AFED SPDRFW QIE+NDI GRS
Sbjct: 362 VYDSRSGNLDLVLETQMIKHSGDNSRGNLYVAPWNYRAFEDASPDRFWLQIEANDIAGRS 541
Query: 469 TLTELRPFSINGHSFRLSWSWKEFFVMGC 497
TLTELRPFSINGHS +LSW WKEF VMGC
Sbjct: 542 TLTELRPFSINGHSLKLSWGWKEFLVMGC 628
>TC233151 weakly similar to UP|Q6YU42 (Q6YU42) Calcineurin-related
phosphoesterase-like protein, partial (12%)
Length = 832
Score = 127 bits (320), Expect(2) = 4e-50
Identities = 63/78 (80%), Positives = 67/78 (85%)
Frame = +2
Query: 58 SDLHLSVHHPNRALDFTNLVGHALSFINPSLVLITGDLTDGKSKDLLTMKQNEDEWVEYR 117
+DLH SVHHPNRALDF LVG ALS INPSLVLITGDLTDGKSKDLLTMKQNED+WVEYR
Sbjct: 5 ADLHFSVHHPNRALDFAKLVGPALSVINPSLVLITGDLTDGKSKDLLTMKQNEDDWVEYR 184
Query: 118 NVMDGVIERSGLHKSLFF 135
V+D VI+RS L KS F
Sbjct: 185 TVLDTVIKRSRLPKSFVF 238
Score = 89.7 bits (221), Expect(2) = 4e-50
Identities = 41/47 (87%), Positives = 45/47 (95%)
Frame = +3
Query: 131 KSLFFDLRGNHDSFGVPVVGGSFDFFSKYSVNGQLGRNRSVNSVTLE 177
++LFF LRGNHDSFGVPVVGGSFDFFSKYS+NGQLGRN S+NSVTLE
Sbjct: 225 RALFFALRGNHDSFGVPVVGGSFDFFSKYSINGQLGRNGSINSVTLE 365
>BF596107
Length = 272
Score = 152 bits (383), Expect = 6e-37
Identities = 74/97 (76%), Positives = 80/97 (82%), Gaps = 1/97 (1%)
Frame = +3
Query: 277 NLKRHHQLSNRFLSLQNFFQFNVHQNSFESTVNCSIGAPP-QEFWEWEIGDWRKSRAIRI 335
NLKRHHQ Q QFN+HQ++FESTVNCS+GAPP QEFWEWE+GDWRKSRA RI
Sbjct: 3 NLKRHHQW-------QKLCQFNIHQSAFESTVNCSLGAPPIQEFWEWEMGDWRKSRAFRI 161
Query: 336 LAIDRGHVSYVDLDFKSGAKHAIILPTFPLDSRFMQT 372
LAIDR HVSYVD+DFKSG K AIILPTFPLDSRFM T
Sbjct: 162 LAIDRSHVSYVDVDFKSGTKRAIILPTFPLDSRFMLT 272
>TC234965 weakly similar to UP|Q6YU42 (Q6YU42) Calcineurin-related
phosphoesterase-like protein, partial (5%)
Length = 728
Score = 131 bits (329), Expect = 1e-30
Identities = 68/95 (71%), Positives = 78/95 (81%)
Frame = +1
Query: 609 WVGFPDIMVLVLPHILFVVLPAILVTGALTAERAIYRERVLALSGKKKDDIDLNSRRPLL 668
+VG PDIMV+VLPH+L VVLPAILVTGALTAERAIYRE +LA KKKDDI ++SR+ +L
Sbjct: 1 YVGSPDIMVVVLPHLLXVVLPAILVTGALTAERAIYREHMLAFLVKKKDDICMDSRKSVL 180
Query: 669 NGNHNSTISTLHLGKRWIRKLLCVVCLAICWKHIM 703
NG S +S +HL KR IRKLL VVCLAICWKH M
Sbjct: 181 NG---SMVSNVHLSKRLIRKLLFVVCLAICWKHFM 276
>TC227051 weakly similar to GB|AAM19768.1|20453045|AY094388
AT3g56200/F18O21_160 {Arabidopsis thaliana;} , partial
(14%)
Length = 568
Score = 32.7 bits (73), Expect = 0.57
Identities = 19/67 (28%), Positives = 33/67 (48%), Gaps = 5/67 (7%)
Frame = +1
Query: 494 VMGCQWASLYYPLLWSALGFMFSFLLVPKALLFFQMNLYTY--RNFIANKGV---VNGAL 548
V+ W SL+ P L +L S L ++ + +Y Y + ++ +GV + GA+
Sbjct: 205 VINTSWFSLFSPSLSLSLSLSLSLSLSLYIYIYIYIYIYIYYRKIYLGREGVTGEILGAI 384
Query: 549 WILQELC 555
W+L LC
Sbjct: 385 WLLWRLC 405
>TC218579 similar to UP|Q9LZ26 (Q9LZ26) Aspartyl aminopeptidase, partial
(33%)
Length = 926
Score = 32.0 bits (71), Expect = 0.97
Identities = 13/39 (33%), Positives = 22/39 (56%)
Frame = -1
Query: 284 LSNRFLSLQNFFQFNVHQNSFESTVNCSIGAPPQEFWEW 322
+SN ++ L+ +F + VH STVNCS ++W +
Sbjct: 518 VSNVYIFLRTYFPYTVH*KLSNSTVNCSDANSRSKYWAY 402
>TC223083
Length = 243
Score = 30.8 bits (68), Expect = 2.2
Identities = 23/57 (40%), Positives = 25/57 (43%), Gaps = 6/57 (10%)
Frame = +2
Query: 240 PLSFSAPSSSGRTLKEVFLKHSISAYLCGHLHS------RFGKNLKRHHQLSNRFLS 290
PL F A SSSGR L V L H + C H+H F N HQ FLS
Sbjct: 74 PLPFIA-SSSGRRLGVVGLTHHHNNPFCSHVHRVKPKPLFFSPNPTNEHQQKQSFLS 241
>TC209141 weakly similar to UP|Q9SW27 (Q9SW27) Geranylgeranylated protein
ATGP4 (At4g24990), complete
Length = 891
Score = 30.4 bits (67), Expect = 2.8
Identities = 22/66 (33%), Positives = 32/66 (48%), Gaps = 14/66 (21%)
Frame = -1
Query: 248 SSGRTLKEVFLKHSISAYLCG-----------HLHSRFGKNLKRHHQLSNRFLSLQNF-- 294
S RT +F H I+ +LC H ++ FGK KR+ L+N + LQNF
Sbjct: 552 SQYRTCTYMFSGHLINFFLCFCFEQRWLDHNMHGYNPFGKLAKRYSTLANCLVVLQNFAS 373
Query: 295 -FQFNV 299
+QF +
Sbjct: 372 TYQFYI 355
>BM094003
Length = 429
Score = 29.6 bits (65), Expect = 4.8
Identities = 29/111 (26%), Positives = 47/111 (42%), Gaps = 14/111 (12%)
Frame = +3
Query: 465 MGRSTLTELRPFSIN------------GHSFRLSWSWKEFFVMGCQWASLYYPLLWSALG 512
+G TEL PF N GHS RLS W E F+ C S ++ L++
Sbjct: 3 VGHFQPTELLPFPRNQPGLRPQVLLLKGHSIRLSGKWIEKFLRTC---SDHFYLVFQVQP 173
Query: 513 FMFSFLLVPKALLFFQMNLYTYRNFIANKGVVNGAL--WILQELCRVPTLW 561
FM ++ L+ ++L A + ++ L WIL+ + + +W
Sbjct: 174 FMLEKQILHIVPLYLGIHLLQQ---AAMRALIKVQLLHWILKGVTIIKMIW 317
>TC232051
Length = 708
Score = 28.9 bits (63), Expect = 8.2
Identities = 12/29 (41%), Positives = 18/29 (61%)
Frame = -1
Query: 274 FGKNLKRHHQLSNRFLSLQNFFQFNVHQN 302
+ KNL HQL+N+ + + F+FN QN
Sbjct: 630 YSKNLTLTHQLTNKIHTTRTGFEFNEEQN 544
>AW395005 GP|5059126|gb| cytochrome P450 monooxygenaseCYP93D1 {Glycine max},
partial (21%)
Length = 419
Score = 28.9 bits (63), Expect = 8.2
Identities = 19/55 (34%), Positives = 26/55 (46%), Gaps = 1/55 (1%)
Frame = +3
Query: 505 PLLWSALGFMFSFLLVPKALLFFQMNLYTYRNFIA-NKGVVNGALWILQELCRVP 558
PLLW G +F F ++ ++ L+T FIA G + W L EL R P
Sbjct: 60 PLLWYVCGNLFLFFVLRSNK*NLKLILHTQDMFIAGTNGPASVLKWSLAELVRNP 224
>TC230418 weakly similar to UP|ALA9_ARATH (Q9SX33) Potential
phospholipid-transporting ATPase 9 (Aminophospholipid
flippase 9) , partial (19%)
Length = 1062
Score = 28.9 bits (63), Expect = 8.2
Identities = 18/57 (31%), Positives = 30/57 (52%), Gaps = 5/57 (8%)
Frame = -1
Query: 17 CSSMNLFIATIVIAFRIYC--CNGETVIEPKGTPDS-AVWVVQLSD--LHLSVHHPN 68
C S N + R+YC C E + E +PD ++V +++D HL++HHP+
Sbjct: 438 CKSFNEALIGCG*EGRVYCSICQEEYMPE*NASPDEYMLYVSEVTDR*CHLAIHHPH 268
>CA784232
Length = 409
Score = 28.9 bits (63), Expect = 8.2
Identities = 15/27 (55%), Positives = 18/27 (66%), Gaps = 1/27 (3%)
Frame = +3
Query: 513 FMFSFLLVPKA-LLFFQMNLYTYRNFI 538
F FSFLL+ LL QMN+ T+ NFI
Sbjct: 87 FFFSFLLITHFFLLSIQMNIITHNNFI 167
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.324 0.138 0.439
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 37,471,979
Number of Sequences: 63676
Number of extensions: 593124
Number of successful extensions: 4476
Number of sequences better than 10.0: 27
Number of HSP's better than 10.0 without gapping: 4416
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 4468
length of query: 706
length of database: 12,639,632
effective HSP length: 104
effective length of query: 602
effective length of database: 6,017,328
effective search space: 3622431456
effective search space used: 3622431456
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 62 (28.5 bits)
Medicago: description of AC147179.5


