

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146972.10 - phase: 0
(106 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
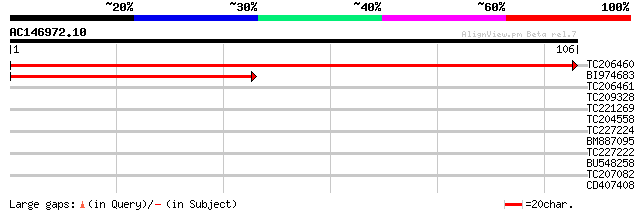
Sequences producing significant alignments: (bits) Value
TC206460 homologue to UP|T2AG_ARATH (Q39236) Transcription initi... 209 2e-55
BI974683 homologue to SP|Q39236|T2AG_ Transcription initiation f... 92 4e-20
TC206461 homologue to UP|T2AG_ARATH (Q39236) Transcription initi... 30 0.006
TC209328 similar to UP|KAPS_CATRO (O49204) Adenylyl-sulfate kina... 26 2.6
TC221269 similar to UP|Q7RHR7 (Q7RHR7) Retinitis pigmentosa GTPa... 26 3.4
TC204558 similar to UP|Q76KW5 (Q76KW5) Class1 chitinase (Fragmen... 26 3.4
TC227224 similar to UP|Q84W10 (Q84W10) At5g22480, partial (20%) 25 4.4
BM887095 25 5.8
TC227222 similar to UP|Q84W10 (Q84W10) At5g22480, partial (17%) 25 7.6
BU548258 24 9.9
TC207082 24 9.9
CD407408 similar to GP|8778633|gb| F5O11.22 {Arabidopsis thalian... 24 9.9
>TC206460 homologue to UP|T2AG_ARATH (Q39236) Transcription initiation factor
IIA gamma chain (TFIIA-gamma), complete
Length = 815
Score = 209 bits (532), Expect = 2e-55
Identities = 103/106 (97%), Positives = 106/106 (99%)
Frame = +3
Query: 1 MATFELYRRSTIGMCLTETLDEMVQNGTLSPEIAIQVLVQFDKSMTEALETQVKSKVSIK 60
MATFELYRRSTIGMCLTETLDEMVQNGTLSPE+AIQVLVQFDKSMTEALETQVKSKVSIK
Sbjct: 144 MATFELYRRSTIGMCLTETLDEMVQNGTLSPELAIQVLVQFDKSMTEALETQVKSKVSIK 323
Query: 61 GHLHTYRFCDNVWTFILQDALFKNEDNQENVGRVKIVACDSKLLSQ 106
GHLHTYRFCDNVWTFILQDALFKNED+QENVGRVKIVACDSKLL+Q
Sbjct: 324 GHLHTYRFCDNVWTFILQDALFKNEDSQENVGRVKIVACDSKLLTQ 461
>BI974683 homologue to SP|Q39236|T2AG_ Transcription initiation factor IIA
gamma chain (TFIIA-gamma). [Mouse-ear cress], partial
(43%)
Length = 426
Score = 92.0 bits (227), Expect = 4e-20
Identities = 45/46 (97%), Positives = 46/46 (99%)
Frame = +1
Query: 1 MATFELYRRSTIGMCLTETLDEMVQNGTLSPEIAIQVLVQFDKSMT 46
MATFELYRRSTIGMCLTETLDEMVQNGTLSPE+AIQVLVQFDKSMT
Sbjct: 289 MATFELYRRSTIGMCLTETLDEMVQNGTLSPELAIQVLVQFDKSMT 426
>TC206461 homologue to UP|T2AG_ARATH (Q39236) Transcription initiation factor
IIA gamma chain (TFIIA-gamma), partial (30%)
Length = 228
Score = 29.6 bits (65), Expect(2) = 0.006
Identities = 13/13 (100%), Positives = 13/13 (100%)
Frame = +1
Query: 1 MATFELYRRSTIG 13
MATFELYRRSTIG
Sbjct: 130 MATFELYRRSTIG 168
Score = 24.3 bits (51), Expect(2) = 0.006
Identities = 11/18 (61%), Positives = 11/18 (61%)
Frame = +2
Query: 14 MCLTETLDEMVQNGTLSP 31
MCLTETLDE V P
Sbjct: 170 MCLTETLDENVSERHSKP 223
>TC209328 similar to UP|KAPS_CATRO (O49204) Adenylyl-sulfate kinase,
chloroplast precursor (APS kinase)
(Adenosine-5'phosphosulfate kinase) (ATP
adenosine-5'-phosphosulfate 3'-phosphotransferase) ,
partial (69%)
Length = 1056
Score = 26.2 bits (56), Expect = 2.6
Identities = 15/39 (38%), Positives = 20/39 (50%)
Frame = +1
Query: 60 KGHLHTYRFCDNVWTFILQDALFKNEDNQENVGRVKIVA 98
KG L DN+ + QD F+ ED EN+ R+ VA
Sbjct: 331 KGKLSYILDGDNIRHGLNQDLSFRAEDRSENIRRIGEVA 447
>TC221269 similar to UP|Q7RHR7 (Q7RHR7) Retinitis pigmentosa GTPase
regulator-like protein, partial (3%)
Length = 802
Score = 25.8 bits (55), Expect = 3.4
Identities = 14/38 (36%), Positives = 21/38 (54%)
Frame = +1
Query: 2 ATFELYRRSTIGMCLTETLDEMVQNGTLSPEIAIQVLV 39
ATF + + + L ++LD+MV EI IQ+LV
Sbjct: 427 ATFPFCNLAFLSLDLFQSLDQMVSMALSISEIFIQILV 540
>TC204558 similar to UP|Q76KW5 (Q76KW5) Class1 chitinase (Fragment), partial
(97%)
Length = 1428
Score = 25.8 bits (55), Expect = 3.4
Identities = 12/49 (24%), Positives = 24/49 (48%)
Frame = +3
Query: 25 QNGTLSPEIAIQVLVQFDKSMTEALETQVKSKVSIKGHLHTYRFCDNVW 73
+N ++P A+ ++V S+ + K +K H+H ++CD W
Sbjct: 195 RNAVVAPIAALLLVVLLAASVVVEAQENSDVKTLVK-HMHGKKYCDKGW 338
>TC227224 similar to UP|Q84W10 (Q84W10) At5g22480, partial (20%)
Length = 588
Score = 25.4 bits (54), Expect = 4.4
Identities = 13/36 (36%), Positives = 19/36 (52%), Gaps = 1/36 (2%)
Frame = +3
Query: 47 EALETQVKSK-VSIKGHLHTYRFCDNVWTFILQDAL 81
++L+ Q K K + K L+ + WT IL DAL
Sbjct: 33 DSLDEQRKGKWIDFKARLNKLLSLEEAWTLILDDAL 140
>BM887095
Length = 400
Score = 25.0 bits (53), Expect = 5.8
Identities = 16/50 (32%), Positives = 27/50 (54%), Gaps = 1/50 (2%)
Frame = -2
Query: 49 LETQVKSKVSIK-GHLHTYRFCDNVWTFILQDALFKNEDNQENVGRVKIV 97
L+T+ SK+ HL+ F N W+ I+ A N ++N+G+V+ V
Sbjct: 216 LKTRSTSKLPYDYSHLNVTGFKKN*WSSIIVAA**GNSFKKKNIGKVQFV 67
>TC227222 similar to UP|Q84W10 (Q84W10) At5g22480, partial (17%)
Length = 706
Score = 24.6 bits (52), Expect = 7.6
Identities = 13/34 (38%), Positives = 18/34 (52%), Gaps = 1/34 (2%)
Frame = +1
Query: 49 LETQVKSK-VSIKGHLHTYRFCDNVWTFILQDAL 81
L+ Q K+K + K L+ + WT IL DAL
Sbjct: 1 LDEQRKNKWIDFKARLNKLLSLEEAWTLILDDAL 102
>BU548258
Length = 628
Score = 24.3 bits (51), Expect = 9.9
Identities = 11/45 (24%), Positives = 22/45 (48%)
Frame = +2
Query: 53 VKSKVSIKGHLHTYRFCDNVWTFILQDALFKNEDNQENVGRVKIV 97
VKSK+ I + RFC + F++ + D + ++ K++
Sbjct: 11 VKSKIVIANSIIRNRFCTFIHCFLID---YDKNDEEGHISETKLI 136
>TC207082
Length = 642
Score = 24.3 bits (51), Expect = 9.9
Identities = 12/46 (26%), Positives = 24/46 (52%)
Frame = -3
Query: 22 EMVQNGTLSPEIAIQVLVQFDKSMTEALETQVKSKVSIKGHLHTYR 67
E V+ G + + + ++FD S++ ++ K I+ +HTYR
Sbjct: 637 ESVRIGYILQPVNLYTRLRFDLSISHLFTISLQMKQVIQYIIHTYR 500
>CD407408 similar to GP|8778633|gb| F5O11.22 {Arabidopsis thaliana}, partial
(4%)
Length = 412
Score = 24.3 bits (51), Expect = 9.9
Identities = 14/38 (36%), Positives = 20/38 (51%)
Frame = -1
Query: 26 NGTLSPEIAIQVLVQFDKSMTEALETQVKSKVSIKGHL 63
NGTLSPE +I + DK + L+ + S+ G L
Sbjct: 388 NGTLSPEESIPSMTTIDK-LRSQLDDAIASECPFCGDL 278
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.320 0.133 0.377
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,816,849
Number of Sequences: 63676
Number of extensions: 36614
Number of successful extensions: 188
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 188
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 188
length of query: 106
length of database: 12,639,632
effective HSP length: 82
effective length of query: 24
effective length of database: 7,418,200
effective search space: 178036800
effective search space used: 178036800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 51 (24.3 bits)
Medicago: description of AC146972.10


