

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147011.10 - phase: 0
(165 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
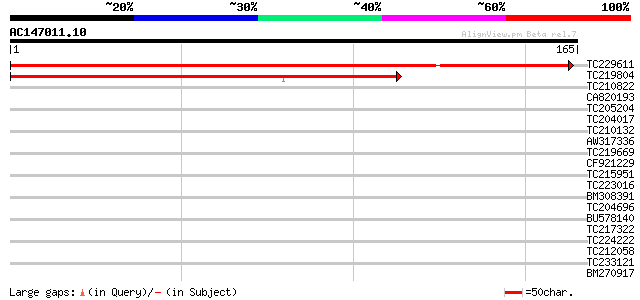
Sequences producing significant alignments: (bits) Value
TC229611 similar to UP|Q9LZ98 (Q9LZ98) ABC transporter-like prot... 286 4e-78
TC219804 similar to UP|Q8LBC2 (Q8LBC2) NBD-like protein, partial... 104 2e-23
TC210822 similar to GB|AAQ89671.1|37202112|BT010649 At4g33460 {A... 33 0.081
CA820193 similar to PIR|T04229|T04 ABC-type transport protein F1... 32 0.11
TC205204 homologue to UP|Q43558 (Q43558) Proline rich protein pr... 32 0.14
TC204017 similar to UP|Q8LGU1 (Q8LGU1) Multidrug-resistance rela... 30 0.40
TC210132 similar to UP|O24510 (O24510) MRP-like ABC transporter,... 30 0.52
AW317336 similar to GP|4586576|dbj| multidrug resistance protein... 29 0.89
TC219669 similar to UP|Q9LU34 (Q9LU34) Multidrug resistance-asso... 29 0.89
CF921229 29 1.2
TC215951 similar to UP|Q9SXT3 (Q9SXT3) Multidrug resistance prot... 28 2.0
TC223016 similar to GB|AAN31116.1|23506209|AY149962 At3g06130/F2... 28 2.6
BM308391 27 3.4
TC204696 similar to UP|Q9LWB4 (Q9LWB4) L1 protein, partial (72%) 27 3.4
BU578140 27 3.4
TC217322 27 3.4
TC224222 similar to UP|O82615 (O82615) T9A4.6 protein (Probable ... 27 3.4
TC212058 27 3.4
TC233121 similar to GB|AAP13385.1|30023704|BT006277 At3g56360 {A... 27 4.4
BM270917 similar to GP|4586576|dbj multidrug resistance protein ... 27 5.8
>TC229611 similar to UP|Q9LZ98 (Q9LZ98) ABC transporter-like protein
(At5g02270), partial (58%)
Length = 857
Score = 286 bits (731), Expect = 4e-78
Identities = 144/164 (87%), Positives = 155/164 (93%)
Frame = +3
Query: 1 MGLLKPFQVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIV 60
MGLLKPF+VLLLDEITVDLDVLARADLL+FLRKECDERGATIIYATHIFDGLEDWPTNIV
Sbjct: 99 MGLLKPFKVLLLDEITVDLDVLARADLLRFLRKECDERGATIIYATHIFDGLEDWPTNIV 278
Query: 61 YVAHGRLELAMPIEKVKETSKLSLMRTVEVWLRKERDEDRRQRKERKAAGLPEFDKQVDG 120
YVAHG+L+LAMP++KVKE SKLSLMRTVE WLRKERDEDR++RKERKAAGLPEF K+V+
Sbjct: 279 YVAHGKLQLAMPMDKVKEISKLSLMRTVESWLRKERDEDRKKRKERKAAGLPEFGKRVEE 458
Query: 121 SRVVGGDPARAPVRVTNNGWAAGRLHSTIAGEENFLLSSNRVLR 164
SRV GDPARA VRV NNGWAAGRL ST+AGEENFLLSSNRVLR
Sbjct: 459 SRVT-GDPARAAVRVINNGWAAGRLTSTVAGEENFLLSSNRVLR 587
>TC219804 similar to UP|Q8LBC2 (Q8LBC2) NBD-like protein, partial (48%)
Length = 711
Score = 104 bits (259), Expect = 2e-23
Identities = 51/115 (44%), Positives = 72/115 (62%), Gaps = 1/115 (0%)
Frame = +2
Query: 1 MGLLKPFQVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIV 60
+GL P++VLLLDE+TVDLDV+ + DLL F ++C++R A YATHIFDGLE W T++
Sbjct: 92 LGLFHPYKVLLLDEVTVDLDVVTQMDLLDFFMEDCEQREAIXXYATHIFDGLETWATHLA 271
Query: 61 YVAHGRLELAMPIEKVKE-TSKLSLMRTVEVWLRKERDEDRRQRKERKAAGLPEF 114
Y+ G L A I VKE S +L+ VE WLR E +++ ++ + F
Sbjct: 272 YIQDGELRRAEKISNVKELKSSTNLLSVVEAWLRAETKLEKKNPVQKTSVASSPF 436
>TC210822 similar to GB|AAQ89671.1|37202112|BT010649 At4g33460 {Arabidopsis
thaliana;} , partial (34%)
Length = 527
Score = 32.7 bits (73), Expect = 0.081
Identities = 20/66 (30%), Positives = 35/66 (52%), Gaps = 1/66 (1%)
Frame = +1
Query: 3 LLKPFQVLLLDEITVDLDVLARADLLKFLRKECDERG-ATIIYATHIFDGLEDWPTNIVY 61
L + +VLLLDE+T LD + ++K +R D T ++ TH + LE + +Y
Sbjct: 28 LAEACKVLLLDELTTFLDEADQVGVIKAVRNSVDTSAEVTALWVTHRLEELE-YADGAIY 204
Query: 62 VAHGRL 67
+ G++
Sbjct: 205MEDGKV 222
>CA820193 similar to PIR|T04229|T04 ABC-type transport protein F14M19.30 -
Arabidopsis thaliana, partial (23%)
Length = 423
Score = 32.3 bits (72), Expect = 0.11
Identities = 18/45 (40%), Positives = 25/45 (55%)
Frame = +1
Query: 3 LLKPFQVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATH 47
LL VLLLDE T LD + +++ L++ C R TII + H
Sbjct: 118 LLHDPAVLLLDEPTSGLDSTSAFKVMRILKQTCVSRNRTIILSIH 252
>TC205204 homologue to UP|Q43558 (Q43558) Proline rich protein precursor,
partial (44%)
Length = 996
Score = 32.0 bits (71), Expect = 0.14
Identities = 17/55 (30%), Positives = 30/55 (53%)
Frame = -3
Query: 70 AMPIEKVKETSKLSLMRTVEVWLRKERDEDRRQRKERKAAGLPEFDKQVDGSRVV 124
A+ +E++ K + E WL +E E+ RQ +E + +PE ++Q + RVV
Sbjct: 520 ALLVEELLVVVKREEEKQEEEWLGEEMPEEGRQGEEMRVVEMPEEERQGEEMRVV 356
>TC204017 similar to UP|Q8LGU1 (Q8LGU1) Multidrug-resistance related protein,
partial (31%)
Length = 1388
Score = 30.4 bits (67), Expect = 0.40
Identities = 24/89 (26%), Positives = 42/89 (46%)
Frame = +3
Query: 3 LLKPFQVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIVYV 62
LLK ++L+LDE T +D A L + +R+E E T+I H + D ++ +
Sbjct: 900 LLKRNRILVLDEATASIDSATDAILQQIIRQEFVE--CTVITVAHRVPTVID-SDMVMVL 1070
Query: 63 AHGRLELAMPIEKVKETSKLSLMRTVEVW 91
++G+L ++ ET+ E W
Sbjct: 1071SYGKLVEYEEPSRLMETNSSFSKLVAEYW 1157
>TC210132 similar to UP|O24510 (O24510) MRP-like ABC transporter, partial
(13%)
Length = 745
Score = 30.0 bits (66), Expect = 0.52
Identities = 22/92 (23%), Positives = 45/92 (48%)
Frame = +2
Query: 3 LLKPFQVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIVYV 62
LLK ++L+LDE T +D + + ++++ E T+I H + D ++++
Sbjct: 323 LLKKSKILVLDEATASVDTATDNTIQQTVKQKFSE--CTVITIAHRITSILD-SDMVLFL 493
Query: 63 AHGRLELAMPIEKVKETSKLSLMRTVEVWLRK 94
G +E +K+ + SL + VE + R+
Sbjct: 494 NQGLIEEYDSPKKLLKNKSSSLAQLVEEYTRR 589
>AW317336 similar to GP|4586576|dbj| multidrug resistance protein {Cicer
arietinum}, partial (37%)
Length = 298
Score = 29.3 bits (64), Expect = 0.89
Identities = 17/53 (32%), Positives = 29/53 (54%)
Frame = -1
Query: 8 QVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIV 60
QVLLLDE T LD ++ ++ + K +G T++ +H + + W +IV
Sbjct: 223 QVLLLDEPTSALDPISTENIEDAMVKLSKHQGMTVLMVSHSIEHIH-WIAHIV 68
>TC219669 similar to UP|Q9LU34 (Q9LU34) Multidrug resistance-associated
protein (MRP)-like; ABC-transporter-like protein,
partial (14%)
Length = 827
Score = 29.3 bits (64), Expect = 0.89
Identities = 24/95 (25%), Positives = 43/95 (45%)
Frame = +1
Query: 3 LLKPFQVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIVYV 62
LLK ++L+LDE T +D L + +R+E E T+I H + D ++ +
Sbjct: 289 LLKRNRILVLDEATASIDSATDVILQQVIRQEFSE--CTVITVAHRVPTVID-SDMVMVL 459
Query: 63 AHGRLELAMPIEKVKETSKLSLMRTVEVWLRKERD 97
++G++ K+ T+ M E W R+
Sbjct: 460 SYGKVVEYDKPSKLMGTNSSFSMLVAEYWSNCNRN 564
>CF921229
Length = 680
Score = 28.9 bits (63), Expect = 1.2
Identities = 18/54 (33%), Positives = 30/54 (55%)
Frame = -3
Query: 60 VYVAHGRLELAMPIEKVKETSKLSLMRTVEVWLRKERDEDRRQRKERKAAGLPE 113
V V G+ E+ + IE+ KET + VEV +R+E+ E + +E+ +PE
Sbjct: 450 VEVTEGKKEVEV-IEEKKETEVTEEKKEVEVEVREEKKESEVKEEEKGQEVVPE 292
>TC215951 similar to UP|Q9SXT3 (Q9SXT3) Multidrug resistance protein
(Fragment), partial (84%)
Length = 1226
Score = 28.1 bits (61), Expect = 2.0
Identities = 16/40 (40%), Positives = 23/40 (57%)
Frame = +2
Query: 8 QVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATH 47
QVLLLDE T LD ++ ++ L K +G T+I +H
Sbjct: 686 QVLLLDEPTSALDPISTENIEDALVKLNKNQGMTVIMVSH 805
>TC223016 similar to GB|AAN31116.1|23506209|AY149962 At3g06130/F28L1_7
{Arabidopsis thaliana;} , partial (6%)
Length = 601
Score = 27.7 bits (60), Expect = 2.6
Identities = 16/52 (30%), Positives = 26/52 (49%)
Frame = -3
Query: 71 MPIEKVKETSKLSLMRTVEVWLRKERDEDRRQRKERKAAGLPEFDKQVDGSR 122
+P+ K +LSL RT + +ERD+ +R RK G + ++ SR
Sbjct: 158 VPLLKYPYCFRLSLTRTTKSHF*RERDKSEERR*LRKMQGNKQLGTRIASSR 3
>BM308391
Length = 428
Score = 27.3 bits (59), Expect = 3.4
Identities = 19/47 (40%), Positives = 21/47 (44%)
Frame = -2
Query: 91 WLRKERDEDRRQRKERKAAGLPEFDKQVDGSRVVGGDPARAPVRVTN 137
W R+ E RR +ER GL G RVV G R RVTN
Sbjct: 169 WERERAREWRRVEEERMVGGLGF------GERVVSGSGRRRS*RVTN 47
>TC204696 similar to UP|Q9LWB4 (Q9LWB4) L1 protein, partial (72%)
Length = 1530
Score = 27.3 bits (59), Expect = 3.4
Identities = 15/46 (32%), Positives = 21/46 (45%)
Frame = +3
Query: 68 ELAMPIEKVKETSKLSLMRTVEVWLRKERDEDRRQRKERKAAGLPE 113
+L I VKET+K + TVE R D ++ R LP+
Sbjct: 453 DLKTAISLVKETAKTKFVETVEAHFRLNIDPKYNDQQLRATVNLPK 590
>BU578140
Length = 434
Score = 27.3 bits (59), Expect = 3.4
Identities = 15/63 (23%), Positives = 28/63 (43%)
Frame = +1
Query: 56 PTNIVYVAHGRLELAMPIEKVKETSKLSLMRTVEVWLRKERDEDRRQRKERKAAGLPEFD 115
P ++VYV G + +P++ ++ L +W +ER++ K +G E
Sbjct: 94 PNSVVYVCLGSICNLIPLQLIELGLALEASEKPFIWFFRERNQTEELNKWINESGFEERT 273
Query: 116 KQV 118
K V
Sbjct: 274 KGV 282
>TC217322
Length = 1077
Score = 27.3 bits (59), Expect = 3.4
Identities = 13/35 (37%), Positives = 20/35 (57%)
Frame = +2
Query: 68 ELAMPIEKVKETSKLSLMRTVEVWLRKERDEDRRQ 102
EL I K +E S LSL R ++WL + + +R+
Sbjct: 842 ELLRGISKTRERSMLSLQRIKDLWLSTKLNYQKRK 946
>TC224222 similar to UP|O82615 (O82615) T9A4.6 protein (Probable
wound-induced protein), partial (40%)
Length = 466
Score = 27.3 bits (59), Expect = 3.4
Identities = 15/35 (42%), Positives = 21/35 (59%), Gaps = 9/35 (25%)
Frame = +3
Query: 118 VDGSRV---------VGGDPARAPVRVTNNGWAAG 143
V+GSRV VGG ++ P+RV+N+G AG
Sbjct: 138 VEGSRVLQNEQHNDEVGGSTSQEPLRVSNSGQEAG 242
>TC212058
Length = 1017
Score = 27.3 bits (59), Expect = 3.4
Identities = 13/35 (37%), Positives = 20/35 (57%)
Frame = +2
Query: 68 ELAMPIEKVKETSKLSLMRTVEVWLRKERDEDRRQ 102
EL I K +E S LSL R ++WL + + +R+
Sbjct: 764 ELLRGISKTRERSMLSLQRIKDLWLSTKLNYQKRK 868
>TC233121 similar to GB|AAP13385.1|30023704|BT006277 At3g56360 {Arabidopsis
thaliana;} , partial (36%)
Length = 990
Score = 26.9 bits (58), Expect = 4.4
Identities = 12/32 (37%), Positives = 21/32 (65%)
Frame = +2
Query: 65 GRLELAMPIEKVKETSKLSLMRTVEVWLRKER 96
GR+ AMP K +ET+ + +M +E+ ++ER
Sbjct: 227 GRITAAMPTVKEQETTPMIMMGAMELLGKRER 322
>BM270917 similar to GP|4586576|dbj multidrug resistance protein {Cicer
arietinum}, partial (46%)
Length = 443
Score = 26.6 bits (57), Expect = 5.8
Identities = 15/40 (37%), Positives = 23/40 (57%)
Frame = +3
Query: 8 QVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATH 47
+VLLLDE T LD ++ ++ L K +G T+I +H
Sbjct: 234 KVLLLDEPTSALDPISTENIEDALVKLNKNQGMTVIMVSH 353
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.320 0.139 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,018,200
Number of Sequences: 63676
Number of extensions: 63741
Number of successful extensions: 327
Number of sequences better than 10.0: 48
Number of HSP's better than 10.0 without gapping: 326
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 326
length of query: 165
length of database: 12,639,632
effective HSP length: 90
effective length of query: 75
effective length of database: 6,908,792
effective search space: 518159400
effective search space used: 518159400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 55 (25.8 bits)
Medicago: description of AC147011.10


