

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146972.11 + phase: 0
(362 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
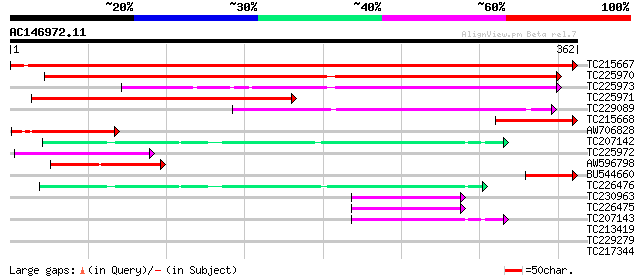
Sequences producing significant alignments: (bits) Value
TC215667 similar to UP|ODPB_PEA (P52904) Pyruvate dehydrogenase ... 658 0.0
TC225970 homologue to GB|AAB86804.1|2454184|ATU80186 pyruvate de... 269 1e-72
TC225973 homologue to GB|AAB86804.1|2454184|ATU80186 pyruvate de... 166 2e-41
TC225971 homologue to GB|AAB86804.1|2454184|ATU80186 pyruvate de... 146 2e-35
TC229089 similar to UP|O82450 (O82450) Branched-chain alpha-keto... 140 1e-33
TC215668 homologue to UP|Q9ZQY1 (Q9ZQY1) Pyruvate dehydrogenase ... 98 5e-21
AW706828 weakly similar to SP|P52904|ODPB_ Pyruvate dehydrogenas... 89 4e-18
TC207142 similar to UP|Q8L692 (Q8L692) 1-deoxy-D-xylulose 5-phos... 63 2e-10
TC225972 homologue to GB|AAB86804.1|2454184|ATU80186 pyruvate de... 59 3e-09
AW596798 similar to PIR|T51835|T518 3-methyl-2-oxobutanoate dehy... 57 1e-08
BU544660 similar to GP|3851001|gb|A pyruvate dehydrogenase E1 be... 57 2e-08
TC226476 homologue to UP|Q8L693 (Q8L693) 1-deoxy-D-xylulose 5-ph... 57 2e-08
TC230963 similar to UP|O64904 (O64904) 1-deoxyxylulose-5-phospha... 55 6e-08
TC226475 homologue to UP|Q8L693 (Q8L693) 1-deoxy-D-xylulose 5-ph... 48 6e-06
TC207143 similar to UP|Q8L692 (Q8L692) 1-deoxy-D-xylulose 5-phos... 43 2e-04
TC213419 similar to UP|Q9LFL9 (Q9LFL9) 1-D-deoxyxylulose 5-phosp... 33 0.15
TC229279 homologue to UP|Q9XH50 (Q9XH50) 1-D-deoxyxylulose 5-pho... 29 3.8
TC217344 28 8.5
>TC215667 similar to UP|ODPB_PEA (P52904) Pyruvate dehydrogenase E1 component
beta subunit, mitochondrial precursor (PDHE1-B) ,
complete
Length = 1464
Score = 658 bits (1698), Expect = 0.0
Identities = 333/362 (91%), Positives = 352/362 (96%)
Frame = +1
Query: 1 MLGVIRNKVNLLRPSFSAFRHLSSSAKQMTVRDALNSALDEEMSADPKVFLMGEEVGEYQ 60
MLGVIR+K +RP+FSA RH SS+AK++TVR+ALNSALDEEMSADPKVFLMGEEVGEYQ
Sbjct: 73 MLGVIRHKS--IRPAFSAIRHFSSAAKEITVREALNSALDEEMSADPKVFLMGEEVGEYQ 246
Query: 61 GAYKISKGLLEKYGPERVLDTPITEAGFTGIGVGAAYYGLKPVVEFMTFNFSMQAIDHII 120
GAYKISKGLL+KYGPERVLDTPITEAGF GIGVGAAYYGL+PVVEFMTFNFSMQAIDHII
Sbjct: 247 GAYKISKGLLDKYGPERVLDTPITEAGFAGIGVGAAYYGLRPVVEFMTFNFSMQAIDHII 426
Query: 121 NSAAKSNYMSAGQINVPIVFRGPNGAAAGVGAQHSHCYASWYGSCPGLKVLAPYSSEDAR 180
NSAAKSNYMSAGQI+VPIVFRGPNGAAAGVGAQHS CYAS YGSCPGLKVL+PYSSEDAR
Sbjct: 427 NSAAKSNYMSAGQISVPIVFRGPNGAAAGVGAQHSQCYASLYGSCPGLKVLSPYSSEDAR 606
Query: 181 GLLKAAIRDPDPVVFLENELLYGESFPVSAEVLDSSFCLPIGKAKIEREGKDVTITAFSK 240
GLLKAAIRDPDPVVFLENELLYGESFPVSAEVLDSSFCLPIGKAKIEREGKDVTITA+SK
Sbjct: 607 GLLKAAIRDPDPVVFLENELLYGESFPVSAEVLDSSFCLPIGKAKIEREGKDVTITAYSK 786
Query: 241 MVGFALKAAETLEKEGISAEVINLRSIRPLDRATINASVRKTNRLVTVEEGFPQHGVGAE 300
MVG+ALKAAETL KEGISAEVINLRSIRPLDR+TINASVRKTNRLVTVEEGFPQHGVGAE
Sbjct: 787 MVGYALKAAETLAKEGISAEVINLRSIRPLDRSTINASVRKTNRLVTVEEGFPQHGVGAE 966
Query: 301 ICASVIEESFGYLDAPVERIAGADVPMPYAANLERLAVPQIEDIVRAAKRACHRSVPMAA 360
IC SVIEESFGYLDAPVERIAGADVPMPYAANLER+AVPQ+EDIVRAAKRAC+RSVP+AA
Sbjct: 967 ICTSVIEESFGYLDAPVERIAGADVPMPYAANLERMAVPQVEDIVRAAKRACYRSVPLAA 1146
Query: 361 TA 362
+A
Sbjct: 1147SA 1152
>TC225970 homologue to GB|AAB86804.1|2454184|ATU80186 pyruvate dehydrogenase
E1 beta subunit {Arabidopsis thaliana;} , partial (80%)
Length = 1579
Score = 269 bits (688), Expect = 1e-72
Identities = 141/330 (42%), Positives = 206/330 (61%)
Frame = +3
Query: 23 SSSAKQMTVRDALNSALDEEMSADPKVFLMGEEVGEYQGAYKISKGLLEKYGPERVLDTP 82
S S ++ + +AL L+EEM DP V +MGE+VG Y G+YK++KGL K+G RVLDTP
Sbjct: 393 SKSGHELLLFEALREGLEEEMERDPCVCVMGEDVGHYGGSYKVTKGLATKFGDLRVLDTP 572
Query: 83 ITEAGFTGIGVGAAYYGLKPVVEFMTFNFSMQAIDHIINSAAKSNYMSAGQINVPIVFRG 142
I E FTG+G+GAA GL+PVVE M F + A + I N+ +Y S GQ +PIV RG
Sbjct: 573 IAENSFTGMGIGAAMTGLRPVVEGMNMGFLLLAFNQISNNCGMLHYTSGGQFKIPIVIRG 752
Query: 143 PNGAAAGVGAQHSHCYASWYGSCPGLKVLAPYSSEDARGLLKAAIRDPDPVVFLENELLY 202
P G +GA+HS S++ S PG++++A + +A+GL+KAAIR +PV+ E+ LLY
Sbjct: 753 PGGVGRQLGAEHSQRLESYFQSIPGIQMVACSTPYNAKGLMKAAIRSENPVILFEHVLLY 932
Query: 203 GESFPVSAEVLDSSFCLPIGKAKIEREGKDVTITAFSKMVGFALKAAETLEKEGISAEVI 262
+ + D + L + +A++ R G+ VTI +S+M ++AA+TL +G EVI
Sbjct: 933 N----LKERIPDEEYVLSLEEAEMVRPGEHVTILTYSRMRYHVMQAAKTLVNKGYDPEVI 1100
Query: 263 NLRSIRPLDRATINASVRKTNRLVTVEEGFPQHGVGAEICASVIEESFGYLDAPVERIAG 322
++RS++P D TI SV+KT+R++ VEE G+GA + A++ E YLDAP+ ++
Sbjct: 1101DIRSLKPFDLHTIGNSVKKTHRVLIVEECMRTGGIGASLTAAITENFHDYLDAPIVCLSS 1280
Query: 323 ADVPMPYAANLERLAVPQIEDIVRAAKRAC 352
DVP PYA LE V Q IV A ++ C
Sbjct: 1281QDVPTPYAGTLEEWTVVQPAQIVTAVEQLC 1370
>TC225973 homologue to GB|AAB86804.1|2454184|ATU80186 pyruvate dehydrogenase
E1 beta subunit {Arabidopsis thaliana;} , partial (58%)
Length = 983
Score = 166 bits (419), Expect = 2e-41
Identities = 104/281 (37%), Positives = 159/281 (56%)
Frame = +1
Query: 72 KYGPERVLDTPITEAGFTGIGVGAAYYGLKPVVEFMTFNFSMQAIDHIINSAAKSNYMSA 131
K+G RVLDTPI E FTG+G+GAA GL+PVVE ++ D + + +Y S
Sbjct: 4 KFGDLRVLDTPIAENSFTGMGIGAAMTGLRPVVEG*HGIPTLD*PD--LYNCGMLHYTSG 177
Query: 132 GQINVPIVFRGPNGAAAGVGAQHSHCYASWYGSCPGLKVLAPYSSEDARGLLKAAIRDPD 191
G F G A + S++ S PG++++A + +A+GL+KAAIR +
Sbjct: 178 GHSRYQSSFG--TW*VGGT*AD-TRTIESYFQSIPGIQMVACSTPYNAKGLMKAAIRSEN 348
Query: 192 PVVFLENELLYGESFPVSAEVLDSSFCLPIGKAKIEREGKDVTITAFSKMVGFALKAAET 251
PV+ E+ LLY + + D + L + +A++ R G+ VTI +S+M ++AA+T
Sbjct: 349 PVILFEHVLLYN----LKERIPDEEYVLSLEEAEMVRPGEHVTILTYSRMRYHVMQAAKT 516
Query: 252 LEKEGISAEVINLRSIRPLDRATINASVRKTNRLVTVEEGFPQHGVGAEICASVIEESFG 311
L +G EVI++RS++P D TI SV+KT+R++ VEE G+GA + A++ E
Sbjct: 517 LVNKGYDPEVIDIRSLKPFDLHTIGNSVKKTHRVLIVEECMRTGGIGASLTAAITENFHD 696
Query: 312 YLDAPVERIAGADVPMPYAANLERLAVPQIEDIVRAAKRAC 352
+LDAP+ ++ DVP PYA LE V Q IV A ++ C
Sbjct: 697 HLDAPIVCLSSQDVPTPYAGTLEEWTVVQPAQIVTAVEQLC 819
>TC225971 homologue to GB|AAB86804.1|2454184|ATU80186 pyruvate dehydrogenase
E1 beta subunit {Arabidopsis thaliana;} , partial (40%)
Length = 803
Score = 146 bits (368), Expect = 2e-35
Identities = 75/169 (44%), Positives = 104/169 (61%)
Frame = +3
Query: 15 SFSAFRHLSSSAKQMTVRDALNSALDEEMSADPKVFLMGEEVGEYQGAYKISKGLLEKYG 74
S +A S S ++ + +AL L+EEM DP V +MGE+VG Y G+YK++KGL K+G
Sbjct: 297 SSAASTSKSGSGHELLLFEALREGLEEEMERDPCVCVMGEDVGHYGGSYKVTKGLAPKFG 476
Query: 75 PERVLDTPITEAGFTGIGVGAAYYGLKPVVEFMTFNFSMQAIDHIINSAAKSNYMSAGQI 134
RVLDTPI E F G+G+GAA GL+PVVE M F + A + I N+ +Y S GQ
Sbjct: 477 DLRVLDTPIAENAFMGMGIGAAMTGLRPVVEGMNMGFLLLAFNQISNNCGMLHYTSGGQF 656
Query: 135 NVPIVFRGPNGAAAGVGAQHSHCYASWYGSCPGLKVLAPYSSEDARGLL 183
+PIV RGP G + A+HS S+Y S PG++++A +A GL+
Sbjct: 657 KIPIVIRGPGGVGRQLEAEHSQRLESYYQSIPGIQMVASSIPYNANGLM 803
>TC229089 similar to UP|O82450 (O82450) Branched-chain alpha-keto acid
decarboxylase E1 beta subunit , partial (59%)
Length = 982
Score = 140 bits (352), Expect = 1e-33
Identities = 80/207 (38%), Positives = 116/207 (55%)
Frame = +3
Query: 143 PNGAAAGVGAQHSHCYASWYGSCPGLKVLAPYSSEDARGLLKAAIRDPDPVVFLENELLY 202
P GA G HS +++ PG+KV+ P S A+GLL + IRDP+PVVF E + LY
Sbjct: 3 PYGAVGHGGHYHSQSPEAFFCHVPGIKVVIPRSPRQAKGLLLSCIRDPNPVVFFEPKWLY 182
Query: 203 GESFPVSAEVLDSSFCLPIGKAKIEREGKDVTITAFSKMVGFALKAAETLEKEGISAEVI 262
+ EV + + LP+ +A++ R+G D+T+ + + +A EKEGIS E+I
Sbjct: 183 RLA---DEEVPEDDYMLPLSEAEVIRQGSDITLVGWGAQLAIMEQACLDAEKEGISCELI 353
Query: 263 NLRSIRPLDRATINASVRKTNRLVTVEEGFPQHGVGAEICASVIEESFGYLDAPVERIAG 322
+L+++ P D+ T+ +SV KT RL+ E G GAEI AS++E F L+APV RI G
Sbjct: 354 DLKTLIPWDKETVESSVNKTGRLLVSHEAPITGGFGAEISASIVERCFSRLEAPVARICG 533
Query: 323 ADVPMPYAANLERLAVPQIEDIVRAAK 349
D P P E +P I+ A K
Sbjct: 534 LDTPFPLV--FEPFYMPTKNKILDAIK 608
>TC215668 homologue to UP|Q9ZQY1 (Q9ZQY1) Pyruvate dehydrogenase E1 beta
subunit isoform 3 , partial (14%)
Length = 479
Score = 98.2 bits (243), Expect = 5e-21
Identities = 46/52 (88%), Positives = 51/52 (97%)
Frame = +2
Query: 311 GYLDAPVERIAGADVPMPYAANLERLAVPQIEDIVRAAKRACHRSVPMAATA 362
GYLDAPVERIAGADVPMPYAANLER+AVPQ+EDIVRAAKR C+RSVP+AA+A
Sbjct: 8 GYLDAPVERIAGADVPMPYAANLERMAVPQVEDIVRAAKRTCYRSVPLAASA 163
>AW706828 weakly similar to SP|P52904|ODPB_ Pyruvate dehydrogenase E1
component beta subunit mitochondrial precursor (EC
1.2.4.1) (PDHE1-B)., partial (18%)
Length = 202
Score = 88.6 bits (218), Expect = 4e-18
Identities = 48/69 (69%), Positives = 59/69 (84%)
Frame = +2
Query: 2 LGVIRNKVNLLRPSFSAFRHLSSSAKQMTVRDALNSALDEEMSADPKVFLMGEEVGEYQG 61
LGVIR+K ++ R + SA RHLSS++K+MTVRDA++SALDEEMSADPK LMGEEVGE G
Sbjct: 2 LGVIRHK-SICR-ALSAIRHLSSASKEMTVRDAVSSALDEEMSADPKGVLMGEEVGEVSG 175
Query: 62 AYKISKGLL 70
AY++ KGLL
Sbjct: 176AYRMMKGLL 202
>TC207142 similar to UP|Q8L692 (Q8L692) 1-deoxy-D-xylulose 5-phosphate
synthase 2 precursor, partial (55%)
Length = 1176
Score = 63.2 bits (152), Expect = 2e-10
Identities = 72/300 (24%), Positives = 116/300 (38%), Gaps = 3/300 (1%)
Frame = +1
Query: 22 LSSSAKQMTVRDALNSALDEEMSADPKVFLMGEEVGEYQGAYKISKGLLEKYGPERVLDT 81
L + + ++ +L +E D K+ + +G G +K PER D
Sbjct: 283 LKAKSSTLSYTQYFAESLIKEAEIDNKIVAIHAAMGGGTGL-----NYFQKKFPERCFDV 447
Query: 82 PITEAGFTGIGVGAAYYGLKPVVEFMTFNFSMQAIDHIINSAAKSNYMSAGQINVPIVFR 141
I E G A GLKP + +F + D +++ +P+ F
Sbjct: 448 GIAEQHAVTFAAGLAAEGLKPFCAIYS-SFLQRGYDQVVHDVDLQK--------LPVRFA 600
Query: 142 GPNGAAAGV-GAQHSHCYASWYGSC-PGLKVLAPYSSEDARGLLK-AAIRDPDPVVFLEN 198
G G H + Y +C P + V+AP + ++ AA D P F
Sbjct: 601 LDRAGLVGADGPTHCGAFDITYMACLPNMVVMAPSDETELMHMVATAAAIDDRPSCF--- 771
Query: 199 ELLYGESFPVSAEVLDSSFCLPIGKAKIEREGKDVTITAFSKMVGFALKAAETLEKEGIS 258
G + + + L IGK +I EG V I + +V +A+ L++ GI
Sbjct: 772 RFPRGNGIGATLPLNNKGTPLEIGKGRILVEGSRVAILGYGSVVQQCRQAS*MLKELGID 951
Query: 259 AEVINLRSIRPLDRATINASVRKTNRLVTVEEGFPQHGVGAEICASVIEESFGYLDAPVE 318
V + R +PLD I ++ L+TVEEG G G+ + S G LD P++
Sbjct: 952 VTVADARFCKPLDTGLIRLLAKEHEILITVEEG-SIGGFGSHV--SQFLSLSGILDGPLK 1122
>TC225972 homologue to GB|AAB86804.1|2454184|ATU80186 pyruvate dehydrogenase
E1 beta subunit {Arabidopsis thaliana;} , partial (13%)
Length = 539
Score = 58.9 bits (141), Expect = 3e-09
Identities = 31/89 (34%), Positives = 49/89 (54%)
Frame = +1
Query: 4 VIRNKVNLLRPSFSAFRHLSSSAKQMTVRDALNSALDEEMSADPKVFLMGEEVGEYQGAY 63
++ N V S +A S S ++ + +AL L+EEM DP V +MGE+VG Y G+Y
Sbjct: 271 LVTNAVATKGASSAASTSKSGSGHELLLFEALREGLEEEMERDPCVCVMGEDVGHYGGSY 450
Query: 64 KISKGLLEKYGPERVLDTPITEAGFTGIG 92
K++KGL K+G + + + G+G
Sbjct: 451 KVTKGLAPKFGDLKFWTPLLXKTPLWGMG 537
>AW596798 similar to PIR|T51835|T518 3-methyl-2-oxobutanoate dehydrogenase
(lipoamide) (EC 1.2.4.4) beta chain [imported] -
Arabidopsis, partial (22%)
Length = 414
Score = 57.0 bits (136), Expect = 1e-08
Identities = 25/73 (34%), Positives = 44/73 (60%)
Frame = +1
Query: 27 KQMTVRDALNSALDEEMSADPKVFLMGEEVGEYQGAYKISKGLLEKYGPERVLDTPITEA 86
K + + A+N AL + DP+ ++ GE+V + G ++ + GL +++G +RV +TP+ E
Sbjct: 196 KSLNLCSAINQALHIALDTDPRSYVFGEDVS-FGGVFRCTTGLADQFGKKRVFNTPLCEQ 372
Query: 87 GFTGIGVGAAYYG 99
G G G+G A G
Sbjct: 373 GIVGFGIGLAAMG 411
>BU544660 similar to GP|3851001|gb|A pyruvate dehydrogenase E1 beta subunit
isoform 2 {Zea mays}, partial (8%)
Length = 553
Score = 56.6 bits (135), Expect = 2e-08
Identities = 26/33 (78%), Positives = 31/33 (93%)
Frame = -1
Query: 330 AANLERLAVPQIEDIVRAAKRACHRSVPMAATA 362
AANLER+AVPQ+ DIVRAAKR C+RSVP+AA+A
Sbjct: 553 AANLERMAVPQVXDIVRAAKRTCYRSVPLAASA 455
>TC226476 homologue to UP|Q8L693 (Q8L693) 1-deoxy-D-xylulose 5-phosphate
synthase 1 precursor, partial (85%)
Length = 2000
Score = 56.6 bits (135), Expect = 2e-08
Identities = 67/289 (23%), Positives = 104/289 (35%), Gaps = 3/289 (1%)
Frame = +3
Query: 20 RHLSSSAKQMTVRDALNSALDEEMSADPKVFLMGEEVGEYQGAYKISKGLLEKYGPERVL 79
+ S A + AL E AD V + +G G L + P R
Sbjct: 1131 KQFKSKATTRSYTTYFAEALIAEAEADKDVVAIHAAMGGGTGM-----NLFHRRFPTRCF 1295
Query: 80 DTPITEAGFTGIGVGAAYYGLKPVVEFMTFNFSMQAIDHIINSAAKSNYMSAGQINVPIV 139
D I E G A GLKP + +F +A D +++ +P+
Sbjct: 1296 DVGIAEQHAVTFAAGLACEGLKPFCAIYS-SFMQRAYDQVVHDVDLQK--------LPVR 1448
Query: 140 FRGPNGAAAGV-GAQHSHCYASWYGSC-PGLKVLAPYSSEDARGLLKAAIRDPD-PVVFL 196
F G G H + + +C P + V+AP + ++ A D P F
Sbjct: 1449 FAMDRAGLVGADGPTHCGSFDVTFMACLPNMVVMAPSDEAELFHMVATAAAINDRPSCFR 1628
Query: 197 ENELLYGESFPVSAEVLDSSFCLPIGKAKIEREGKDVTITAFSKMVGFALKAAETLEKEG 256
G V + L IGK +I EG+ V + + L A +E G
Sbjct: 1629 YPR---GNGIGVQLPTGNKGTPLEIGKGRILIEGERVALLGYGSAGQNCLPATSLVEHHG 1799
Query: 257 ISAEVINLRSIRPLDRATINASVRKTNRLVTVEEGFPQHGVGAEICASV 305
+ V + R +PLDR+ + + L+T EEG G G+ + S+
Sbjct: 1800 LRVTVADARFCKPLDRSLMRTLTKSHEVLITPEEG-SIGGFGSHVAXSL 1943
>TC230963 similar to UP|O64904 (O64904) 1-deoxyxylulose-5-phosphate synthase,
partial (26%)
Length = 947
Score = 54.7 bits (130), Expect = 6e-08
Identities = 27/73 (36%), Positives = 42/73 (56%)
Frame = +1
Query: 219 LPIGKAKIEREGKDVTITAFSKMVGFALKAAETLEKEGISAEVINLRSIRPLDRATINAS 278
L +GK ++ +EG V + + MV ++AA+ LE GIS V++ R +PLD +
Sbjct: 163 LEVGKGRVLKEGSRVALVGYGTMVQSCMEAAKVLEAHGISTTVVDARFCKPLDGDLMTRL 342
Query: 279 VRKTNRLVTVEEG 291
R+ L+TVEEG
Sbjct: 343 AREHEILITVEEG 381
>TC226475 homologue to UP|Q8L693 (Q8L693) 1-deoxy-D-xylulose 5-phosphate
synthase 1 precursor, partial (27%)
Length = 757
Score = 48.1 bits (113), Expect = 6e-06
Identities = 27/73 (36%), Positives = 39/73 (52%)
Frame = +3
Query: 219 LPIGKAKIEREGKDVTITAFSKMVGFALKAAETLEKEGISAEVINLRSIRPLDRATINAS 278
L IGK +I EG+ V + + V L AA +E G+ V + R +PLDR+ I +
Sbjct: 150 LEIGKGRILIEGERVALLGYGSAVQNCLAAASLVECHGLRLTVADARFCKPLDRSLIRSL 329
Query: 279 VRKTNRLVTVEEG 291
+ L+TVEEG
Sbjct: 330 AKSHEVLITVEEG 368
>TC207143 similar to UP|Q8L692 (Q8L692) 1-deoxy-D-xylulose 5-phosphate
synthase 2 precursor, partial (21%)
Length = 752
Score = 43.1 bits (100), Expect = 2e-04
Identities = 32/100 (32%), Positives = 46/100 (46%)
Frame = +1
Query: 219 LPIGKAKIEREGKDVTITAFSKMVGFALKAAETLEKEGISAEVINLRSIRPLDRATINAS 278
L IGK +I G V I + +V E L++ GI V + R +PLD I
Sbjct: 28 LEIGKGRILVXGSRVAILGYGSVVQQCRXXXEMLKELGIDVTVADARFCKPLDTGLIRLL 207
Query: 279 VRKTNRLVTVEEGFPQHGVGAEICASVIEESFGYLDAPVE 318
++ L+TVEEG G G+ + S G LD P++
Sbjct: 208 AKEHEILITVEEG-SIGGFGSHV--SQFLSLSGILDGPLK 318
>TC213419 similar to UP|Q9LFL9 (Q9LFL9) 1-D-deoxyxylulose 5-phosphate
synthase-like protein, partial (21%)
Length = 492
Score = 33.5 bits (75), Expect = 0.15
Identities = 20/54 (37%), Positives = 27/54 (49%)
Frame = +3
Query: 238 FSKMVGFALKAAETLEKEGISAEVINLRSIRPLDRATINASVRKTNRLVTVEEG 291
+ MV LKA L K G+ V + R +PLD + + + LVTVEEG
Sbjct: 72 YGSMVQNCLKAHSLLAKLGMEVTVADARFCKPLDIMLLRQLCKHHSFLVTVEEG 233
>TC229279 homologue to UP|Q9XH50 (Q9XH50) 1-D-deoxyxylulose 5-phosphate
synthase, partial (15%)
Length = 1074
Score = 28.9 bits (63), Expect = 3.8
Identities = 14/33 (42%), Positives = 20/33 (60%)
Frame = +1
Query: 259 AEVINLRSIRPLDRATINASVRKTNRLVTVEEG 291
A V + R +PLDR+ I + + L+TVEEG
Sbjct: 1 ATVADARFCKPLDRSLIRSLAQSHEVLITVEEG 99
>TC217344
Length = 518
Score = 27.7 bits (60), Expect = 8.5
Identities = 16/80 (20%), Positives = 38/80 (47%), Gaps = 1/80 (1%)
Frame = +1
Query: 265 RSIRPLDRATINASVRKTNR-LVTVEEGFPQHGVGAEICASVIEESFGYLDAPVERIAGA 323
R + ++ ++IN +R + +++ + GVGAE+ ++ + YL+ ++ +
Sbjct: 274 RRLGLINHSSINTQIRPSRHPIISFQSEVEDSGVGAELLDIILPKENCYLERSGGQVVAS 453
Query: 324 DVPMPYAANLERLAVPQIED 343
P + R + P I+D
Sbjct: 454 SPPFFCGSPPSRASNPLIQD 513
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.318 0.134 0.383
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,949,232
Number of Sequences: 63676
Number of extensions: 153424
Number of successful extensions: 761
Number of sequences better than 10.0: 36
Number of HSP's better than 10.0 without gapping: 755
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 756
length of query: 362
length of database: 12,639,632
effective HSP length: 98
effective length of query: 264
effective length of database: 6,399,384
effective search space: 1689437376
effective search space used: 1689437376
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 59 (27.3 bits)
Medicago: description of AC146972.11


