

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147178.10 + phase: 0
(161 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
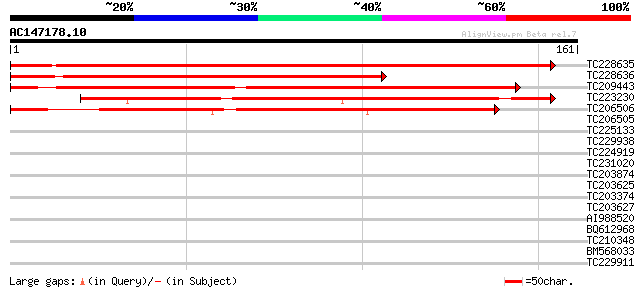
Sequences producing significant alignments: (bits) Value
TC228635 similar to UP|Q6NLX7 (Q6NLX7) At4g36660, partial (83%) 268 6e-73
TC228636 similar to UP|Q6NLX7 (Q6NLX7) At4g36660, partial (51%) 201 1e-52
TC209443 similar to UP|Q6NLX7 (Q6NLX7) At4g36660, partial (64%) 199 6e-52
TC223230 similar to UP|Q9LSL0 (Q9LSL0) Emb|CAB16809.1, partial (... 144 1e-35
TC206506 similar to UP|Q9FYX2 (Q9FYX2) BAC19.4, partial (33%) 143 4e-35
TC206505 similar to UP|Q6NLX7 (Q6NLX7) At4g36660, partial (11%) 32 0.13
TC225133 UP|Q8JUF1 (Q8JUF1) Large polyprotein 2, complete 29 0.85
TC229938 similar to UP|Q9FKW9 (Q9FKW9) Alpha-mannosidase, partia... 28 1.4
TC224919 similar to UP|O04945 (O04945) Enoyl-ACP reductase precu... 28 2.5
TC231020 27 3.2
TC203874 GB|AAA34124.1|170354|TOBUBI4A pentameric polyubiquitin ... 27 4.2
TC203625 UP|Q39257 (Q39257) Ubiquitin, partial (83%) 27 4.2
TC203374 GB|AAA34124.1|170354|TOBUBI4A pentameric polyubiquitin ... 27 4.2
TC203627 UP|Q41067 (Q41067) Polyubiquitin, partial (47%) 27 4.2
AI988520 similar to GP|1332579|emb polyubiquitin {Pinus sylvestr... 27 5.5
BQ612968 similar to GP|15983390|gb At2g47790/F17A22.18 {Arabidop... 26 7.2
TC210348 similar to UP|Q9FGX6 (Q9FGX6) Gb|AAF36742.1, partial (65%) 26 9.4
BM568033 homologue to GP|13877525|gb| S18.A ribosomal protein {A... 26 9.4
TC229911 similar to UP|Q9FMK9 (Q9FMK9) Emb|CAB76911.1 (At5g63140... 26 9.4
>TC228635 similar to UP|Q6NLX7 (Q6NLX7) At4g36660, partial (83%)
Length = 858
Score = 268 bits (686), Expect = 6e-73
Identities = 129/155 (83%), Positives = 143/155 (92%)
Frame = +2
Query: 1 MKDDDGLPTTTAAINKKENMDSGLFGKGRYKFWALAAILLLAFWSMFTGTVSLRWSGNLN 60
MKD+DGLPTTTA + KKEN+DS LFGKGRYKFWALAAILLLAFWSMFTGTVSLRWSG LN
Sbjct: 74 MKDEDGLPTTTA-VTKKENLDSSLFGKGRYKFWALAAILLLAFWSMFTGTVSLRWSGTLN 250
Query: 61 TLSSDLDTPIHDDLDVLEMEEREKVVRHMWDVYTNSRRIRLPRFWQEAFEAAYEELTSDV 120
+LS+D+DTPIHDDLDVLEMEEREKVVRHMWDVYTNSR +RLPRFWQEAFEAAYE+LTSDV
Sbjct: 251 SLSNDIDTPIHDDLDVLEMEEREKVVRHMWDVYTNSRSVRLPRFWQEAFEAAYEDLTSDV 430
Query: 121 PGVRDDAITEIAKMSVRSVNYDPPPIQSTVSFPFS 155
VRD A+ EIAKMSVRS+++DPPP+QST + FS
Sbjct: 431 AEVRDAAVAEIAKMSVRSIHFDPPPLQSTSAREFS 535
>TC228636 similar to UP|Q6NLX7 (Q6NLX7) At4g36660, partial (51%)
Length = 446
Score = 201 bits (512), Expect = 1e-52
Identities = 96/107 (89%), Positives = 101/107 (93%)
Frame = +1
Query: 1 MKDDDGLPTTTAAINKKENMDSGLFGKGRYKFWALAAILLLAFWSMFTGTVSLRWSGNLN 60
MKD+DGLPTT A KKE+MDS LFGKGRYKFWALAAILLLAFWSMFTGTVSLRWSG LN
Sbjct: 130 MKDEDGLPTTAAT--KKESMDSSLFGKGRYKFWALAAILLLAFWSMFTGTVSLRWSGTLN 303
Query: 61 TLSSDLDTPIHDDLDVLEMEEREKVVRHMWDVYTNSRRIRLPRFWQE 107
+LSSD+DTPIHDDLDVLEMEEREKVVRHMWDVYTNSRR+RLPRFWQE
Sbjct: 304 SLSSDIDTPIHDDLDVLEMEEREKVVRHMWDVYTNSRRVRLPRFWQE 444
>TC209443 similar to UP|Q6NLX7 (Q6NLX7) At4g36660, partial (64%)
Length = 1078
Score = 199 bits (505), Expect = 6e-52
Identities = 100/145 (68%), Positives = 114/145 (77%)
Frame = +2
Query: 1 MKDDDGLPTTTAAINKKENMDSGLFGKGRYKFWALAAILLLAFWSMFTGTVSLRWSGNLN 60
MKD + P I KKE DSGLFG+G YKFWALAA+LLLAFWSMFTGTVSLRWSG LN
Sbjct: 140 MKDKETFP-----IAKKEASDSGLFGRGSYKFWALAAMLLLAFWSMFTGTVSLRWSGTLN 304
Query: 61 TLSSDLDTPIHDDLDVLEMEEREKVVRHMWDVYTNSRRIRLPRFWQEAFEAAYEELTSDV 120
T S +HD LDVLE+EEREK+VRHMWDVYTN+RRIRLPRFW++AFEAAYE+LTSD
Sbjct: 305 TFSH---ASLHDHLDVLEIEEREKLVRHMWDVYTNNRRIRLPRFWEKAFEAAYEDLTSDA 475
Query: 121 PGVRDDAITEIAKMSVRSVNYDPPP 145
VRD A+ EI MS+RS++ PP
Sbjct: 476 ADVRDVAVAEIVNMSLRSLDLQLPP 550
>TC223230 similar to UP|Q9LSL0 (Q9LSL0) Emb|CAB16809.1, partial (43%)
Length = 715
Score = 144 bits (364), Expect = 1e-35
Identities = 79/138 (57%), Positives = 101/138 (72%), Gaps = 3/138 (2%)
Frame = +1
Query: 21 DSGLFGKGRYKF-WALAAILLLAFWSMFTGTVSLRWSGNLNTLSSDLDTPIHDDLDVLEM 79
DSG R+ W LAA++L+A WSMF G+V+L+WS + N DLD+ I +DLDVLE+
Sbjct: 52 DSGNTNSKRHLLLWILAAVILIALWSMFAGSVTLKWSLSNN---DDLDSTILEDLDVLEV 222
Query: 80 EEREKVVRHMWDVY--TNSRRIRLPRFWQEAFEAAYEELTSDVPGVRDDAITEIAKMSVR 137
EEREKVVRHMWD+Y TN++ + LP+FW EAFEAAY++L SDVP VRD A++EIAKMS+R
Sbjct: 223 EEREKVVRHMWDLYSHTNTKSVGLPKFWWEAFEAAYQQLVSDVPAVRDAAVSEIAKMSLR 402
Query: 138 SVNYDPPPIQSTVSFPFS 155
S+ P IQS S S
Sbjct: 403 SL---PIQIQSHSSITAS 447
>TC206506 similar to UP|Q9FYX2 (Q9FYX2) BAC19.4, partial (33%)
Length = 865
Score = 143 bits (360), Expect = 4e-35
Identities = 77/143 (53%), Positives = 96/143 (66%), Gaps = 4/143 (2%)
Frame = +2
Query: 1 MKDDDGLPTTTAAINKKENMDSGLFGKGRYKFWALAAILLLAFWSMFTGTVSLRWS-GNL 59
MKD+DG P + GKG+YK W L AI+LLA WSMFT +++L+WS GNL
Sbjct: 83 MKDEDGTPQSA--------------GKGQYKLWVLGAIILLALWSMFTASLTLKWSAGNL 220
Query: 60 NTLSSDLDTPIHDDLDVLEMEEREKVVRHMWDVYTNSRRIR---LPRFWQEAFEAAYEEL 116
N D D+ +D DVLE+EEREKVVR MWDVYT S+ R LPRFW +AF AAY+ L
Sbjct: 221 NR---DFDSATLNDFDVLEVEEREKVVRRMWDVYTQSKSGRGSGLPRFWSDAFHAAYDHL 391
Query: 117 TSDVPGVRDDAITEIAKMSVRSV 139
SDV VRD A++EIAK+S+ S+
Sbjct: 392 VSDVQSVRDAAVSEIAKISLHSL 460
>TC206505 similar to UP|Q6NLX7 (Q6NLX7) At4g36660, partial (11%)
Length = 482
Score = 32.0 bits (71), Expect = 0.13
Identities = 15/26 (57%), Positives = 20/26 (76%)
Frame = +1
Query: 114 EELTSDVPGVRDDAITEIAKMSVRSV 139
+ L SDV VRD A++EIAKMS+ S+
Sbjct: 13 QALVSDVRSVRDAAVSEIAKMSLHSL 90
>TC225133 UP|Q8JUF1 (Q8JUF1) Large polyprotein 2, complete
Length = 4042
Score = 29.3 bits (64), Expect = 0.85
Identities = 15/27 (55%), Positives = 16/27 (58%), Gaps = 4/27 (14%)
Frame = +2
Query: 11 TAAINKKENMD----SGLFGKGRYKFW 33
TA I K N+ SGLF GRYKFW
Sbjct: 635 TAEIKYKRNLSNIHISGLFYDGRYKFW 715
>TC229938 similar to UP|Q9FKW9 (Q9FKW9) Alpha-mannosidase, partial (18%)
Length = 883
Score = 28.5 bits (62), Expect = 1.4
Identities = 16/39 (41%), Positives = 21/39 (53%)
Frame = +1
Query: 44 WSMFTGTVSLRWSGNLNTLSSDLDTPIHDDLDVLEMEER 82
WS FT S WSGNL ++ S T L+VL ++R
Sbjct: 169 WSWFT-XASYNWSGNLFSILSGFHT*ELGKLEVLSFDKR 282
>TC224919 similar to UP|O04945 (O04945) Enoyl-ACP reductase precursor ,
partial (89%)
Length = 1597
Score = 27.7 bits (60), Expect = 2.5
Identities = 17/41 (41%), Positives = 21/41 (50%)
Frame = +3
Query: 36 AAILLLAFWSMFTGTVSLRWSGNLNTLSSDLDTPIHDDLDV 76
AA L S TGTV L LN + +D+PI DLD+
Sbjct: 1089 AAFLASPLASAITGTV-LYVDNGLNAMGVGVDSPIFKDLDI 1208
>TC231020
Length = 704
Score = 27.3 bits (59), Expect = 3.2
Identities = 15/52 (28%), Positives = 24/52 (45%), Gaps = 3/52 (5%)
Frame = -3
Query: 7 LPTTTAAI---NKKENMDSGLFGKGRYKFWALAAILLLAFWSMFTGTVSLRW 55
+PT T + +KE + G+ +G ++ W WS+ TG LRW
Sbjct: 288 IPTPTKVMLGSARKETREGGI--EGEWRIWKTRQRSATQSWSVATGFCMLRW 139
>TC203874 GB|AAA34124.1|170354|TOBUBI4A pentameric polyubiquitin {Nicotiana
sylvestris;} , partial (42%)
Length = 482
Score = 26.9 bits (58), Expect = 4.2
Identities = 13/32 (40%), Positives = 18/32 (55%)
Frame = -1
Query: 32 FWALAAILLLAFWSMFTGTVSLRWSGNLNTLS 63
FW L + +L S F +SL WSG + +LS
Sbjct: 290 FWML*SARVLPSSSCFPAKISLCWSGGIPSLS 195
>TC203625 UP|Q39257 (Q39257) Ubiquitin, partial (83%)
Length = 1427
Score = 26.9 bits (58), Expect = 4.2
Identities = 13/32 (40%), Positives = 18/32 (55%)
Frame = -2
Query: 32 FWALAAILLLAFWSMFTGTVSLRWSGNLNTLS 63
FW L + +L S F +SL WSG + +LS
Sbjct: 727 FWML*SARVLPSSSCFPAKISLCWSGGIPSLS 632
>TC203374 GB|AAA34124.1|170354|TOBUBI4A pentameric polyubiquitin {Nicotiana
sylvestris;} , partial (40%)
Length = 654
Score = 26.9 bits (58), Expect = 4.2
Identities = 13/32 (40%), Positives = 18/32 (55%)
Frame = -2
Query: 32 FWALAAILLLAFWSMFTGTVSLRWSGNLNTLS 63
FW L + +L S F +SL WSG + +LS
Sbjct: 182 FWML*SARVLPSSSCFPAKISLCWSGGIPSLS 87
>TC203627 UP|Q41067 (Q41067) Polyubiquitin, partial (47%)
Length = 1191
Score = 26.9 bits (58), Expect = 4.2
Identities = 13/32 (40%), Positives = 18/32 (55%)
Frame = -3
Query: 32 FWALAAILLLAFWSMFTGTVSLRWSGNLNTLS 63
FW L + +L S F +SL WSG + +LS
Sbjct: 502 FWIL*SAKVLPSSSCFPAKISLCWSGGIPSLS 407
>AI988520 similar to GP|1332579|emb polyubiquitin {Pinus sylvestris},
partial (16%)
Length = 451
Score = 26.6 bits (57), Expect = 5.5
Identities = 14/38 (36%), Positives = 20/38 (51%)
Frame = +3
Query: 26 GKGRYKFWALAAILLLAFWSMFTGTVSLRWSGNLNTLS 63
G+G FW L++ +L S +SL WSG + LS
Sbjct: 18 GEGSDFFWMLSSARVLPSSSCLPVKISLCWSGGMPYLS 131
>BQ612968 similar to GP|15983390|gb At2g47790/F17A22.18 {Arabidopsis
thaliana}, partial (35%)
Length = 444
Score = 26.2 bits (56), Expect = 7.2
Identities = 15/52 (28%), Positives = 23/52 (43%), Gaps = 11/52 (21%)
Frame = +3
Query: 10 TTAAINKKENMDS-----------GLFGKGRYKFWALAAILLLAFWSMFTGT 50
TT IN ++++S G+FG+ K W L I L W+ G+
Sbjct: 228 TTGDINDDDHLESVINMGTSIAKVGIFGETFQKLWCLTHIETLGIWNWKDGS 383
>TC210348 similar to UP|Q9FGX6 (Q9FGX6) Gb|AAF36742.1, partial (65%)
Length = 644
Score = 25.8 bits (55), Expect = 9.4
Identities = 10/39 (25%), Positives = 20/39 (50%)
Frame = +3
Query: 80 EEREKVVRHMWDVYTNSRRIRLPRFWQEAFEAAYEELTS 118
E+R+ + R ++D+ I + W+ E ++LTS
Sbjct: 174 EQRQTITRTLFDIIKEHGPITVANTWERVQEVGLKDLTS 290
>BM568033 homologue to GP|13877525|gb| S18.A ribosomal protein {Arabidopsis
thaliana}, partial (36%)
Length = 284
Score = 25.8 bits (55), Expect = 9.4
Identities = 11/34 (32%), Positives = 18/34 (52%)
Frame = +3
Query: 57 GNLNTLSSDLDTPIHDDLDVLEMEEREKVVRHMW 90
G +S+ LD + DDL+ L+ + +RH W
Sbjct: 39 GYSQVVSNALDMKLRDDLERLKKIRNHRGLRHYW 140
>TC229911 similar to UP|Q9FMK9 (Q9FMK9) Emb|CAB76911.1 (At5g63140), partial
(45%)
Length = 778
Score = 25.8 bits (55), Expect = 9.4
Identities = 9/16 (56%), Positives = 11/16 (68%)
Frame = -1
Query: 143 PPPIQSTVSFPFSTHG 158
PPP+ S + PF THG
Sbjct: 439 PPPLLSWATLPFHTHG 392
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.318 0.133 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,160,613
Number of Sequences: 63676
Number of extensions: 83775
Number of successful extensions: 508
Number of sequences better than 10.0: 39
Number of HSP's better than 10.0 without gapping: 491
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 501
length of query: 161
length of database: 12,639,632
effective HSP length: 90
effective length of query: 71
effective length of database: 6,908,792
effective search space: 490524232
effective search space used: 490524232
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 55 (25.8 bits)
Medicago: description of AC147178.10


