

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147000.4 + phase: 1 /pseudo
(187 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
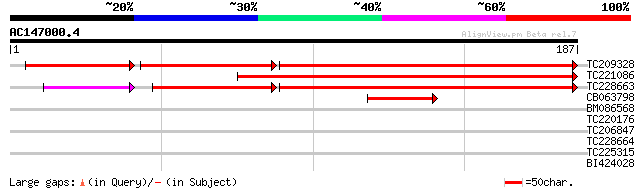
Score E
Sequences producing significant alignments: (bits) Value
TC209328 similar to UP|KAPS_CATRO (O49204) Adenylyl-sulfate kina... 186 2e-74
TC221086 similar to UP|Q9SE92 (Q9SE92) Adenosine-5'-phosphosulfa... 159 7e-40
TC228663 similar to UP|Q9SE92 (Q9SE92) Adenosine-5'-phosphosulfa... 145 8e-37
CB063798 GP|6563285|gb| adenosine-5'-phosphosulfate kinase {Zea ... 47 4e-06
BM086568 27 7.2
TC220176 27 7.2
TC206847 similar to GB|AAF76441.1|8569096|F2J10 ESTs gb|AI994059... 27 7.2
TC228664 weakly similar to UP|PPS_URECA (Q27128) Bifunctional 3'... 23 8.3
TC225315 similar to UP|Q41414 (Q41414) Epoxide hydrolase, partia... 26 9.4
BI424028 26 9.4
>TC209328 similar to UP|KAPS_CATRO (O49204) Adenylyl-sulfate kinase,
chloroplast precursor (APS kinase)
(Adenosine-5'phosphosulfate kinase) (ATP
adenosine-5'-phosphosulfate 3'-phosphotransferase) ,
partial (69%)
Length = 1056
Score = 186 bits (471), Expect(3) = 2e-74
Identities = 86/98 (87%), Positives = 94/98 (95%)
Frame = +1
Query: 90 LISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPY 149
LISPYQKDRDACRAL+P+GDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPY
Sbjct: 487 LISPYQKDRDACRALMPKGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPY 666
Query: 150 EPPCCCEIILQQKGSDCKSPKDMAETVISYLEKSGHLQ 187
EPP CEI+LQQKGS+CKSP DMAE VISYLE++G+L+
Sbjct: 667 EPPSSCEIVLQQKGSNCKSPSDMAEEVISYLEENGYLR 780
Score = 73.9 bits (180), Expect(3) = 2e-74
Identities = 39/45 (86%), Positives = 39/45 (86%)
Frame = +3
Query: 44 CMCFESKLALQRKTDLHP*R*QYSAWSKP*S*F*SRRSFRKHTKD 88
CMCFESKLALQRKT LHP* *QYSAWSKP S*F SRRSF KH KD
Sbjct: 300 CMCFESKLALQRKTVLHP*W*QYSAWSKPRS*FQSRRSF*KH*KD 434
Score = 57.8 bits (138), Expect(3) = 2e-74
Identities = 28/36 (77%), Positives = 30/36 (82%)
Frame = +2
Query: 6 TRRTFCGMIVQFKNVIDSSCFSKKDVLYG*LVSVVQ 41
TR+ CGM VQF+N IDSSCFSKK VLYG*L SVVQ
Sbjct: 179 TRQILCGMTVQFRNKIDSSCFSKKAVLYG*LASVVQ 286
>TC221086 similar to UP|Q9SE92 (Q9SE92) Adenosine-5'-phosphosulfate kinase
(Fragment) , partial (39%)
Length = 518
Score = 159 bits (402), Expect = 7e-40
Identities = 73/112 (65%), Positives = 93/112 (82%)
Frame = +3
Query: 76 F*SRRSFRKHTKDCLISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLAR 135
F SRRS K++++CLISPY++DRD CRA+LP+ +FIEVF+++PL +CEARDPKGLYKLAR
Sbjct: 6 FQSRRSR*KYSQNCLISPYRRDRDTCRAMLPDANFIEVFMNMPLELCEARDPKGLYKLAR 185
Query: 136 AGKIKGFTGIDDPYEPPCCCEIILQQKGSDCKSPKDMAETVISYLEKSGHLQ 187
AGKIKGFTGIDDPYEPP CEI ++Q+ DC +P MA V++YLE G L+
Sbjct: 186 AGKIKGFTGIDDPYEPPINCEIEIKQENGDCPTPTLMAGQVVTYLENKGFLE 341
>TC228663 similar to UP|Q9SE92 (Q9SE92) Adenosine-5'-phosphosulfate kinase
(Fragment) , partial (68%)
Length = 940
Score = 145 bits (366), Expect(3) = 8e-37
Identities = 65/98 (66%), Positives = 82/98 (83%)
Frame = +3
Query: 90 LISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPY 149
LISPY++DRD CRA+LP+ +FIEVF+++PL +CEARDPKGLYKLARAGKIKGFTGIDDPY
Sbjct: 444 LISPYRRDRDTCRAMLPDANFIEVFMNMPLELCEARDPKGLYKLARAGKIKGFTGIDDPY 623
Query: 150 EPPCCCEIILQQKGSDCKSPKDMAETVISYLEKSGHLQ 187
EPP CEI ++Q+ +C +P MA V++YLE G L+
Sbjct: 624 EPPINCEIEIKQENGNCPTPTMMAGQVVTYLENKGFLE 737
Score = 22.7 bits (47), Expect(3) = 8e-37
Identities = 17/41 (41%), Positives = 25/41 (60%)
Frame = +2
Query: 48 ESKLALQRKTDLHP*R*QYSAWSKP*S*F*SRRSFRKHTKD 88
+ + AL+ K L P* SAW+K S F*S RS K++++
Sbjct: 269 KQRTALKGKVILCP*WR*PSAWTKQGSWF*S*RSH*KYSQN 391
Score = 22.7 bits (47), Expect(3) = 8e-37
Identities = 13/30 (43%), Positives = 18/30 (59%)
Frame = +1
Query: 12 GMIVQFKNVIDSSCFSKKDVLYG*LVSVVQ 41
G IV+ + + S +K+DVLYG L S Q
Sbjct: 154 GKIVK*EGLKGRSYLTKRDVLYGLLDSADQ 243
>CB063798 GP|6563285|gb| adenosine-5'-phosphosulfate kinase {Zea mays},
partial (7%)
Length = 378
Score = 47.4 bits (111), Expect = 4e-06
Identities = 21/23 (91%), Positives = 22/23 (95%)
Frame = +1
Query: 119 LHVCEARDPKGLYKLARAGKIKG 141
L +CEARDPKGLYKLARAGKIKG
Sbjct: 1 LELCEARDPKGLYKLARAGKIKG 69
>BM086568
Length = 431
Score = 26.6 bits (57), Expect = 7.2
Identities = 11/31 (35%), Positives = 19/31 (60%), Gaps = 2/31 (6%)
Frame = +3
Query: 152 PCCCEIILQQKGSDC--KSPKDMAETVISYL 180
P C ++ ++KGS C SP+ ++ + SYL
Sbjct: 168 PFWCLLLTEEKGSQCIRSSPQPLSHVLFSYL 260
>TC220176
Length = 991
Score = 26.6 bits (57), Expect = 7.2
Identities = 10/25 (40%), Positives = 14/25 (56%)
Frame = +3
Query: 148 PYEPPCCCEIILQQKGSDCKSPKDM 172
P E CCC + Q+K K+P D+
Sbjct: 423 PMECICCCNLFRQEKLKFMKNPNDL 497
>TC206847 similar to GB|AAF76441.1|8569096|F2J10 ESTs gb|AI994059, gb|T43740
come from this gene. {Arabidopsis thaliana;} , partial
(46%)
Length = 1571
Score = 26.6 bits (57), Expect = 7.2
Identities = 16/54 (29%), Positives = 25/54 (45%)
Frame = +1
Query: 132 KLARAGKIKGFTGIDDPYEPPCCCEIILQQKGSDCKSPKDMAETVISYLEKSGH 185
K + K F+G P P C+I+LQ S+ SP+ + +IS + H
Sbjct: 46 KSPASSKPSSFSGKVHPSSSPFFCKILLQGLDSNSTSPRVSSVPLISDSDPHPH 207
>TC228664 weakly similar to UP|PPS_URECA (Q27128) Bifunctional
3'-phosphoadenosine 5'-phosphosulfate synthethase (PAPS
synthethase) (PAPSS) (Sulfurylase kinase) (SK)
[Includes: Sulfate adenylyltransferase (Sulfate
adenylate transferase) (SAT) (ATP-sulfurylase);
Adenylyl-s , partial (9%)
Length = 567
Score = 22.7 bits (47), Expect(2) = 8.3
Identities = 13/30 (43%), Positives = 18/30 (59%)
Frame = +1
Query: 12 GMIVQFKNVIDSSCFSKKDVLYG*LVSVVQ 41
G IV+ + + S +K+DVLYG L S Q
Sbjct: 253 GKIVK*EGLKGRSYLTKRDVLYGLLDSADQ 342
Score = 21.9 bits (45), Expect(2) = 8.3
Identities = 16/41 (39%), Positives = 24/41 (58%)
Frame = +2
Query: 48 ESKLALQRKTDLHP*R*QYSAWSKP*S*F*SRRSFRKHTKD 88
+ + AL+ K L P* S W+K S F SRRS K++++
Sbjct: 368 KQRTALKGKVILCP*WR*PSTWTKQGSWFQSRRSR*KYSQN 490
>TC225315 similar to UP|Q41414 (Q41414) Epoxide hydrolase, partial (34%)
Length = 643
Score = 26.2 bits (56), Expect = 9.4
Identities = 10/20 (50%), Positives = 11/20 (55%)
Frame = -3
Query: 147 DPYEPPCCCEIILQQKGSDC 166
D Y PCCCE+ K DC
Sbjct: 290 DLYSSPCCCELPGYVKRIDC 231
>BI424028
Length = 421
Score = 26.2 bits (56), Expect = 9.4
Identities = 11/29 (37%), Positives = 17/29 (57%)
Frame = +3
Query: 155 CEIILQQKGSDCKSPKDMAETVISYLEKS 183
C L KGS+CKS + ++SYL ++
Sbjct: 144 CSHQLCDKGSNCKSSQQQCPYMVSYLSEN 230
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.340 0.150 0.512
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,013,054
Number of Sequences: 63676
Number of extensions: 137923
Number of successful extensions: 906
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 900
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 904
length of query: 187
length of database: 12,639,632
effective HSP length: 92
effective length of query: 95
effective length of database: 6,781,440
effective search space: 644236800
effective search space used: 644236800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 56 (26.2 bits)
Medicago: description of AC147000.4


