

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147014.15 - phase: 0
(138 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
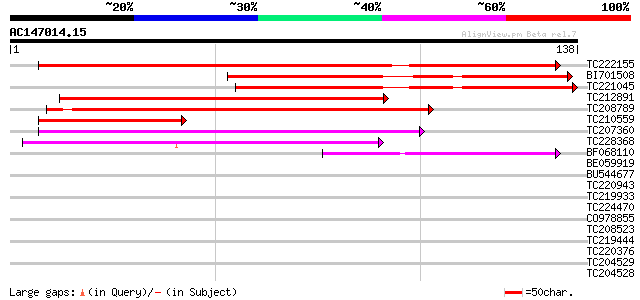
Sequences producing significant alignments: (bits) Value
TC222155 weakly similar to UP|O81662 (O81662) Transcription acti... 114 2e-26
BI701508 94 2e-20
TC221045 similar to UP|Q41275 (Q41275) Transcription factor SaMA... 92 8e-20
TC212891 homologue to UP|Q9AR13 (Q9AR13) MADS-box transcription ... 69 7e-13
TC208789 similar to UP|Q9SBQ1 (Q9SBQ1) MADS box transcription fa... 68 1e-12
TC210559 similar to UP|Q84MI1 (Q84MI1) MADS-box protein (Fragmen... 64 2e-11
TC207360 similar to UP|Q8VWZ3 (Q8VWZ3) C-type MADS box protein, ... 52 9e-08
TC228368 similar to UP|O49173 (O49173) MADS-box protein NMH 7, p... 43 5e-05
BF068110 similar to GP|27804375|gb| MADS-box transcription facto... 42 9e-05
BE059919 similar to GP|602900|emb|C SLM1 {Silene latifolia}, par... 39 0.001
BU544677 weakly similar to SP|Q38840|AG17_ Agamous-like MADS box... 37 0.002
TC220943 weakly similar to UP|Q9ZSL4 (Q9ZSL4) MADS box protein (... 36 0.005
TC219933 weakly similar to UP|Q9EQF1 (Q9EQF1) GABA-A epsilon sub... 35 0.015
TC224470 similar to UP|Q9AR13 (Q9AR13) MADS-box transcription fa... 34 0.019
CO978855 32 0.096
TC208523 similar to UP|TOB1_MOUSE (Q61471) Tob1 protein (Transdu... 30 0.37
TC219444 weakly similar to UP|Q9FXT6 (Q9FXT6) PEX14 protein, par... 30 0.37
TC220376 30 0.48
TC204529 30 0.48
TC204528 30 0.48
>TC222155 weakly similar to UP|O81662 (O81662) Transcription activator,
partial (65%)
Length = 915
Score = 114 bits (284), Expect = 2e-26
Identities = 64/127 (50%), Positives = 84/127 (65%)
Frame = +2
Query: 8 QNLKHETASLMKKIELLEASKRKLMGEGLGSCSLDELQQIEQQLEKSVSVVRARKNQAYK 67
Q LK ++ASL KKIELLE SKR+L+G+ + SCS DEL+ IE+QL+ S+ VR RK Q Y
Sbjct: 410 QQLKLDSASLAKKIELLEHSKRELLGQSVSSCSYDELKGIEEQLQISLQRVRQRKTQLYT 589
Query: 68 HQIDQLKEKEKNLVAENARLSKQVKRNYCPPQPQPQPTTKDHQREDQQPYAESSPSSDVV 127
QIDQL+ +E NL+ ENA+LS +R + Q H + + +P+ SS S DV
Sbjct: 590 EQIDQLRSQESNLLKENAKLSAMWQR----AEKSSQQQWPRHTQAEAEPHCSSSQSLDVD 757
Query: 128 TELFIGL 134
TELFIGL
Sbjct: 758 TELFIGL 778
>BI701508
Length = 360
Score = 94.0 bits (232), Expect = 2e-20
Identities = 56/84 (66%), Positives = 61/84 (71%)
Frame = +1
Query: 54 SVSVVRARKNQAYKHQIDQLKEKEKNLVAENARLSKQVKRNYCPPQPQPQPTTKDHQRED 113
SVS VRARKNQ YK QIDQLKEKE+ L AENARL +Q QPQP TKD ++
Sbjct: 1 SVSNVRARKNQVYKEQIDQLKEKERALYAENARLCEQY-------GIQPQPATKD--PKE 153
Query: 114 QQPYAESSPSSDVVTELFIGLHRS 137
QPYAESSPSS+V TELFIGL RS
Sbjct: 154 IQPYAESSPSSEVETELFIGLPRS 225
>TC221045 similar to UP|Q41275 (Q41275) Transcription factor SaMADS A,
partial (24%)
Length = 409
Score = 92.0 bits (227), Expect = 8e-20
Identities = 55/83 (66%), Positives = 60/83 (72%)
Frame = +1
Query: 56 SVVRARKNQAYKHQIDQLKEKEKNLVAENARLSKQVKRNYCPPQPQPQPTTKDHQREDQQ 115
S VRARKNQ YK QIDQLKEKE+ L AENARL +Q QPQP TKD ++ Q
Sbjct: 10 SSVRARKNQVYKEQIDQLKEKERALYAENARLCEQY------GGIQPQPATKD--PKEIQ 165
Query: 116 PYAESSPSSDVVTELFIGLHRSS 138
PYAESSPSS+V TELFIGL RSS
Sbjct: 166 PYAESSPSSEVETELFIGLLRSS 234
>TC212891 homologue to UP|Q9AR13 (Q9AR13) MADS-box transcription factor
MADS4, partial (57%)
Length = 688
Score = 68.9 bits (167), Expect = 7e-13
Identities = 35/80 (43%), Positives = 52/80 (64%)
Frame = +3
Query: 13 ETASLMKKIELLEASKRKLMGEGLGSCSLDELQQIEQQLEKSVSVVRARKNQAYKHQIDQ 72
E L KI+LL+ + R MGE LGS SL ELQ +EQQL+ ++ +R R+NQ I +
Sbjct: 15 EYTRLKAKIDLLQRNHRHYMGEDLGSMSLKELQSLEQQLDTALKQIRTRRNQLMYESISE 194
Query: 73 LKEKEKNLVAENARLSKQVK 92
L++KEK + +N L+K++K
Sbjct: 195LQKKEKVIQEQNNMLAKKIK 254
>TC208789 similar to UP|Q9SBQ1 (Q9SBQ1) MADS box transcription factor,
partial (72%)
Length = 995
Score = 68.2 bits (165), Expect = 1e-12
Identities = 36/94 (38%), Positives = 58/94 (61%)
Frame = +3
Query: 10 LKHETASLMKKIELLEASKRKLMGEGLGSCSLDELQQIEQQLEKSVSVVRARKNQAYKHQ 69
++HE L ++E+L+ ++R MGE L S +L LQ +EQQL+ ++ +R+RKNQA
Sbjct: 495 IEHE--KLKARVEVLQRNQRNFMGEDLDSLNLRGLQSLEQQLDSALKHIRSRKNQAMNES 668
Query: 70 IDQLKEKEKNLVAENARLSKQVKRNYCPPQPQPQ 103
I +L++K++ L N LSK++K PQ Q
Sbjct: 669 ISELQKKDRTLREHNNLLSKKIKEKEKELTPQEQ 770
>TC210559 similar to UP|Q84MI1 (Q84MI1) MADS-box protein (Fragment), partial
(64%)
Length = 710
Score = 64.3 bits (155), Expect = 2e-11
Identities = 31/36 (86%), Positives = 35/36 (97%)
Frame = +1
Query: 8 QNLKHETASLMKKIELLEASKRKLMGEGLGSCSLDE 43
Q+LK ETA+LMKKIELLEASKRKL+GEGLGSCSL+E
Sbjct: 601 QHLKQETANLMKKIELLEASKRKLLGEGLGSCSLEE 708
>TC207360 similar to UP|Q8VWZ3 (Q8VWZ3) C-type MADS box protein, partial
(88%)
Length = 1075
Score = 52.0 bits (123), Expect = 9e-08
Identities = 29/94 (30%), Positives = 54/94 (56%)
Frame = +2
Query: 8 QNLKHETASLMKKIELLEASKRKLMGEGLGSCSLDELQQIEQQLEKSVSVVRARKNQAYK 67
Q + E++ L ++I ++ R ++GE LGS SL EL+ +E +LEK +S VR+RK++
Sbjct: 356 QFYQQESSKLRRQIRDIQNLNRHILGEALGSLSLKELKNLEGRLEKGLSRVRSRKHETLF 535
Query: 68 HQIDQLKEKEKNLVAENARLSKQVKRNYCPPQPQ 101
++ ++++E L N L ++ + Q Q
Sbjct: 536 ADVEFMQKREIELQNHNNYLRAKIAEHERAQQQQ 637
>TC228368 similar to UP|O49173 (O49173) MADS-box protein NMH 7, partial (97%)
Length = 1042
Score = 42.7 bits (99), Expect = 5e-05
Identities = 22/92 (23%), Positives = 51/92 (54%), Gaps = 4/92 (4%)
Frame = +3
Query: 4 LNIWQNLKHETASLMKKIELLEASKRKLMGEGLGSC----SLDELQQIEQQLEKSVSVVR 59
+++W + +KK++ + + RK + + +G C +++L+ +E++++K+ VVR
Sbjct: 309 VDLWNSHYENMQENLKKLKEVNRNLRKEIRQRMGDCLNELGMEDLKLLEEEMDKAAKVVR 488
Query: 60 ARKNQAYKHQIDQLKEKEKNLVAENARLSKQV 91
RK + +QID ++K N + RL + +
Sbjct: 489 ERKYKVITNQIDTQRKKFNNEKEVHNRLLRDL 584
>BF068110 similar to GP|27804375|gb| MADS-box transcription factor CDM36
{Chrysanthemum x morifolium}, partial (9%)
Length = 415
Score = 42.0 bits (97), Expect = 9e-05
Identities = 27/58 (46%), Positives = 34/58 (58%)
Frame = +2
Query: 77 EKNLVAENARLSKQVKRNYCPPQPQPQPTTKDHQREDQQPYAESSPSSDVVTELFIGL 134
EK L+ ENA+L++ + + QP TK+ Q AESS SSDV TELFIGL
Sbjct: 2 EKALLDENAKLTENARLSE-KHDIHLQPATKNQNVNQPQCNAESSSSSDVETELFIGL 172
>BE059919 similar to GP|602900|emb|C SLM1 {Silene latifolia}, partial (31%)
Length = 426
Score = 38.5 bits (88), Expect = 0.001
Identities = 18/58 (31%), Positives = 37/58 (63%)
Frame = +2
Query: 34 EGLGSCSLDELQQIEQQLEKSVSVVRARKNQAYKHQIDQLKEKEKNLVAENARLSKQV 91
EGL + + +L+ +E +LEK +S +R++KN+ +I+ +K++E L +N L ++
Sbjct: 5 EGLSTMNGKDLKNLETKLEKGISRIRSKKNEMLFAEIEHMKKREIYLHNDNQLLRAKI 178
>BU544677 weakly similar to SP|Q38840|AG17_ Agamous-like MADS box protein
AGL17. [Mouse-ear cress] {Arabidopsis thaliana}, partial
(38%)
Length = 653
Score = 37.4 bits (85), Expect = 0.002
Identities = 19/63 (30%), Positives = 36/63 (56%)
Frame = -1
Query: 31 LMGEGLGSCSLDELQQIEQQLEKSVSVVRARKNQAYKHQIDQLKEKEKNLVAENARLSKQ 90
LMGE L + ELQ +E S+ V +K+Q ++I++L++K + EN L ++
Sbjct: 653 LMGEELMGLGIKELQSLENXXXMSLKGVXMKKDQILTNEIEELRQKGNLIHQENVELYQK 474
Query: 91 VKR 93
+++
Sbjct: 473 MEQ 465
>TC220943 weakly similar to UP|Q9ZSL4 (Q9ZSL4) MADS box protein (Fragment),
partial (15%)
Length = 638
Score = 36.2 bits (82), Expect = 0.005
Identities = 18/53 (33%), Positives = 34/53 (63%)
Frame = +1
Query: 40 SLDELQQIEQQLEKSVSVVRARKNQAYKHQIDQLKEKEKNLVAENARLSKQVK 92
+L ELQ++E++L+ S++ V K + + +I LKEK K L+ +N + + +K
Sbjct: 10 TLRELQKLEERLDSSLNRVYKAKVENFIKEIGILKEKGKKLMEDNMLIKQMIK 168
>TC219933 weakly similar to UP|Q9EQF1 (Q9EQF1) GABA-A epsilon subunit splice
variant (Fragment), partial (7%)
Length = 490
Score = 34.7 bits (78), Expect = 0.015
Identities = 17/33 (51%), Positives = 20/33 (60%)
Frame = +1
Query: 98 PQPQPQPTTKDHQREDQQPYAESSPSSDVVTEL 130
P+PQPQP TK E QP+ E P S+ V EL
Sbjct: 169 PEPQPQPETKPEPTEQTQPHLE--PKSESVPEL 261
>TC224470 similar to UP|Q9AR13 (Q9AR13) MADS-box transcription factor MADS4,
partial (33%)
Length = 485
Score = 34.3 bits (77), Expect = 0.019
Identities = 22/75 (29%), Positives = 33/75 (43%), Gaps = 20/75 (26%)
Frame = +2
Query: 52 EKSVSVVRARKNQAYKHQIDQLKEKEKNLVAENARLSKQVKR---------------NY- 95
E + +R R+N I +L++KEK + EN L+K++K NY
Sbjct: 5 EAGIKNIRTRRNDLMSESISELQKKEKRIQEENNTLAKKIKEKEQAVAQQAAQWEQPNYR 184
Query: 96 ----CPPQPQPQPTT 106
PQ QP PT+
Sbjct: 185 VDTSFMPQQQPLPTS 229
>CO978855
Length = 826
Score = 32.0 bits (71), Expect = 0.096
Identities = 18/52 (34%), Positives = 29/52 (55%), Gaps = 4/52 (7%)
Frame = -2
Query: 34 EGLGSCSLDELQQIEQQLEKSVSVVRARKNQAYKHQIDQL----KEKEKNLV 81
+ LG C + +QI QLE+ +R R++Q Y ID L K+KE+ ++
Sbjct: 825 DSLGGCEGNTPEQISAQLERLNQTIR-RESQRYSESIDDLRMLYKKKERKII 673
>TC208523 similar to UP|TOB1_MOUSE (Q61471) Tob1 protein (Transducer of
erbB-2 1), partial (5%)
Length = 665
Score = 30.0 bits (66), Expect = 0.37
Identities = 12/27 (44%), Positives = 16/27 (58%)
Frame = +1
Query: 98 PQPQPQPTTKDHQREDQQPYAESSPSS 124
PQPQPQP + Q++ Q +PSS
Sbjct: 106 PQPQPQPQQQQEQQQQQAMLPSQAPSS 186
>TC219444 weakly similar to UP|Q9FXT6 (Q9FXT6) PEX14 protein, partial (23%)
Length = 1053
Score = 30.0 bits (66), Expect = 0.37
Identities = 28/85 (32%), Positives = 36/85 (41%), Gaps = 18/85 (21%)
Frame = +2
Query: 61 RKNQAYKHQIDQLKEKEKNLVAENARLSKQ-----VKRNYCPPQP-------------QP 102
RKN + +ID E E N V A S+Q V+R + PPQP +P
Sbjct: 524 RKNVRIR-EIDN--ENESNGVPNGAASSQQPIQQPVQRVWVPPQPPPIAMPEAAEAIRRP 694
Query: 103 QPTTKDHQREDQQPYAESSPSSDVV 127
+P + Q D Q A S SD V
Sbjct: 695 KPAAQKEQMSDNQSVAHSLDGSDDV 769
>TC220376
Length = 740
Score = 29.6 bits (65), Expect = 0.48
Identities = 18/67 (26%), Positives = 33/67 (48%), Gaps = 5/67 (7%)
Frame = +3
Query: 60 ARKNQAYKHQIDQLKEKEKNLV-----AENARLSKQVKRNYCPPQPQPQPTTKDHQREDQ 114
+R + +K ++D KE E + + A +ARL + ++ + Q Q P K + +DQ
Sbjct: 126 SRAERKFKSELDHFKEVELDALHSSVDAVSARLRRHLQASKANHQQQKTPGKKSYAGDDQ 305
Query: 115 QPYAESS 121
+SS
Sbjct: 306 MSLLKSS 326
>TC204529
Length = 854
Score = 29.6 bits (65), Expect = 0.48
Identities = 16/62 (25%), Positives = 32/62 (50%)
Frame = +1
Query: 43 ELQQIEQQLEKSVSVVRARKNQAYKHQIDQLKEKEKNLVAENARLSKQVKRNYCPPQPQP 102
+L+ Q L++ + ++A KN +L+++++ L AE +L +Q+K P P
Sbjct: 184 KLKDTNQGLQEKIKELKAEKN--------ELRDEKQRLKAEKEKLEQQLKSLNAQPSFMP 339
Query: 103 QP 104
P
Sbjct: 340 PP 345
>TC204528
Length = 793
Score = 29.6 bits (65), Expect = 0.48
Identities = 16/62 (25%), Positives = 32/62 (50%)
Frame = +2
Query: 43 ELQQIEQQLEKSVSVVRARKNQAYKHQIDQLKEKEKNLVAENARLSKQVKRNYCPPQPQP 102
+L+ Q L++ + ++A KN +L+++++ L AE +L +Q+K P P
Sbjct: 524 KLKDTNQGLQEKIKELKAEKN--------ELRDEKQRLKAEKEKLEQQLKSLNAQPSFMP 679
Query: 103 QP 104
P
Sbjct: 680 PP 685
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.310 0.126 0.343
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,966,923
Number of Sequences: 63676
Number of extensions: 83752
Number of successful extensions: 842
Number of sequences better than 10.0: 106
Number of HSP's better than 10.0 without gapping: 760
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 820
length of query: 138
length of database: 12,639,632
effective HSP length: 87
effective length of query: 51
effective length of database: 7,099,820
effective search space: 362090820
effective search space used: 362090820
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 54 (25.4 bits)
Medicago: description of AC147014.15


