

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147013.10 - phase: 0 /pseudo
(339 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
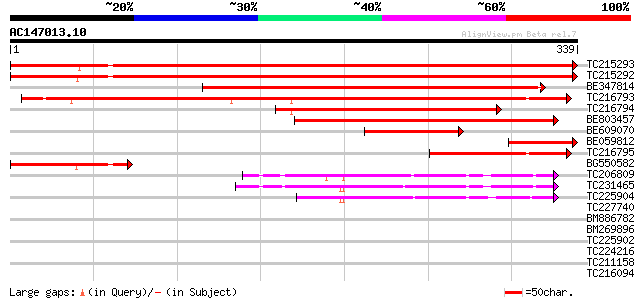
Score E
Sequences producing significant alignments: (bits) Value
TC215293 similar to UP|FENR_TOBAC (O04977) Ferredoxin--NADP redu... 613 e-176
TC215292 similar to UP|FENR_PEA (P10933) Ferredoxin--NADP reduct... 608 e-175
BE347814 similar to SP|P41346|FENR Ferredoxin--NADP reductase c... 300 8e-82
TC216793 similar to UP|FENS_PEA (Q41014) Ferredoxin--NADP reduct... 278 2e-75
TC216794 homologue to UP|FENS_PEA (Q41014) Ferredoxin--NADP redu... 155 2e-38
BE803457 weakly similar to GP|10177134|dbj ferredoxin-NADP+ redu... 138 4e-33
BE609070 similar to SP|P41346|FENR_ Ferredoxin--NADP reductase ... 94 7e-20
BE059812 GP|294092|gb|AA ferredoxin NADP+ reductase {Pisum sativ... 87 1e-17
TC216795 homologue to UP|FENS_PEA (Q41014) Ferredoxin--NADP redu... 79 2e-15
BG550582 similar to SP|O04977|FENR_ Ferredoxin--NADP reductase ... 74 7e-14
TC206809 homologue to UP|NCPR_PHAAU (P37116) NADPH--cytochrome P... 71 6e-13
TC231465 UP|Q8GUS1 (Q8GUS1) NADPH:P450 reductase, partial (77%) 66 2e-11
TC225904 similar to UP|O48937 (O48937) NADPH cytochrome P450 red... 60 1e-09
TC227740 similar to UP|Q8LBD3 (Q8LBD3) Cytochrome-b5 reductase-l... 40 0.002
BM886782 35 0.049
BM269896 similar to SP|P51414|R26A_ 60S ribosomal protein L26A. ... 31 0.70
TC225902 similar to UP|O48937 (O48937) NADPH cytochrome P450 red... 31 0.92
TC224216 similar to GB|AAP68290.1|31711868|BT008851 At3g12156 {A... 29 2.7
TC211158 weakly similar to UP|Q8LG88 (Q8LG88) Sodium-dicarboxyla... 28 4.5
TC216094 similar to UP|Q6SJQ9 (Q6SJQ9) TFIID component TAF2 (Fra... 28 7.7
>TC215293 similar to UP|FENR_TOBAC (O04977) Ferredoxin--NADP reductase,
leaf-type isozyme, chloroplast precursor (FNR) ,
complete
Length = 1488
Score = 613 bits (1580), Expect = e-176
Identities = 302/342 (88%), Positives = 326/342 (95%), Gaps = 3/342 (0%)
Frame = +1
Query: 1 MAAAVTAAVSFPYSNSTSLPIRTSVISPDRLVFKKVSLNN--VSISGRL-TVRAEVATEA 57
MAAAV+AAVSFP + STSLP RTS+I+P+R+VFKK SL N VS SGR+ ++RA+V TEA
Sbjct: 139 MAAAVSAAVSFPSTKSTSLPSRTSLIAPERVVFKKASLQNRDVSSSGRVVSIRAQVTTEA 318
Query: 58 PAPVKVEKISKKQEEGIVVNKFKPKTPYIGRCLLNTKITGDDAPGETWHMVFSTEGEVPY 117
PA KVEK SKKQ+EG+VVNKFKPKTPYIGRCLLNTKITGDDAPGETWHMVFSTEGE+PY
Sbjct: 319 PA--KVEKESKKQDEGVVVNKFKPKTPYIGRCLLNTKITGDDAPGETWHMVFSTEGEIPY 492
Query: 118 REGQSIGIVPDGIDKNGKPHKLRLYSIASSALGDFGDSKTVSLCVKRLVYTNDAGEVVKG 177
REGQSIG++PDGIDKNGKPHKLRLYSIASSA+GDFGDSKTVSLCVKRLVYTN+ GE+VKG
Sbjct: 493 REGQSIGVIPDGIDKNGKPHKLRLYSIASSAIGDFGDSKTVSLCVKRLVYTNENGEIVKG 672
Query: 178 VCSNFLCDLRPGSEVQITGPVGKEMLMPKDPNATVIMLGTGTGIAPFRSFLWKMFFEKHE 237
VCSNFLCDL+PG+EV ITGPVGKEMLMPKDPNAT+IMLGTGTGIAPFRSFLWKMFFEKHE
Sbjct: 673 VCSNFLCDLKPGAEVTITGPVGKEMLMPKDPNATIIMLGTGTGIAPFRSFLWKMFFEKHE 852
Query: 238 DYKFNGLAWLFLGVPTSSSLLYKEEFEKMKEKAPENFRLDFAVSREQVNDKGEKMYIQTR 297
DYKFNGLAWLFLGVPTSSSLLYKEEFEKM+EK+PENFRLDFAVSREQ N+KGEKMYIQTR
Sbjct: 853 DYKFNGLAWLFLGVPTSSSLLYKEEFEKMQEKSPENFRLDFAVSREQTNEKGEKMYIQTR 1032
Query: 298 MAQYAEELWELLKKDNTFVYMCGLKGMEKGIDDIMVSLAAKD 339
MAQYAEELWELLKKDNTFVYMCGLKGMEKGIDDIMVSLAAKD
Sbjct: 1033MAQYAEELWELLKKDNTFVYMCGLKGMEKGIDDIMVSLAAKD 1158
>TC215292 similar to UP|FENR_PEA (P10933) Ferredoxin--NADP reductase, leaf
isozyme, chloroplast precursor (FNR) , complete
Length = 1435
Score = 608 bits (1569), Expect = e-175
Identities = 301/342 (88%), Positives = 323/342 (94%), Gaps = 3/342 (0%)
Frame = +2
Query: 1 MAAAVTAAVSFPYSNSTSLPIRTSVISPDRLVFKKVSLN--NVSISGRL-TVRAEVATEA 57
MA AVTAAVSFP + STSL RTS+ +PDR+VFKKVSL +VS SGR+ ++RA+V TEA
Sbjct: 131 MATAVTAAVSFPATKSTSLSSRTSITAPDRIVFKKVSLQYRDVSTSGRVVSIRAQVTTEA 310
Query: 58 PAPVKVEKISKKQEEGIVVNKFKPKTPYIGRCLLNTKITGDDAPGETWHMVFSTEGEVPY 117
PA KVEK SKKQEEG++VNKFKPK PYIGRCLLNTKITGDDAPGETWHMVFSTEGEVPY
Sbjct: 311 PA--KVEKESKKQEEGVIVNKFKPKNPYIGRCLLNTKITGDDAPGETWHMVFSTEGEVPY 484
Query: 118 REGQSIGIVPDGIDKNGKPHKLRLYSIASSALGDFGDSKTVSLCVKRLVYTNDAGEVVKG 177
REGQSIG++PDG+DKNGKPHKLRLYSIASSALGDFGDSKTVSLCVKRLVYTN+ GE+VKG
Sbjct: 485 REGQSIGVIPDGVDKNGKPHKLRLYSIASSALGDFGDSKTVSLCVKRLVYTNENGELVKG 664
Query: 178 VCSNFLCDLRPGSEVQITGPVGKEMLMPKDPNATVIMLGTGTGIAPFRSFLWKMFFEKHE 237
VCSNFLCDL+PG+EV+ITGPVGKEMLMPKDPNATVIML TGTGIAPFRSFLWKMFFEKH+
Sbjct: 665 VCSNFLCDLKPGAEVKITGPVGKEMLMPKDPNATVIMLATGTGIAPFRSFLWKMFFEKHD 844
Query: 238 DYKFNGLAWLFLGVPTSSSLLYKEEFEKMKEKAPENFRLDFAVSREQVNDKGEKMYIQTR 297
DYKFNGLAWLFLGVPTSSSLLYKEEFEKMKEKAP+NFRLDFAVSREQ N+KGEKMYIQTR
Sbjct: 845 DYKFNGLAWLFLGVPTSSSLLYKEEFEKMKEKAPDNFRLDFAVSREQTNEKGEKMYIQTR 1024
Query: 298 MAQYAEELWELLKKDNTFVYMCGLKGMEKGIDDIMVSLAAKD 339
MAQYAEELWELLKKDNTFVYMCGLKGMEKGIDDIMVSLAAKD
Sbjct: 1025MAQYAEELWELLKKDNTFVYMCGLKGMEKGIDDIMVSLAAKD 1150
>BE347814 similar to SP|P41346|FENR Ferredoxin--NADP reductase chloroplast
precursor (EC 1.18.1.2) (FNR). [Broad bean] {Vicia
faba}, partial (51%)
Length = 613
Score = 300 bits (767), Expect = 8e-82
Identities = 150/205 (73%), Positives = 172/205 (83%)
Frame = +2
Query: 116 PYREGQSIGIVPDGIDKNGKPHKLRLYSIASSALGDFGDSKTVSLCVKRLVYTNDAGEVV 175
P RE SIG++PDG+DKNG+PH LRLYSIASSA+GDFGDSKTVSLCVKRL +TN+ GE+V
Sbjct: 2 P*RE*PSIGVIPDGMDKNGEPH*LRLYSIASSAIGDFGDSKTVSLCVKRLGFTNENGEMV 181
Query: 176 KGVCSNFLCDLRPGSEVQITGPVGKEMLMPKDPNATVIMLGTGTGIAPFRSFLWKMFFEK 235
GVCSNFLCDL+P +EV IT PVGKEMLMPKDPNAT+IMLGTGTGIAP+RSFL KMFF K
Sbjct: 182 YGVCSNFLCDLKPAAEVTITVPVGKEMLMPKDPNATIIMLGTGTGIAPYRSFLCKMFFAK 361
Query: 236 HEDYKFNGLAWLFLGVPTSSSLLYKEEFEKMKEKAPENFRLDFAVSREQVNDKGEKMYIQ 295
H+DYKFNG AWLFLGVPTSSSLLYKEE EKM+EK+P+N LD+AV R N+ G+ MYIQ
Sbjct: 362 HQDYKFNGEAWLFLGVPTSSSLLYKEECEKMQEKSPDNVMLDYAVWR*HTNNIGDGMYIQ 541
Query: 296 TRMAQYAEELWELLKKDNTFVYMCG 320
+R+AQYA L + NT V CG
Sbjct: 542 SRVAQYAR*ALGLTDRGNTIV-ACG 613
>TC216793 similar to UP|FENS_PEA (Q41014) Ferredoxin--NADP reductase, root
isozyme, chloroplast precursor (FNR) , partial (97%)
Length = 1408
Score = 278 bits (712), Expect = 2e-75
Identities = 156/345 (45%), Positives = 208/345 (60%), Gaps = 16/345 (4%)
Frame = +3
Query: 8 AVSFPYSNSTSLPIRTSVISPDRLVFKK---------VSLNNVSISGRLTVRAEVATEAP 58
AV+ P N SL R++V +P+ + K + NN S+ R V V +
Sbjct: 234 AVTVPVGNDLSLR-RSAVKAPNLNFWDKSWAPVFTLDLKPNNPSLRSRHVVCMSVQQASV 410
Query: 59 APVKVEKISKKQEEGIVVNKFKPKTPYIGRCLLNTKITGDDAPGETWHMVFSTEGEVPYR 118
+ V V + + + +N +KPK PY + ++ G APGET H+V G VPY
Sbjct: 411 SKVNVSPLELEDAKEPPLNLYKPKEPYTATIVSVDRLVGPKAPGETCHIVIDHGGNVPYW 590
Query: 119 EGQSIGIVPDGID--KNGKPHKLRLYSIASSALGDFGDSKTVSLCVKRLVY----TNDAG 172
EGQS G++P G + K G PH +RLYSIAS+ GDF D KT SLCV+R VY T
Sbjct: 591 EGQSYGVIPPGENPKKPGAPHNVRLYSIASTRYGDFFDGKTASLCVRRAVYYDPETGKED 770
Query: 173 EVVKGVCSNFLCDLRPGSEVQITGPVGKEMLMPKD-PNATVIMLGTGTGIAPFRSFLWKM 231
G+CSNFLC+ +PG ++QITGP GK ML+P+D PNAT IM+ TGTG+APFR +L +M
Sbjct: 771 PSKNGICSNFLCNSKPGDKIQITGPSGKIMLLPEDDPNATHIMIATGTGVAPFRGYLRRM 950
Query: 232 FFEKHEDYKFNGLAWLFLGVPTSSSLLYKEEFEKMKEKAPENFRLDFAVSREQVNDKGEK 291
F E YKF GLAWLFLGV + SLLY EEF K +NFR D A+SREQ N G K
Sbjct: 951 FMESVPTYKFGGLAWLFLGVANTDSLLYDEEFSKYLNDYSDNFRYDRALSREQKNKNGGK 1130
Query: 292 MYIQTRMAQYAEELWELLKKDNTFVYMCGLKGMEKGIDDIMVSLA 336
MY+Q ++ +Y++E+++LL + +Y CGLKGM GI D + +A
Sbjct: 1131MYVQDKIEEYSDEIFKLL-DNGAHIYFCGLKGMMPGIQDTLKKVA 1262
>TC216794 homologue to UP|FENS_PEA (Q41014) Ferredoxin--NADP reductase, root
isozyme, chloroplast precursor (FNR) , partial (37%)
Length = 422
Score = 155 bits (392), Expect = 2e-38
Identities = 78/140 (55%), Positives = 97/140 (68%), Gaps = 5/140 (3%)
Frame = +2
Query: 160 LCVKRLVY----TNDAGEVVKGVCSNFLCDLRPGSEVQITGPVGKEMLMPKD-PNATVIM 214
LCV+R VY T G+CSNFLC+ +PG ++QITGP GK ML+P+D PNAT IM
Sbjct: 2 LCVRRAVYYDPETGKEDPSKNGICSNFLCNSKPGDKIQITGPSGKIMLLPEDDPNATHIM 181
Query: 215 LGTGTGIAPFRSFLWKMFFEKHEDYKFNGLAWLFLGVPTSSSLLYKEEFEKMKEKAPENF 274
+ TGTG+APFR +L +MF E YKF GLAWLFLGV + SLLY +EF K + P+NF
Sbjct: 182 IATGTGVAPFRGYLRRMFMESVPAYKFGGLAWLFLGVANTDSLLYDDEFSKYLKDYPDNF 361
Query: 275 RLDFAVSREQVNDKGEKMYI 294
R + A+SREQ N G KMY+
Sbjct: 362 RYNRALSREQKNKSGGKMYV 421
>BE803457 weakly similar to GP|10177134|dbj ferredoxin-NADP+ reductase
{Arabidopsis thaliana}, partial (15%)
Length = 478
Score = 138 bits (347), Expect = 4e-33
Identities = 73/158 (46%), Positives = 102/158 (64%)
Frame = +2
Query: 171 AGEVVKGVCSNFLCDLRPGSEVQITGPVGKEMLMPKDPNATVIMLGTGTGIAPFRSFLWK 230
+G++ K V CDL+P + V TGP KE+LM +P+AT+ + T +AP R+ LW
Sbjct: 5 SGQIGKVVS*YSDCDLKP*AIVTHTGPRVKELLMR*NPSATIYVTRTTAALAPVRALLWN 184
Query: 231 MFFEKHEDYKFNGLAWLFLGVPTSSSLLYKEEFEKMKEKAPENFRLDFAVSREQVNDKGE 290
++KHE +F+G W++L P SSLL KE+ +KM + +NF+LD A RE N+ GE
Sbjct: 185 EIYKKHEKNEFDGK*WIYLRDPACSSLLSKEDMDKMHKTYADNFKLDSARCRELTNEHGE 364
Query: 291 KMYIQTRMAQYAEELWELLKKDNTFVYMCGLKGMEKGI 328
KMYI TR AQYA +L ++LKK+ T +MCGL GM GI
Sbjct: 365 KMYIPTREAQYA*QLCDILKKEYTTAHMCGLLGMHVGI 478
>BE609070 similar to SP|P41346|FENR_ Ferredoxin--NADP reductase chloroplast
precursor (EC 1.18.1.2) (FNR). [Broad bean] {Vicia
faba}, partial (14%)
Length = 178
Score = 94.4 bits (233), Expect = 7e-20
Identities = 42/59 (71%), Positives = 51/59 (86%)
Frame = +2
Query: 213 IMLGTGTGIAPFRSFLWKMFFEKHEDYKFNGLAWLFLGVPTSSSLLYKEEFEKMKEKAP 271
IMLG + I P RS +WKMFFEKH++YKFNG+AWLFLGVPTS SLLYKEEF+K++ K+P
Sbjct: 2 IMLGAFSWIDPLRSLIWKMFFEKHKNYKFNGVAWLFLGVPTSRSLLYKEEFDKIQYKSP 178
>BE059812 GP|294092|gb|AA ferredoxin NADP+ reductase {Pisum sativum}, partial
(46%)
Length = 258
Score = 87.0 bits (214), Expect = 1e-17
Identities = 41/41 (100%), Positives = 41/41 (100%)
Frame = +3
Query: 299 AQYAEELWELLKKDNTFVYMCGLKGMEKGIDDIMVSLAAKD 339
AQYAEELWELLKKDNTFVYMCGLKGMEKGIDDIMVSLAAKD
Sbjct: 6 AQYAEELWELLKKDNTFVYMCGLKGMEKGIDDIMVSLAAKD 128
>TC216795 homologue to UP|FENS_PEA (Q41014) Ferredoxin--NADP reductase, root
isozyme, chloroplast precursor (FNR) , partial (34%)
Length = 764
Score = 79.3 bits (194), Expect = 2e-15
Identities = 38/85 (44%), Positives = 57/85 (66%)
Frame = +1
Query: 252 PTSSSLLYKEEFEKMKEKAPENFRLDFAVSREQVNDKGEKMYIQTRMAQYAEELWELLKK 311
P+ SLLY +EF K + P+NFR + A+SREQ N G KMY+Q ++ +Y++E+++LL
Sbjct: 91 PSLRSLLYDDEFSKYLKDYPDNFRYNRALSREQKNKSGGKMYVQDKIEEYSDEIFKLL-D 267
Query: 312 DNTFVYMCGLKGMEKGIDDIMVSLA 336
+ +Y CGLKGM GI D + +A
Sbjct: 268 NGAHIYFCGLKGMMPGIQDTLKKVA 342
>BG550582 similar to SP|O04977|FENR_ Ferredoxin--NADP reductase leaf-type
isozyme chloroplast precursor (EC 1.18.1.2) (FNR).,
partial (14%)
Length = 251
Score = 74.3 bits (181), Expect = 7e-14
Identities = 48/76 (63%), Positives = 56/76 (73%), Gaps = 3/76 (3%)
Frame = +2
Query: 1 MAAAVTAAVSFPYSNSTSLPIRTSVISPDRLVFKKVSL--NNVSISGR-LTVRAEVATEA 57
MAAAV+AAVSFP + STSLP RT I+PDR+V KK S +V SGR + RA+V TEA
Sbjct: 26 MAAAVSAAVSFPSTKSTSLPSRTFFITPDRVVGKKASFFYRDVCSSGRKVGRRAQVTTEA 205
Query: 58 PAPVKVEKISKKQEEG 73
PA KVEK SKKQ+EG
Sbjct: 206PA--KVEKESKKQDEG 247
>TC206809 homologue to UP|NCPR_PHAAU (P37116) NADPH--cytochrome P450
reductase (CPR) (P450R) , partial (70%)
Length = 1867
Score = 71.2 bits (173), Expect = 6e-13
Identities = 65/197 (32%), Positives = 94/197 (46%), Gaps = 8/197 (4%)
Frame = +2
Query: 140 RLYSIASSALGDFGDSKTVSLCVKRLVY-TNDAGEVVKGVCSNFLCDLRP--GSEVQITG 196
R YSI+SS F + C LVY G + KGVCS ++ + P S
Sbjct: 788 RYYSISSSPR--FAPQRVHVTCA--LVYGPTPTGRIHKGVCSTWMKNAIPLEKSPDCSWA 955
Query: 197 PV---GKEMLMPKDPNATVIMLGTGTGIAPFRSFLWKMFFEKHEDYKFNGLAWLFLGVPT 253
P+ +P D + +IM+G GTG+APFR FL + F K + G A LF G
Sbjct: 956 PIFIRPSNFKLPVDHSIPIIMVGPGTGLAPFRGFLQERFALKEAGVQ-QGPAILFFGCRN 1132
Query: 254 SS-SLLYKEEFEKMKEKAPENFRLDFAVSREQVNDKGEKMYIQTRMAQYAEELWELLKKD 312
+Y+EE + E+ + L A SRE EK Y+Q +M A +LW L+ +
Sbjct: 1133RRLDFIYEEELKNFVEQGSLS-ELIVAFSRE----GAEKEYVQHKMMDQAAQLWSLISQG 1297
Query: 313 NTFVYMCG-LKGMEKGI 328
++Y+CG KGM + +
Sbjct: 1298G-YLYVCGDAKGMARDV 1345
>TC231465 UP|Q8GUS1 (Q8GUS1) NADPH:P450 reductase, partial (77%)
Length = 1933
Score = 66.2 bits (160), Expect = 2e-11
Identities = 63/201 (31%), Positives = 94/201 (46%), Gaps = 8/201 (3%)
Frame = +1
Query: 136 PH-KLRLYSIASSALGDFGDSKTVSLCVKRLVYTNDAGEVVKGVCSNFLCDLRPGSEVQI 194
PH + R YSI+SS F K C + G + KGVCS ++ + P + +
Sbjct: 910 PHLQPRYYSISSSPR--FSPQKVHVTCAL-VCGPTPTGRIHKGVCSTWMKNAIPLEKSRD 1080
Query: 195 TG--PV---GKEMLMPKDPNATVIMLGTGTGIAPFRSFLWKMFFEKHEDYKFNGLAWLFL 249
P+ +P D + +IM+G GTG+APFR FL + K ED G A LF
Sbjct: 1081 CSWAPIFIRTSNFKLPADHSIPIIMVGPGTGLAPFRGFLQERLALK-EDAVQLGPALLFF 1257
Query: 250 GVPT-SSSLLYKEEFEKMKEKAPENFRLDFAVSREQVNDKGEKMYIQTRMAQYAEELWEL 308
G +Y++E + E+ + L SRE EK Y+Q +M A LW L
Sbjct: 1258 GCRNRQMDFIYEDELKNFMEQGALS-ELIVTFSRE----GPEKEYVQHKMMDKAANLWNL 1422
Query: 309 LKKDNTFVYMCG-LKGMEKGI 328
+ + ++Y+CG KGM + +
Sbjct: 1423 ISQGG-YLYVCGDAKGMARDV 1482
>TC225904 similar to UP|O48937 (O48937) NADPH cytochrome P450 reductase ,
partial (68%)
Length = 1704
Score = 60.5 bits (145), Expect = 1e-09
Identities = 48/164 (29%), Positives = 77/164 (46%), Gaps = 7/164 (4%)
Frame = +1
Query: 172 GEVVKGVCSNFLCDLRPGSEVQITG--PV---GKEMLMPKDPNATVIMLGTGTGIAPFRS 226
G + KGVCS ++ + P + Q P+ +P D +IM+G GTG+APFR
Sbjct: 841 GRIHKGVCSTWMKNSVPLEKSQDCSWAPIFVRTSNFRLPSDNKVPIIMIGPGTGLAPFRG 1020
Query: 227 FLWKMFFEKHEDYKFNGLAWLFLGVPT-SSSLLYKEEFEKMKEKAPENFRLDFAVSREQV 285
FL + K + G + LF G +Y++E + L A SRE
Sbjct: 1021FLQERLALKEGGAEL-GPSVLFFGCRNRQMDYIYEDELSHFVNTGALD-ELILAFSREGP 1194
Query: 286 NDKGEKMYIQTRMAQYAEELWELLKKDNTFVYMCG-LKGMEKGI 328
K Y+Q +M + A E+W ++ + ++Y+CG KGM + +
Sbjct: 1195T----KEYVQHKMMEKASEIWSMISQ-GAYIYVCGDAKGMARDV 1311
>TC227740 similar to UP|Q8LBD3 (Q8LBD3) Cytochrome-b5 reductase-like protein,
partial (78%)
Length = 1368
Score = 40.0 bits (92), Expect = 0.002
Identities = 37/153 (24%), Positives = 66/153 (42%)
Frame = +1
Query: 176 KGVCSNFLCDLRPGSEVQITGPVGKEMLMPKDPNATVIMLGTGTGIAPFRSFLWKMFFEK 235
+G S L+PG V++ GP+ K P + + M+ GTGI P + + +
Sbjct: 529 EGKMSQHFASLKPGDVVEVKGPIEKLRYTP-NMKKHIGMIAGGTGITPMLQVIEAIL--R 699
Query: 236 HEDYKFNGLAWLFLGVPTSSSLLYKEEFEKMKEKAPENFRLDFAVSREQVNDKGEKMYIQ 295
+ D K ++ L+ V + +L K++ + + P N ++ + V N +G YI
Sbjct: 700 NPDDK-TQISLLYANV-SPDDILLKQKLDILATSHP-NLKIFYTVDNPTKNWRGGAGYIS 870
Query: 296 TRMAQYAEELWELLKKDNTFVYMCGLKGMEKGI 328
+ D+T + +CG GM K I
Sbjct: 871 KDVVVKGLP----SPSDDTLILVCGPPGMMKAI 957
>BM886782
Length = 385
Score = 35.0 bits (79), Expect = 0.049
Identities = 17/39 (43%), Positives = 24/39 (60%), Gaps = 4/39 (10%)
Frame = -1
Query: 162 VKRLVY----TNDAGEVVKGVCSNFLCDLRPGSEVQITG 196
V+R VY T G+CSNFLC+ +P +++QITG
Sbjct: 385 VRRAVYYDPVTGKEDPSKNGICSNFLCNSKP*NKIQITG 269
>BM269896 similar to SP|P51414|R26A_ 60S ribosomal protein L26A. [Mouse-ear
cress] {Arabidopsis thaliana}, partial (19%)
Length = 421
Score = 31.2 bits (69), Expect = 0.70
Identities = 14/39 (35%), Positives = 23/39 (58%)
Frame = -3
Query: 133 NGKPHKLRLYSIASSALGDFGDSKTVSLCVKRLVYTNDA 171
+G+P RL+ + D+GD +T LC++ +YTN A
Sbjct: 320 HGEPLVTRLHRRGHAPREDYGDQRTRHLCLRLRLYTNHA 204
>TC225902 similar to UP|O48937 (O48937) NADPH cytochrome P450 reductase ,
partial (11%)
Length = 581
Score = 30.8 bits (68), Expect = 0.92
Identities = 12/37 (32%), Positives = 23/37 (61%), Gaps = 1/37 (2%)
Frame = +2
Query: 293 YIQTRMAQYAEELWELLKKDNTFVYMCG-LKGMEKGI 328
Y+Q +M + A E+W ++ ++Y+CG KGM + +
Sbjct: 2 YVQHKMMEKASEIWSMI-SXGAYIYVCGDAKGMARNV 109
>TC224216 similar to GB|AAP68290.1|31711868|BT008851 At3g12156 {Arabidopsis
thaliana;} , partial (64%)
Length = 889
Score = 29.3 bits (64), Expect = 2.7
Identities = 18/62 (29%), Positives = 31/62 (49%)
Frame = +3
Query: 204 MPKDPNATVIMLGTGTGIAPFRSFLWKMFFEKHEDYKFNGLAWLFLGVPTSSSLLYKEEF 263
+PK+PNA + + T G P S L E + + + + W+ G SS +L+ +EF
Sbjct: 489 IPKNPNAVIFVAATDDGYIPKHSVL-----ELQKAWPGSEVRWV-TGGHVSSFILHNDEF 650
Query: 264 EK 265
+
Sbjct: 651 RR 656
>TC211158 weakly similar to UP|Q8LG88 (Q8LG88) Sodium-dicarboxylate
cotransporter-like, partial (32%)
Length = 656
Score = 28.5 bits (62), Expect = 4.5
Identities = 14/42 (33%), Positives = 20/42 (47%)
Frame = +2
Query: 217 TGTGIAPFRSFLWKMFFEKHEDYKFNGLAWLFLGVPTSSSLL 258
TGTG+ +WK F + + FN W F G P + +L
Sbjct: 362 TGTGVNLIIIGMWKSLFPEAKPISFN--TWFFYGFPVAVLIL 481
>TC216094 similar to UP|Q6SJQ9 (Q6SJQ9) TFIID component TAF2 (Fragment),
partial (20%)
Length = 2400
Score = 27.7 bits (60), Expect = 7.7
Identities = 17/56 (30%), Positives = 23/56 (40%), Gaps = 3/56 (5%)
Frame = -2
Query: 197 PVGKEMLMPKDPNATVIMLGTGTGIAPFRSFLWKM---FFEKHEDYKFNGLAWLFL 249
P G L D N ++ + G I+ R+F+W F K Y F L FL
Sbjct: 1274 PTGGRRLFDLDFNGSLSFMAIGYSISLLRNFIWSFNWCFLGKFSKYLFGLLKGRFL 1107
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.317 0.135 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,753,794
Number of Sequences: 63676
Number of extensions: 176987
Number of successful extensions: 820
Number of sequences better than 10.0: 41
Number of HSP's better than 10.0 without gapping: 807
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 808
length of query: 339
length of database: 12,639,632
effective HSP length: 98
effective length of query: 241
effective length of database: 6,399,384
effective search space: 1542251544
effective search space used: 1542251544
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 59 (27.3 bits)
Medicago: description of AC147013.10


