

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146972.8 + phase: 0
(198 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
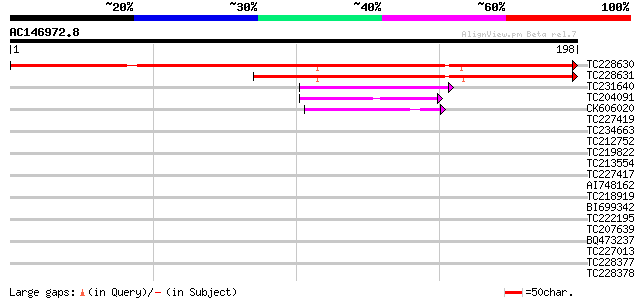
Sequences producing significant alignments: (bits) Value
TC228630 similar to UP|Q7XBE3 (Q7XBE3) Humj1, partial (64%) 266 5e-72
TC228631 similar to UP|Q7XBE3 (Q7XBE3) Humj1, partial (50%) 173 5e-44
TC231640 homologue to UP|Q918P0 (Q918P0) Latency associated anti... 44 5e-05
TC204091 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1... 40 7e-04
CK606020 40 7e-04
TC227419 39 0.002
TC234663 39 0.002
TC212752 weakly similar to UP|Q7M4A3 (Q7M4A3) Spermatid-specific... 38 0.003
TC219822 homologue to UP|CENB_HUMAN (P07199) Major centromere au... 38 0.003
TC213554 weakly similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protei... 37 0.005
TC227417 37 0.006
AI748162 similar to PIR|S20007|S200 beta-conglycinin alpha chain... 37 0.008
TC218919 homologue to UP|Q39892 (Q39892) Nucleosome assembly pro... 35 0.017
BI699342 35 0.017
TC222195 35 0.030
TC207639 similar to UP|O48592 (O48592) GT2 protein, partial (24%) 35 0.030
BQ473237 34 0.039
TC227013 similar to UP|GAC1_YEAST (P28006) Protein phosphatase 1... 34 0.039
TC228377 34 0.039
TC228378 34 0.039
>TC228630 similar to UP|Q7XBE3 (Q7XBE3) Humj1, partial (64%)
Length = 782
Score = 266 bits (680), Expect = 5e-72
Identities = 144/201 (71%), Positives = 153/201 (75%), Gaps = 3/201 (1%)
Frame = +3
Query: 1 MEAITASLERSLQNCSLNNNNQNEEGSATIDVVAGEGRGGGGGGIGISSSSSSSDENHIS 60
MEAITASLERSLQNCSLNN N N + S + G GG IGISS S + S
Sbjct: 111 MEAITASLERSLQNCSLNNTNNNHQQSRRPEEDDGSATDGG---IGISSCSDDVPDPDNS 281
Query: 61 NNSDATLELNSNISLPYHWEQCLDLKTGEIYYINWRNGMKAKEDPR-RAAERECEESEEE 119
+ SD TLELNS+ISLPYHWEQCLDLKTGEIYYINWRNGMKAKEDPR R A+ E
Sbjct: 282 HISDTTLELNSHISLPYHWEQCLDLKTGEIYYINWRNGMKAKEDPRSRRADYSVSRDCES 461
Query: 120 EEEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQ--NVLVVAGCKICLMYFMVPKQV 177
EEEEEEESWYDSEESSSES SKE YD+ +V EKQ NVLVVAGCK CLMYFMVPKQV
Sbjct: 462 EEEEEEESWYDSEESSSESCPSSSKEHYDEGQV-EKQNNNVLVVAGCKGCLMYFMVPKQV 638
Query: 178 EDCPKCSGQLLHFDRSENCSP 198
EDCPKC+GQLLHFDRSEN SP
Sbjct: 639 EDCPKCTGQLLHFDRSENGSP 701
>TC228631 similar to UP|Q7XBE3 (Q7XBE3) Humj1, partial (50%)
Length = 632
Score = 173 bits (438), Expect = 5e-44
Identities = 90/116 (77%), Positives = 95/116 (81%), Gaps = 3/116 (2%)
Frame = +1
Query: 86 KTGEIYYINWRNGMKAKEDPR-RAAERECEESEEEEEEEEEESWYDSEESSSESSTIISK 144
+TGEIYYINWRNGMKAKEDPR R A+ E EEEEEEESWYDSEESSSES SK
Sbjct: 130 QTGEIYYINWRNGMKAKEDPRSRRADYSVSRDCESEEEEEEESWYDSEESSSESCPSSSK 309
Query: 145 EQYDQREVIEKQN--VLVVAGCKICLMYFMVPKQVEDCPKCSGQLLHFDRSENCSP 198
E YD+ +V EKQN VLVVAGCK CLMYFMVPKQVEDCPKC+GQLLHFDRSEN SP
Sbjct: 310 EHYDEGQV-EKQNNNVLVVAGCKGCLMYFMVPKQVEDCPKCTGQLLHFDRSENGSP 474
>TC231640 homologue to UP|Q918P0 (Q918P0) Latency associated antigen, partial
(7%)
Length = 466
Score = 43.9 bits (102), Expect = 5e-05
Identities = 22/54 (40%), Positives = 31/54 (56%)
Frame = +1
Query: 102 KEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEK 155
+E+ E E EE EEEEEEEEEE + EE II ++++D E+ +K
Sbjct: 31 EEEKEEEEEEEEEEEEEEEEEEEEEEKEEEEEEEGNVLVIIDEDKHDTEELNKK 192
Score = 33.5 bits (75), Expect = 0.066
Identities = 17/46 (36%), Positives = 26/46 (55%)
Frame = +1
Query: 116 SEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQNVLVV 161
S+ +EEEEE+E + EE E +E+ + E E+ NVLV+
Sbjct: 13 SDTQEEEEEKEEEEEEEEEEEEEEEEEEEEEEKEEEEEEEGNVLVI 150
>TC204091 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1, partial
(70%)
Length = 2075
Score = 40.0 bits (92), Expect = 7e-04
Identities = 19/50 (38%), Positives = 29/50 (58%)
Frame = +2
Query: 102 KEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQRE 151
+E E E EE EEEEEE+EEE + EE E + ++ ++Q Q++
Sbjct: 140 EEVEEEEVEEEVEEEEEEEEEDEEEE--EEEEEEEEENDVVKQQQQQQQD 283
Score = 33.9 bits (76), Expect = 0.050
Identities = 19/54 (35%), Positives = 30/54 (55%)
Frame = +2
Query: 103 EDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQ 156
E+P + E E EE EEEE EEE + EE E +E+ ++ +V+++Q
Sbjct: 107 ENPPQEYE-EVEEEVEEEEVEEEVEEEEEEEEEDEEEEEEEEEEEEENDVVKQQ 265
Score = 32.7 bits (73), Expect = 0.11
Identities = 20/62 (32%), Positives = 32/62 (51%)
Frame = +2
Query: 99 MKAKEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQNV 158
+K ++ +E EE EEE EEEE E + EE E +E+ ++ E E++N
Sbjct: 83 LKQQQPKPENPPQEYEEVEEEVEEEEVEEEVEEEEEEEEE----DEEEEEEEEEEEEEND 250
Query: 159 LV 160
+V
Sbjct: 251 VV 256
Score = 29.6 bits (65), Expect = 0.95
Identities = 17/46 (36%), Positives = 24/46 (51%), Gaps = 1/46 (2%)
Frame = +2
Query: 100 KAKEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTI-ISK 144
+ +E+ E E EE EEEEEEEE + ++ + I ISK
Sbjct: 170 EVEEEEEEEEEDEEEEEEEEEEEEENDVVKQQQQQQQDEDDIPISK 307
>CK606020
Length = 340
Score = 40.0 bits (92), Expect = 7e-04
Identities = 22/49 (44%), Positives = 27/49 (54%)
Frame = +1
Query: 104 DPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREV 152
DP E E EE EEEEEEEEEE + EE E +E+ ++ EV
Sbjct: 184 DPSLDEEEEEEEEEEEEEEEEEEEEIEEEEEEEEEEV---EEEVEEEEV 321
Score = 38.9 bits (89), Expect = 0.002
Identities = 20/43 (46%), Positives = 25/43 (57%)
Frame = +1
Query: 110 ERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREV 152
E E EE EEEEEEEEEE + EE E + +E+ + EV
Sbjct: 208 EEEEEEEEEEEEEEEEEIEEEEEEEEEEVEEEVEEEEVEGEEV 336
Score = 34.3 bits (77), Expect = 0.039
Identities = 18/40 (45%), Positives = 22/40 (55%)
Frame = +1
Query: 102 KEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTI 141
+E+ E E EE EEEEEEEEEE + EE E +
Sbjct: 217 EEEEEEEEEEEEEEIEEEEEEEEEEVEEEVEEEEVEGEEV 336
Score = 33.5 bits (75), Expect = 0.066
Identities = 19/62 (30%), Positives = 29/62 (46%)
Frame = +1
Query: 97 NGMKAKEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQ 156
N M + P + +EEEEEEEEE + EE E +E+ + E +E++
Sbjct: 136 NPMSLPDSPVANDDEIDPSLDEEEEEEEEEEEEEEEEEEEEIEEEEEEEEEEVEEEVEEE 315
Query: 157 NV 158
V
Sbjct: 316 EV 321
Score = 32.0 bits (71), Expect = 0.19
Identities = 17/44 (38%), Positives = 22/44 (49%)
Frame = +1
Query: 102 KEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKE 145
+E+ E E EE EEEE EEEEE + E E + +E
Sbjct: 202 EEEEEEEEEEEEEEEEEEEIEEEEEEEEEEVEEEVEEEEVEGEE 333
Score = 28.1 bits (61), Expect = 2.8
Identities = 21/77 (27%), Positives = 37/77 (47%)
Frame = -3
Query: 5 TASLERSLQNCSLNNNNQNEEGSATIDVVAGEGRGGGGGGIGISSSSSSSDENHISNNSD 64
T+S S + S + ++ + S++I + SSSSSSS + ++S
Sbjct: 335 TSSPSTSSSSTSSSTSSSSSSSSSSISSSSSSSSSSSSS----SSSSSSSSSDGSISSSL 168
Query: 65 ATLELNSNISLPYHWEQ 81
AT E S+I L + W++
Sbjct: 167 ATGESGSDIGLNWIWKR 117
>TC227419
Length = 688
Score = 38.5 bits (88), Expect = 0.002
Identities = 33/119 (27%), Positives = 51/119 (42%), Gaps = 5/119 (4%)
Frame = +2
Query: 71 SNISLPYHWEQCLDLK--TGEIYYINWRNGMKAKEDPRRAAERECEESEEEEEEEEEESW 128
SN+ L E LDLK + + N + + + A ER +E + +S
Sbjct: 71 SNVLLNLFDETSLDLKLVSSTLPSSNNYQSVCTLDKVKSALERAEKEPIVRKRTSFLKSP 250
Query: 129 YDSEESSSESSTIISKEQYDQREVIEKQNVL---VVAGCKICLMYFMVPKQVEDCPKCS 184
+ S SS+ +E + E E+ NV + AGC CL Y M+ K CP+C+
Sbjct: 251 LSASSPSYSSSSSSIRETIQEEECEERLNVSSSPLAAGCPGCLSYVMIMKNNPKCPRCN 427
>TC234663
Length = 773
Score = 38.5 bits (88), Expect = 0.002
Identities = 25/82 (30%), Positives = 35/82 (42%), Gaps = 4/82 (4%)
Frame = +3
Query: 121 EEEEEESW---YDSEESSSESSTIISKEQYDQREVIEKQNVLVVAGCKICLMYFMVPKQV 177
E EE SW S SSS I +E + + + + GC C MY M+ +
Sbjct: 237 ELPEEFSWSSCLSSSVSSSSKIPIEDEENVQRANASPETKAMDLVGCPRCFMYVMLSEVD 416
Query: 178 EDCPKCSGQL-LHFDRSENCSP 198
CPKC + L + EN +P
Sbjct: 417 PKCPKCKTTVFLELFKDENFNP 482
>TC212752 weakly similar to UP|Q7M4A3 (Q7M4A3) Spermatid-specific protein T2
precursor, partial (35%)
Length = 533
Score = 37.7 bits (86), Expect = 0.003
Identities = 20/33 (60%), Positives = 22/33 (66%), Gaps = 2/33 (6%)
Frame = +3
Query: 102 KEDPRRAAER--ECEESEEEEEEEEEESWYDSE 132
+E+P R E E EE EEEEEEEEEE W D E
Sbjct: 39 EEEPHRMEEDDDESEEEEEEEEEEEEEVWDDWE 137
Score = 28.5 bits (62), Expect = 2.1
Identities = 16/51 (31%), Positives = 22/51 (42%), Gaps = 9/51 (17%)
Frame = +3
Query: 100 KAKEDPRRAAERECEESEEEEEEEEEE---------SWYDSEESSSESSTI 141
K K++ R E+ +E EEEEEEE W +E ES +
Sbjct: 21 KQKQEEEEEPHRMEEDDDESEEEEEEEEEEEEEVWDDWEGEDEGERESEFV 173
>TC219822 homologue to UP|CENB_HUMAN (P07199) Major centromere autoantigen B
(Centromere protein B) (CENP-B), partial (5%)
Length = 710
Score = 37.7 bits (86), Expect = 0.003
Identities = 24/65 (36%), Positives = 30/65 (45%)
Frame = +1
Query: 97 NGMKAKEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQ 156
N A +P E E EE EE EEEEEEE + EE + S E+ +
Sbjct: 397 NYAPALAEPEEEEEEEEEEEEENEEEEEEEEGEEEEEEEARSMV---------NELAQLD 549
Query: 157 NVLVV 161
N+LVV
Sbjct: 550 NMLVV 564
Score = 29.3 bits (64), Expect = 1.2
Identities = 21/71 (29%), Positives = 33/71 (45%)
Frame = +1
Query: 102 KEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQNVLVV 161
+E+ E E EE E EEEEEEE +E + ++ ++ + E I K +V +
Sbjct: 442 EEEEEEENEEEEEEEEGEEEEEEEARSMVNELAQLDNMLVVIRSLKGTLETI-KVDVQAI 618
Query: 162 AGCKICLMYFM 172
CL F+
Sbjct: 619 KERVKCLQSFL 651
>TC213554 weakly similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1,
partial (18%)
Length = 451
Score = 37.4 bits (85), Expect = 0.005
Identities = 20/52 (38%), Positives = 27/52 (51%)
Frame = +2
Query: 105 PRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQ 156
P AA+ EE EEE EEEE E + EE E +E+ DQ++ + Q
Sbjct: 167 PENAAQEYEEEVEEEVEEEEVEEEVEEEEEEEEDEEDEEEEEEDQKDQYQHQ 322
Score = 34.7 bits (78), Expect = 0.030
Identities = 19/50 (38%), Positives = 23/50 (46%)
Frame = +2
Query: 102 KEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQRE 151
+E+ E E E E EEEEEEEE D EE + +Q Q E
Sbjct: 191 EEEVEEEVEEEEVEEEVEEEEEEEEDEEDEEEEEEDQKDQYQHQQQQQDE 340
Score = 33.1 bits (74), Expect = 0.086
Identities = 19/54 (35%), Positives = 27/54 (49%)
Frame = +2
Query: 103 EDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQ 156
E+ + E E EE EEEE EEE + EE E ++Q DQ + ++Q
Sbjct: 170 ENAAQEYEEEVEEEVEEEEVEEEVEEEEEEEEDEEDEEEEEEDQKDQYQHQQQQ 331
Score = 33.1 bits (74), Expect = 0.086
Identities = 20/52 (38%), Positives = 26/52 (49%), Gaps = 1/52 (1%)
Frame = +2
Query: 101 AKEDPRRAAERECEESE-EEEEEEEEESWYDSEESSSESSTIISKEQYDQRE 151
A ++ E E EE E EEE EEEEE D E+ E K+QY ++
Sbjct: 176 AAQEYEEEVEEEVEEEEVEEEVEEEEEEEEDEEDEEEEEED--QKDQYQHQQ 325
Score = 32.3 bits (72), Expect = 0.15
Identities = 14/35 (40%), Positives = 21/35 (60%)
Frame = +2
Query: 110 ERECEESEEEEEEEEEESWYDSEESSSESSTIISK 144
E E EE EEEEEE++++ + ++ E ISK
Sbjct: 257 EEEDEEDEEEEEEDQKDQYQHQQQQQDEDDDPISK 361
Score = 30.8 bits (68), Expect = 0.43
Identities = 18/53 (33%), Positives = 26/53 (48%)
Frame = +2
Query: 103 EDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEK 155
E+ E E E EEEEEEE+EE + EE + ++Q + + I K
Sbjct: 203 EEEVEEEEVEEEVEEEEEEEEDEEDEEEEEEDQKDQYQHQQQQQDEDDDPISK 361
>TC227417
Length = 1067
Score = 37.0 bits (84), Expect = 0.006
Identities = 21/57 (36%), Positives = 29/57 (50%), Gaps = 3/57 (5%)
Frame = +2
Query: 131 SEESSSESSTIISKEQYDQREVIEKQNVL---VVAGCKICLMYFMVPKQVEDCPKCS 184
S S S SS+ I + + E E+ NV + AGC CL Y M+ K CP+C+
Sbjct: 617 SSPSYSSSSSSIRETTIQEDECEERLNVSSSPMAAGCPGCLSYVMIMKNNPKCPRCN 787
Score = 30.0 bits (66), Expect = 0.73
Identities = 22/88 (25%), Positives = 41/88 (46%), Gaps = 4/88 (4%)
Frame = +2
Query: 8 LERSLQNCSLNNNNQNEEGSATIDVVAGEGRGGGGGGIGISSSSSSSD----ENHISNNS 63
L SL++ E S+ I V+ G + +G S S + + +S
Sbjct: 50 LSLSLESSVRTRTAMAAEVSSLIRVMGGGYKEEQHRTVGNESHGEKSTALITRDLLGGSS 229
Query: 64 DATLELNSNISLPYHWEQCLDLKTGEIY 91
+ EL+ ++ +P WE+ LDL++G++Y
Sbjct: 230 IESQELDLDLQVPSGWEKRLDLQSGKVY 313
>AI748162 similar to PIR|S20007|S200 beta-conglycinin alpha chain precursor -
soybean, partial (13%)
Length = 253
Score = 36.6 bits (83), Expect = 0.008
Identities = 21/63 (33%), Positives = 32/63 (50%), Gaps = 10/63 (15%)
Frame = +2
Query: 105 PRRAAERECEESEEEEEEEEEESWYDSEESSSESST----------IISKEQYDQREVIE 154
P AER+ EE E+EE + E E DSE + ++ T + K QYD+ V++
Sbjct: 29 PHENAERKREEDEDEEHQPESEDSEDSESRTHKNKTPFLFGSNRLDTLFKHQYDRIRVLQ 208
Query: 155 KQN 157
+ N
Sbjct: 209 RIN 217
>TC218919 homologue to UP|Q39892 (Q39892) Nucleosome assembly protein 1,
partial (33%)
Length = 729
Score = 35.4 bits (80), Expect = 0.017
Identities = 17/54 (31%), Positives = 30/54 (55%)
Frame = +1
Query: 93 INWRNGMKAKEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQ 146
++W G A+ D + E +E EEE+E+E+E+ YD +E E +K++
Sbjct: 139 VSWFTGEXAQGD--EFEDLEDDEDEEEDEDEDEDEEYDEDEDDEEEDDTKTKKK 294
>BI699342
Length = 325
Score = 35.4 bits (80), Expect = 0.017
Identities = 18/33 (54%), Positives = 22/33 (66%)
Frame = -1
Query: 109 AERECEESEEEEEEEEEESWYDSEESSSESSTI 141
AE E EE EEEEEEEEEE + EE + + T+
Sbjct: 268 AEEEEEEEEEEEEEEEEEEEEEEEEEKAINLTM 170
Score = 34.3 bits (77), Expect = 0.039
Identities = 17/28 (60%), Positives = 19/28 (67%)
Frame = -1
Query: 108 AAERECEESEEEEEEEEEESWYDSEESS 135
A E E EE EEEEEEEEEE + EE +
Sbjct: 268 AEEEEEEEEEEEEEEEEEEEEEEEEEKA 185
Score = 33.5 bits (75), Expect = 0.066
Identities = 16/26 (61%), Positives = 18/26 (68%)
Frame = -1
Query: 101 AKEDPRRAAERECEESEEEEEEEEEE 126
A+E+ E E EE EEEEEEEEEE
Sbjct: 268 AEEEEEEEEEEEEEEEEEEEEEEEEE 191
Score = 31.6 bits (70), Expect = 0.25
Identities = 15/29 (51%), Positives = 17/29 (57%)
Frame = -1
Query: 115 ESEEEEEEEEEESWYDSEESSSESSTIIS 143
E EEEEEEEEEE + EE E I+
Sbjct: 265 EEEEEEEEEEEEEEEEEEEEEEEEEKAIN 179
Score = 30.8 bits (68), Expect = 0.43
Identities = 14/26 (53%), Positives = 18/26 (68%)
Frame = -1
Query: 102 KEDPRRAAERECEESEEEEEEEEEES 127
+E+ E E EE EEEEEEEEE++
Sbjct: 262 EEEEEEEEEEEEEEEEEEEEEEEEKA 185
Score = 28.1 bits (61), Expect = 2.8
Identities = 13/28 (46%), Positives = 17/28 (60%)
Frame = -1
Query: 97 NGMKAKEDPRRAAERECEESEEEEEEEE 124
N + +E+ E E EE EEEEEEE+
Sbjct: 271 NAEEEEEEEEEEEEEEEEEEEEEEEEEK 188
Score = 27.3 bits (59), Expect = 4.7
Identities = 16/46 (34%), Positives = 26/46 (55%)
Frame = +3
Query: 45 IGISSSSSSSDENHISNNSDATLELNSNISLPYHWEQCLDLKTGEI 90
I SSSSSSS + S++S ++ +S+ YH + L +G+I
Sbjct: 183 IAFSSSSSSSSSSSSSSSSSSSSSSSSSAFEVYHLKMRLKFISGKI 320
>TC222195
Length = 844
Score = 34.7 bits (78), Expect = 0.030
Identities = 23/76 (30%), Positives = 32/76 (41%), Gaps = 8/76 (10%)
Frame = +1
Query: 116 SEEEEEEEEEESWYDSEESSSESSTII-------SKEQYDQRE-VIEKQNVLVVAGCKIC 167
S E W + SSS SS S+E+ +QRE + + + GC C
Sbjct: 193 SAE*SSSTSSGEWPELSSSSSASSVSCTSKMPKESEEENEQRENSTPETKAMDLVGCPRC 372
Query: 168 LMYFMVPKQVEDCPKC 183
MY M+ + CPKC
Sbjct: 373 FMYVMLSEVDPKCPKC 420
>TC207639 similar to UP|O48592 (O48592) GT2 protein, partial (24%)
Length = 1689
Score = 34.7 bits (78), Expect = 0.030
Identities = 19/44 (43%), Positives = 25/44 (56%)
Frame = +1
Query: 95 WRNGMKAKEDPRRAAERECEESEEEEEEEEEESWYDSEESSSES 138
+ G + + ERE EE +EEEEEEEEE ++EE ES
Sbjct: 874 YEQGAAKENNNNERKEREEEEEDEEEEEEEEED--ENEEGDLES 999
Score = 29.3 bits (64), Expect = 1.2
Identities = 16/34 (47%), Positives = 21/34 (61%)
Frame = +1
Query: 112 ECEESEEEEEEEEEESWYDSEESSSESSTIISKE 145
E +E EEEEE+EEEE + EE +E + S E
Sbjct: 910 ERKEREEEEEDEEEEE--EEEEDENEEGDLESVE 1005
Score = 28.1 bits (61), Expect = 2.8
Identities = 14/28 (50%), Positives = 16/28 (57%)
Frame = +1
Query: 110 ERECEESEEEEEEEEEESWYDSEESSSE 137
E E +E EEEEEEE+E D E E
Sbjct: 928 EEEEDEEEEEEEEEDENEEGDLESVEDE 1011
Score = 26.9 bits (58), Expect = 6.2
Identities = 12/43 (27%), Positives = 21/43 (47%)
Frame = +1
Query: 95 WRNGMKAKEDPRRAAERECEESEEEEEEEEEESWYDSEESSSE 137
WR + ++ + + EEEEE+EEE + E+ + E
Sbjct: 856 WRPPSQYEQGAAKENNNNERKEREEEEEDEEEEEEEEEDENEE 984
>BQ473237
Length = 408
Score = 34.3 bits (77), Expect = 0.039
Identities = 20/57 (35%), Positives = 29/57 (50%), Gaps = 2/57 (3%)
Frame = -2
Query: 134 SSSESSTIISKEQYDQREVIEKQNVLVVAGCKICLMYFMVPKQVED--CPKCSGQLL 188
SSS+ + +S E D E+ +V+ GC CLMY + K + CPKC +L
Sbjct: 386 SSSDPDSCVSSETND-----EEIRSMVLVGCLRCLMYVLFSKDDPNPKCPKCKSTVL 231
>TC227013 similar to UP|GAC1_YEAST (P28006) Protein phosphatase 1 regulatory
subunit GAC1, partial (3%)
Length = 1149
Score = 34.3 bits (77), Expect = 0.039
Identities = 19/50 (38%), Positives = 30/50 (60%)
Frame = +2
Query: 103 EDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREV 152
E+ + E E EE EEEEEEEEEE + ++S+ S I + + +D ++
Sbjct: 206 ENEEKHNEDEIEE-EEEEEEEEEEEEFSFVLTNSDGSPISADDAFDNGQI 352
>TC228377
Length = 958
Score = 34.3 bits (77), Expect = 0.039
Identities = 13/30 (43%), Positives = 18/30 (59%)
Frame = +1
Query: 159 LVVAGCKICLMYFMVPKQVEDCPKCSGQLL 188
+V+ GC CLMY M+ + CPKC +L
Sbjct: 397 MVLVGCPRCLMYVMLSEDDPKCPKCKSTVL 486
>TC228378
Length = 602
Score = 34.3 bits (77), Expect = 0.039
Identities = 13/30 (43%), Positives = 18/30 (59%)
Frame = +3
Query: 159 LVVAGCKICLMYFMVPKQVEDCPKCSGQLL 188
+V+ GC CLMY M+ + CPKC +L
Sbjct: 36 MVLVGCPRCLMYVMLSEDDPKCPKCKSTVL 125
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.306 0.126 0.358
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,224,698
Number of Sequences: 63676
Number of extensions: 158915
Number of successful extensions: 4072
Number of sequences better than 10.0: 283
Number of HSP's better than 10.0 without gapping: 2099
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3141
length of query: 198
length of database: 12,639,632
effective HSP length: 92
effective length of query: 106
effective length of database: 6,781,440
effective search space: 718832640
effective search space used: 718832640
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 43 (22.0 bits)
S2: 56 (26.2 bits)
Medicago: description of AC146972.8


