

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146971.7 - phase: 0 /pseudo
(2360 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
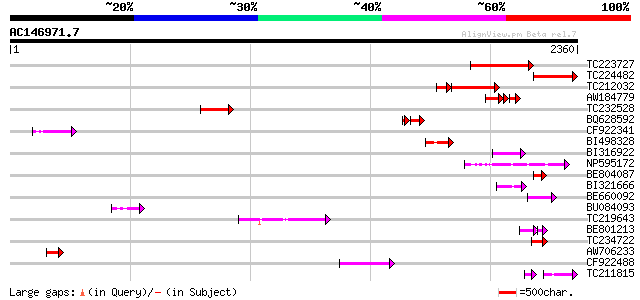
Score E
Sequences producing significant alignments: (bits) Value
TC223727 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 373 e-103
TC224482 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 285 1e-76
TC212032 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 219 2e-71
AW184779 111 2e-47
TC232528 weakly similar to UP|Q6WAY5 (Q6WAY5) Gag/pol polyprotei... 146 9e-35
BQ628592 111 1e-32
CF922341 125 2e-28
BI498328 119 1e-26
BI316922 103 9e-22
NP595172 polyprotein [Glycine max] 96 2e-19
BE804087 84 6e-16
BI321666 82 3e-15
BE660092 weakly similar to GP|9884624|dbj retroelement pol polyp... 77 1e-13
BU084093 72 2e-12
TC219643 weakly similar to UP|Q6WAY7 (Q6WAY7) Gag/pol polyprotei... 70 8e-12
BE801213 weakly similar to GP|6691193|gb| F7F22.17 {Arabidopsis ... 48 5e-11
TC234722 similar to UP|Q6WAY9 (Q6WAY9) Pol (Fragment), partial (... 64 8e-10
AW706233 63 2e-09
CF922488 62 3e-09
TC211815 45 5e-09
>TC223727 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (9%)
Length = 843
Score = 373 bits (958), Expect = e-103
Identities = 172/266 (64%), Positives = 209/266 (77%)
Frame = +1
Query: 1916 NHHNDVPLISVKFLDRPAYVFAAEVVFDDKPWFHDIKVFLQTREYPPGASNKDKKTLRRL 1975
N D+P I +PA+ E D KPW+ DIK ++ ++EY P ++ DK+TLRRL
Sbjct: 46 NAARDLPYIEFWCRGKPAHCCQVEEERDGKPWYFDIKRYVISKEYLPEIADNDKRTLRRL 225
Query: 1976 SSNFFLNGDILYKRNFDTVLLRCVDKYEADLLIHEIHEGSFGIHPNGHTMAKKILRAGYY 2035
++ FF++G ILYKRN D LRCVD EA+ +I E+HEGSFG H NGH MA+KILRAGYY
Sbjct: 226 AAGFFMSGSILYKRNHDMKPLRCVDAREANHMIEEVHEGSFGTHANGHAMARKILRAGYY 405
Query: 2036 WMTMESDCYKHTRKCHKCQIYADKIHMPPTTLNLLSSPWPFSMWGIDMIGRIEPKASNGH 2095
W+TMESDC H RKCHKCQ +AD ++ PP LN++SSPWPFSMWGID+IG IEPKASNGH
Sbjct: 406 WLTMESDCCVHVRKCHKCQAFADNVNAPPHPLNVMSSPWPFSMWGIDVIGAIEPKASNGH 585
Query: 2096 RFILVAIDYFTKWVEAASYANVTKQVVVKFIKNHIICRYGIPNRIITDNGTNLNNKMMKE 2155
RFILVAIDYFTKWVEAASY +V + VVV+FIK IICRYG+P +IITDNGTNLNNKMM E
Sbjct: 586 RFILVAIDYFTKWVEAASYTDVMRGVVVRFIKKEIICRYGLPRKIITDNGTNLNNKMMGE 765
Query: 2156 LCDDFKIEHHNSSPYRPQMNGAVEAA 2181
+C++FKI+HHN +PYRP+MN AVE A
Sbjct: 766 ICEEFKIQHHNPTPYRPKMN*AVEVA 843
>TC224482 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (6%)
Length = 669
Score = 285 bits (730), Expect = 1e-76
Identities = 136/182 (74%), Positives = 155/182 (84%)
Frame = +1
Query: 2179 EAANKNIKRIVQKMVVTYKDWHEMLPFALHGYRTSVRTSIGATPFSLVYGMEAVLPVEVE 2238
EAANKNIK+I+QKM V+YKDWHEMLPFALHGYRTSVRTS GATPFSLVYGMEAVLP EVE
Sbjct: 1 EAANKNIKKIIQKMTVSYKDWHEMLPFALHGYRTSVRTSTGATPFSLVYGMEAVLPFEVE 180
Query: 2239 IPSLRVLMEVDLSEAEWVQNRYDQLNLIEEKRMAALCHGQLYQKRMKQAFDKKVRPREFK 2298
+PSLR+L E L E+EW Q RYDQLNLIE KR+ A+ HG+LYQ+RMK AFDKKV R+F
Sbjct: 181 VPSLRILAESGLKESEWAQTRYDQLNLIEGKRLTAMSHGRLYQQRMKSAFDKKVCLRKFH 360
Query: 2299 EGDLVLKKIFSFQPDSRGKWAPNYEGPYVVKKAFSGGAMTLQTMDGEELPRPVNTDTVKK 2358
EGDLVLKK+ D RGKWAPNYEGP+VVK+AFSGGA+ L MDGEELP P+N+D VK+
Sbjct: 361 EGDLVLKKMSHAVKDHRGKWAPNYEGPFVVKRAFSGGALVLTNMDGEELPSPMNSDVVKR 540
Query: 2359 YF 2360
Y+
Sbjct: 541 YY 546
>TC212032 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (3%)
Length = 803
Score = 219 bits (559), Expect(2) = 2e-71
Identities = 102/199 (51%), Positives = 142/199 (71%)
Frame = +2
Query: 1838 IEEAIDLRIKNIEIYGDSALVINQIKGKWETLHAGLIPYRDYARRLLTFFNKVELHHIPR 1897
++ AID +K +++YGDSALVI+Q++G+ ET LIPY+ Y + L FF+++ HH+
Sbjct: 206 VQAAIDSNVKLLKVYGDSALVIHQLRGECETRDPNLIPYQAYIKELAGFFDEISFHHVA* 385
Query: 1898 DENQMADALATLSSMIKVNHHNDVPLISVKFLDRPAYVFAAEVVFDDKPWFHDIKVFLQT 1957
+ENQMADALATL SM ++ H D+P I + RPA+ E D KPW+ DIK ++++
Sbjct: 386 EENQMADALATLVSMFQLTPHGDLPYIEFRCRGRPAHCCLVEEERDGKPWYFDIKRYVES 565
Query: 1958 REYPPGASNKDKKTLRRLSSNFFLNGDILYKRNFDTVLLRCVDKYEADLLIHEIHEGSFG 2017
+EYP AS+ DK+ RRL++ FF++G ILYKRN D VLL CV+ E + ++ E+HEGSFG
Sbjct: 566 KEYPLEASDNDKRRKRRLAAGFFMSGSILYKRNHDMVLLHCVNGKEVENMLGEVHEGSFG 745
Query: 2018 IHPNGHTMAKKILRAGYYW 2036
H NGH MA+KILRAGYYW
Sbjct: 746 THSNGHAMARKILRAGYYW 802
Score = 70.5 bits (171), Expect(2) = 2e-71
Identities = 35/62 (56%), Positives = 43/62 (68%), Gaps = 1/62 (1%)
Frame = +1
Query: 1776 LFGEGPDPD-SVWGLIFDGAVNVYGNGIGAVLLTPKGTHIPFTARLRFDCTNNIAEYEAC 1834
LF E D D W + FD A NV G+G+GA+L++P IPFT RL FDCTNN+AEYEAC
Sbjct: 16 LFEEKLDEDRDKWIVWFDRASNVLGHGVGAILVSPDNQCIPFTTRLGFDCTNNMAEYEAC 195
Query: 1835 IM 1836
+
Sbjct: 196 AL 201
>AW184779
Length = 432
Score = 111 bits (277), Expect(3) = 2e-47
Identities = 50/74 (67%), Positives = 58/74 (77%)
Frame = +1
Query: 1981 LNGDILYKRNFDTVLLRCVDKYEADLLIHEIHEGSFGIHPNGHTMAKKILRAGYYWMTME 2040
L+ +ILYKRN D VLLRCVD EA+ ++ E+HEGSFG H N H MA+KILR GYYW+TME
Sbjct: 1 LSRNILYKRNHDMVLLRCVDAREAEQMLVEVHEGSFGTHANIHAMAQKILRVGYYWLTME 180
Query: 2041 SDCYKHTRKCHKCQ 2054
SDC H KCHKCQ
Sbjct: 181 SDCCIHVWKCHKCQ 222
Score = 85.5 bits (210), Expect(3) = 2e-47
Identities = 38/47 (80%), Positives = 43/47 (90%)
Frame = +3
Query: 2078 MWGIDMIGRIEPKASNGHRFILVAIDYFTKWVEAASYANVTKQVVVK 2124
MWGID+IG IEPKASNGH FILVAIDYFTKWVEA SYA+VT+ VV++
Sbjct: 291 MWGIDVIGAIEPKASNGHHFILVAIDYFTKWVEAVSYASVTRSVVIR 431
Score = 34.3 bits (77), Expect(3) = 2e-47
Identities = 11/23 (47%), Positives = 18/23 (77%)
Frame = +2
Query: 2054 QIYADKIHMPPTTLNLLSSPWPF 2076
Q +AD ++ PP LN+L++PWP+
Sbjct: 224 QTFADNVNAPPIPLNVLAAPWPY 292
>TC232528 weakly similar to UP|Q6WAY5 (Q6WAY5) Gag/pol polyprotein
(Fragment), partial (3%)
Length = 449
Score = 146 bits (369), Expect = 9e-35
Identities = 67/137 (48%), Positives = 104/137 (75%)
Frame = +1
Query: 795 SKISIMSLLLNSEAHKDALMKVLEQAFVDYDVTVGQFGGIVGNITACNNLSFSDEELPAE 854
+++S++ LL++SE H+ L+KVL +A V D++V FGG+V NITA N L+F++EE+PAE
Sbjct: 31 ARVSLLELLMSSEPHRALLVKVLNEAHVAQDISVEGFGGLVNNITANNYLAFAEEEIPAE 210
Query: 855 GRNHNRALHISVNCKTDALSKVLVDTGSSLNVLSKTTYTQLTYQDAPLRRSGVMVKAFDG 914
GR HN+ALH+SV C ++KVL+D G SLNV+ K+T +L + + L+ S ++V+AFDG
Sbjct: 211 GRGHNKALHVSVKCMDHIVAKVLIDNGYSLNVMPKSTLDKLPFNASHLKPSSMVVRAFDG 390
Query: 915 SRKDVLGEVVLPITVGP 931
+R++V GE+ LP+ +GP
Sbjct: 391 TRREVRGEIDLPVQIGP 441
>BQ628592
Length = 423
Score = 111 bits (277), Expect(2) = 1e-32
Identities = 48/61 (78%), Positives = 56/61 (91%)
Frame = -3
Query: 1667 YSMLEKTCCALAWAAKRLRHYMINHTTWLISKMDPIKYIFEKPALIGRIARWQMLLSEYD 1726
YSMLE+TCC L WA+ RLR YM++HTTWLISKMDP+KYIFEKPAL GRIARWQ+LLSE++
Sbjct: 184 YSMLERTCCTLVWASHRLRQYMLSHTTWLISKMDPVKYIFEKPALTGRIARWQVLLSEFN 5
Query: 1727 I 1727
I
Sbjct: 4 I 2
Score = 49.3 bits (116), Expect(2) = 1e-32
Identities = 21/30 (70%), Positives = 26/30 (86%)
Frame = -1
Query: 1633 ENSMGCVLGQQDETGRKEHAIYYLSKKFTE 1662
+ SMGCVL Q D++G+KE AIYYLSKKFT+
Sbjct: 285 DESMGCVLVQHDDSGKKEQAIYYLSKKFTD 196
>CF922341
Length = 675
Score = 125 bits (315), Expect = 2e-28
Identities = 73/185 (39%), Positives = 104/185 (55%), Gaps = 1/185 (0%)
Frame = +1
Query: 93 TCEAQMPTHQYTGQVPLPAMRVTPATMTYSAPVIHTIPQTEEPIFHSGNVEAYEEVSDLQ 152
TC Q G +PL T P H PQ + HS + ++++
Sbjct: 160 TCLTYATEGQAVGGIPL--------RNTLEGPQYH--PQLH--LLHSTTSKNPHVMAEMG 303
Query: 153 EKYDELRRDMKALRDKGKFG-KTAYDLCLVPSVQVPHKFKIPDFEKYKGSSCPEEHLKMY 211
K D L ++A+ + +L LVP++ P KFK+ DF+KYKG++CP+ HLKMY
Sbjct: 304 -KLDHLEEGLRAIEGGEDYAFANLEELFLVPNIITPPKFKVLDFDKYKGTTCPKNHLKMY 480
Query: 212 VRRMPAYAQDDQILIYYFQESLTGPASKWYTNLDKTRVQTFRDLCEAFVEQYSYNVDMTP 271
++M AYA+D+++LI+ FQESLTG A WYTNL+ +RV +++DL AFV QY YN DM
Sbjct: 481 CQKMGAYAKDEELLIHSFQESLTGVAVTWYTNLEPSRVHSWKDLMVAFVRQYQYNFDMVL 660
Query: 272 DRSDL 276
R L
Sbjct: 661 SRMQL 675
>BI498328
Length = 335
Score = 119 bits (299), Expect = 1e-26
Identities = 62/119 (52%), Positives = 81/119 (67%), Gaps = 2/119 (1%)
Frame = +1
Query: 1731 SQKAIKGSILADHLAHQPLEDYRPIKFDFPDEEIMYLKMKDCDEPLFGEGPDPDSV--WG 1788
+QKA+KGS LAD+LA PL+ YRP+ +FPDE+IM LF E + + W
Sbjct: 4 TQKAVKGSALADYLAQ*PLQGYRPMHPEFPDEDIM---------ALFEEKRTHEDINKWI 156
Query: 1789 LIFDGAVNVYGNGIGAVLLTPKGTHIPFTARLRFDCTNNIAEYEACIMGIEEAIDLRIK 1847
+ FDGA N G+G+GAVL++P IPFTARL FDCTNN+AEYEAC +G++ AID +K
Sbjct: 157 VCFDGASNALGHGVGAVLVSPDDQCIPFTARLGFDCTNNMAEYEACALGVQAAIDFDVK 333
>BI316922
Length = 405
Score = 103 bits (257), Expect = 9e-22
Identities = 49/135 (36%), Positives = 81/135 (59%)
Frame = +3
Query: 2010 EIHEGSFGIHPNGHTMAKKILRAGYYWMTMESDCYKHTRKCHKCQIYADKIHMPPTTLNL 2069
E+H G G+H M ++LR GYY TM C ++ +KC +C + + H+ L+
Sbjct: 3 EMHRGICGMHSKSQLMTTRVLRVGYY**TMRKYCTEYVKKCEEC*KFGNISHLLVEELHN 182
Query: 2070 LSSPWPFSMWGIDMIGRIEPKASNGHRFILVAIDYFTKWVEAASYANVTKQVVVKFIKNH 2129
+ +PWPF++ G+D++ R P + +++LV ID FTKW+E A ++ V KF+ +
Sbjct: 183 IVAPWPFAI*GVDIL-RPFPLSKRQVKYLLVGIDQFTKWIETEHIAIISIANVRKFV*RN 359
Query: 2130 IICRYGIPNRIITDN 2144
I+C +GIPN +I+DN
Sbjct: 360 IVC*FGIPNTLISDN 404
>NP595172 polyprotein [Glycine max]
Length = 4659
Score = 95.5 bits (236), Expect = 2e-19
Identities = 113/451 (25%), Positives = 188/451 (41%), Gaps = 13/451 (2%)
Frame = +1
Query: 1891 ELHHIPRDENQMADALATLSSMIKVNHHNDVPLISVKFLDRPAYVFAAEVVFDDKPWFHD 1950
++ + P +NQ ADAL+ + + H+ FL+ A ++ D
Sbjct: 2965 KIEYKPGKDNQAADALSRMFMLAWSEPHSI-------FLEE----LRARLISDPH----- 3096
Query: 1951 IKVFLQTREYPPGASNKDKKTLRRLSSNFFLNGDILYKRNFDTVLLRCVDKYEADLLIHE 2010
+K ++T Y GA +S++ + +LY + D V++ ++ + ++ E
Sbjct: 3097 LKQLMET--YKQGAD----------ASHYTVREGLLYWK--DRVVIPAEEEI-VNKILQE 3231
Query: 2011 IHEGSFGIHPNGHTMAKKILRAGYYWMTMESDCYKHTRKCHKCQIYADKIHMPPTTLNLL 2070
H G H G T L+A +YW M+ D + +KC CQ +P L L
Sbjct: 3232 YHSSPIGGHA-GITRTLARLKAQFYWPKMQEDVKAYIQKCLICQQAKSNNTLPAGLLQPL 3408
Query: 2071 SSPWPFSMW---GIDMIGRIEPKASNGHRFILVAIDYFTKWVEAASY-ANVTKQVVVKFI 2126
P P +W +D I + S G I+V ID TK+ A+ +VV +
Sbjct: 3409 --PIPQQVWEDVAMDFITGLPN--SFGLSVIMVVIDRLTKYAHFIPLKADYNSKVVAEAF 3576
Query: 2127 KNHIICRYGIPNRIITDNGTNLNNKMMKELCDDFKIEHHN---SSPYRPQMNGAVEAANK 2183
+HI+ +GIP I++D + + L FK++ SS Y PQ +G E NK
Sbjct: 3577 MSHIVKLHGIPRSIVSDRDRVFTSTFWQHL---FKLQGTTLAMSSAYHPQSDGQSEVLNK 3747
Query: 2184 NIKRIVQKMVVTY-KDWHEMLPFALHGYRTSVRTSIGATPFSLVYGMEAVLPVEVEIPSL 2242
++ ++ + K W + LP+A Y T+ S+G TPF +YG E P
Sbjct: 3748 CLEMYLRCFTYEHPKGWVKALPWAEFWYNTAYHMSLGMTPFRALYGRE---------PPT 3900
Query: 2243 RVLMEVDLSEAEWVQNRYDQLNLIEEKRMAALCHGQLYQKRMKQAFDKKVRPREFKEGDL 2302
+ + V+ + + + K L Q+ MK+ DKK F+ GD
Sbjct: 3901 LTRQACSIDDPAEVREQLTDRDALLAKLKINLTRA---QQVMKRQADKKRLDVSFQIGDE 4071
Query: 2303 VLKKIFSFQPDS-----RGKWAPNYEGPYVV 2328
VL K+ ++ S K + Y GP+ V
Sbjct: 4072 VLVKLQPYRQHSAVLRKNQKLSMRYFGPFKV 4164
Score = 33.5 bits (75), Expect = 1.1
Identities = 24/100 (24%), Positives = 42/100 (42%), Gaps = 2/100 (2%)
Frame = +1
Query: 1636 MGCVLGQQDETGRKEHAIYYLSKKFTECESRYSMLEKTCCALAWAAKRLRHYMINHTTWL 1695
+G VLGQ H I Y SKK + S + A+ A + RHY++ + +
Sbjct: 2707 VGAVLGQNG------HPIAYFSKKLAPRMQKQSAYTRELLAITEALSKFRHYLLGNKFII 2868
Query: 1696 ISKMDPIKYIFEKPALIGRIARW--QMLLSEYDIEYRSQK 1733
+ +K + ++ W + L ++ IEY+ K
Sbjct: 2869 RTDQRSLKSLMDQSLQTPEQQAWLHKFLGYDFKIEYKPGK 2988
>BE804087
Length = 160
Score = 84.3 bits (207), Expect = 6e-16
Identities = 40/53 (75%), Positives = 44/53 (82%)
Frame = +2
Query: 2179 EAANKNIKRIVQKMVVTYKDWHEMLPFALHGYRTSVRTSIGATPFSLVYGMEA 2231
EAANKNIK+ + KM V+YKDWHEM FALH YRT VRTS GATP+SLVYG EA
Sbjct: 2 EAANKNIKKNI*KMTVSYKDWHEMFSFALHMYRTLVRTSTGATPYSLVYGKEA 160
>BI321666
Length = 430
Score = 82.0 bits (201), Expect = 3e-15
Identities = 41/126 (32%), Positives = 67/126 (52%)
Frame = +2
Query: 2025 MAKKILRAGYYWMTMESDCYKHTRKCHKCQIYADKIHMPPTTLNLLSSPWPFSMWGIDMI 2084
++ +L++ +Y ++ D Y H + C+KCQ L+ + F WGID +
Sbjct: 2 ISTNVLQSRFYLPSIFKDAYVHAQSCNKCQRTRSVSKRNELPLHTILEVEIFDYWGIDFV 181
Query: 2085 GRIEPKASNGHRFILVAIDYFTKWVEAASYANVTKQVVVKFIKNHIICRYGIPNRIITDN 2144
G P SN +ILV +DY +KWVEA + ++V+KF+K I R G+P +I +
Sbjct: 182 GPFPPSFSN--EYILVVVDYVSKWVEAVACQKSDAKIVIKFLKKQIFSRLGVPWVLIDNG 355
Query: 2145 GTNLNN 2150
G++L N
Sbjct: 356 GSHLCN 373
>BE660092 weakly similar to GP|9884624|dbj retroelement pol polyprotein-like
{Arabidopsis thaliana}, partial (13%)
Length = 378
Score = 76.6 bits (187), Expect = 1e-13
Identities = 43/122 (35%), Positives = 69/122 (56%), Gaps = 1/122 (0%)
Frame = -3
Query: 2153 MKELCDDFKIEHHNSSPYRPQMNGAVEAANKNIKRIVQKMVV-TYKDWHEMLPFALHGYR 2211
M L + + H S+PY PQ NG E +N+ IKRI++K+V + KDW L AL +R
Sbjct: 376 MHALLKKYGVVHRVSTPYHPQTNGQAEISNREIKRILEKIVQPSRKDWSTRLDDALWAHR 197
Query: 2212 TSVRTSIGATPFSLVYGMEAVLPVEVEIPSLRVLMEVDLSEAEWVQNRYDQLNLIEEKRM 2271
T+ + IG +P+ +V+G LPVE+E + + + S + + R QL+ ++E R+
Sbjct: 196 TAYKAPIGMSPYRVVFGKACHLPVEIEHKAYWAVKTCNFSMDQAGEERKLQLSELDEIRL 17
Query: 2272 AA 2273
A
Sbjct: 16 EA 11
>BU084093
Length = 421
Score = 72.4 bits (176), Expect = 2e-12
Identities = 40/137 (29%), Positives = 61/137 (44%)
Frame = -1
Query: 422 PQYPQIPQYPQFPQNPSPQNTQQQNFQQQPYQQFPYQQYPQQNFQQQPYQQRPQQPRPPR 481
P P+ P + PQ P+P T+ +NF +P+Q+
Sbjct: 346 PVQPKAPAQREAPQVPTPNTTRPVG-----------NSNTTRNFPPRPFQE--------- 227
Query: 482 MPINPIPVTYAELLPGLLKKNLVQTRTAPPIPEKLPSWYRLDQTCDFHEGGRGHNIETCY 541
P+P+TY +LLP L+ +L + P WY + TC +H G GH++E C
Sbjct: 226 --FTPLPMTYEDLLPSLIANHLAVVTPGRVLQPPFPKWYDPNATCKYHGGVPGHSVEKCL 53
Query: 542 AFKSTVQRLINDGKITF 558
A K VQ L++ G +TF
Sbjct: 52 ALKYKVQHLMDAGWLTF 2
>TC219643 weakly similar to UP|Q6WAY7 (Q6WAY7) Gag/pol polyprotein (Fragment),
partial (8%)
Length = 1320
Score = 70.5 bits (171), Expect = 8e-12
Identities = 112/412 (27%), Positives = 170/412 (41%), Gaps = 31/412 (7%)
Frame = +1
Query: 954 PWIHEAGAVTSTLHQKLKFVKNGKLVTVNGEEALLVSHLSSFSFI-GADDVEGTPFQGFT 1012
PWIH G V STLHQKLKFV G LV V+GEE +LVS SS ++ A++ T FQ F
Sbjct: 1 PWIHSVGVVPSTLHQKLKFVVEGHLVIVSGEEDILVSCPSSMPYVEAAEESLETAFQSFE 180
Query: 1013 IEDKNTKRNEASISSLKDAQKVI-----------------QAGGSTSWGKLIELPENKHR 1055
+ ++ + L DA ++ GG TS LI N+ +
Sbjct: 181 VVSISSVDSLFGQPCLSDAAVMMARVMLGNGYEPGMGLGKDNGGITS---LINTQGNRGK 351
Query: 1056 EGLGFFPSTG----LSTAKKGTFHSSGFIHAIIEDDPESVPRGFITPGVSSHNWVAV--- 1108
GLG+ P+ +K + SS + P + R FI+ G+ V
Sbjct: 352 YGLGYKPTQADMKRSIAGRKNSGQSSRWRQESEGSPPCHISRSFISAGLGDKGQVFAICE 531
Query: 1109 -DVPFVAHLSK*CICLVYAFLFKKKKNNILSPAPSESELI*GFLFQEIIINKIKTSFFSR 1167
DVP L + C ++ ++ + + + ++* F ++ + R
Sbjct: 532 DDVPSTLDLVRPCPPDFQLGNWRVEERSGI----YATSIM*TF----TVLKAL*LGLGFR 687
Query: 1168 CFVSFFLCFSEKW*YK-KPNMNFQKNLHVSSF*KS--SITCAD*KSTNPLNNITL*SLPT 1224
F+ LC Y + ++ L+ SF S AD ++ P I L T
Sbjct: 688 VFLLLRLCVF*F*IYNTRTFLHLFLRLYPFSFICMFISFETADPMTSPPKVLIPGTHLST 867
Query: 1225 LSSLCMRRRKK--RMKRSLTKSLDYSNKKRRPFSLMGTN*K*LTWAPKKTRKKSRLGHRS 1282
LS ++R+ K RM ++S + + + L+ * TW + K R
Sbjct: 868 LSKK*IKRKMKETRMWDFPQN*KEWSPMRTKKWGLIKKKQN**TWEVAVEKGK*R*AQVL 1047
Query: 1283 EQVLKSK**SSSKNMLMCLPGPTKICRV*TLILWYITYL*NLSVRRSNKS*E 1334
+ **S + L G TKIC V* L L+ YL* SV R +++*E
Sbjct: 1048 PHLSVKN**SC*ETTKTSLLGHTKICPV*VLTLYSTDYL*IPSVPR*SRN*E 1203
>BE801213 weakly similar to GP|6691193|gb| F7F22.17 {Arabidopsis thaliana},
partial (3%)
Length = 416
Score = 48.1 bits (113), Expect(2) = 5e-11
Identities = 22/81 (27%), Positives = 43/81 (52%)
Frame = +3
Query: 2120 QVVVKFIKNHIICRYGIPNRIITDNGTNLNNKMMKELCDDFKIEHHNSSPYRPQMNGAVE 2179
++V+KF+K +I R+G+P +I+D G++ + ++ + H + Y PQ NG +
Sbjct: 12 KIVIKFLKKNIFSRFGMPRILISDGGSHFYYSQLNKVLKHDSVRHKVETSYHPQTNGQAK 191
Query: 2180 AANKNIKRIVQKMVVTYKDWH 2200
+N + + +K K WH
Sbjct: 192 VSNIHKENS*EK-----KMWH 239
Score = 39.7 bits (91), Expect(2) = 5e-11
Identities = 19/42 (45%), Positives = 25/42 (59%)
Frame = +2
Query: 2197 KDWHEMLPFALHGYRTSVRTSIGATPFSLVYGMEAVLPVEVE 2238
KDW L AL +T+ +T IG TPF +VY LPVE++
Sbjct: 248 KDWSSKLEDALWACKTAKKTPIGLTPFQMVYRKACHLPVELK 373
>TC234722 similar to UP|Q6WAY9 (Q6WAY9) Pol (Fragment), partial (32%)
Length = 482
Score = 63.9 bits (154), Expect = 8e-10
Identities = 33/70 (47%), Positives = 43/70 (61%), Gaps = 1/70 (1%)
Frame = -3
Query: 2170 YRPQMNGAVEAANKNIKRIVQKMVVTY-KDWHEMLPFALHGYRTSVRTSIGATPFSLVYG 2228
Y PQ NG E +NK IKR+++ +VV+ KDW L A YR + +T IG +PF LVYG
Sbjct: 234 YHPQTNGQAEVSNKEIKRVLENIVVSSRKDWALKLDDAFWAYRIAFKTPIGLSPFQLVYG 55
Query: 2229 MEAVLPVEVE 2238
L VE+E
Sbjct: 54 KACHLSVELE 25
>AW706233
Length = 376
Score = 62.8 bits (151), Expect = 2e-09
Identities = 30/72 (41%), Positives = 47/72 (64%), Gaps = 1/72 (1%)
Frame = -3
Query: 153 EKYDELRRDMKALRDKGKFG-KTAYDLCLVPSVQVPHKFKIPDFEKYKGSSCPEEHLKMY 211
EK D L+ KA+ + +L LV ++ P KFK+ +F+KYKG++CP+ HLKMY
Sbjct: 347 EKLDHLKERFKAIEGGQDYAFANLEELFLVXNIISPPKFKVLNFDKYKGTTCPKNHLKMY 168
Query: 212 VRRMPAYAQDDQ 223
++M AYA+D++
Sbjct: 167 CQKMGAYAKDEK 132
>CF922488
Length = 741
Score = 62.0 bits (149), Expect = 3e-09
Identities = 71/227 (31%), Positives = 101/227 (44%)
Frame = +1
Query: 1374 RKMVRSECVLTTVI*TKLARKTTSLYLILTCWSTVLQSLKSSHSWMDSPVIIKSRWHPKI 1433
++M R EC T I*TK ++ + LY I T +S SWMD VI + R H +I
Sbjct: 10 KRMGRCECAWTIEI*TKPVQRISFLYRISTFSWITRPVFPNSPSWMDFQVITR*R*HQRI 189
Query: 1434 ERKHHLLHLGALSVTK*CRSV*SMPELLIRGV*LLFSTI*YIKRLKSMWMI**SSRLLKK 1493
++ L G S + CR * M G +S * +R +S WM * ++ ++
Sbjct: 190 WKRQLSLLYGEPSAIRLCRLG*RMLGQHTSGPWWHYSRT*CTRR*RSTWMT*S*NQERRR 369
Query: 1494 TMSSIYRRCSSA*GSTNFV*IPTNALLVSDPESF*VSLSARKASK*ILTKSKPSERCLLR 1553
SI C +T *IP + L +PES L AR+ + I T+ K S R
Sbjct: 370 NTLSICESCLGDYVNTG*D*IPQSVCLR*NPESCSTLLIAREE*RWIRTR*K*SLRWPSH 549
Query: 1554 EQKKK*GVFLDD*ITSPGSFLI*LQLVGRYSNYSAKNKVLCGPKIAR 1600
Q+ K V * TS S+ *L L +S A+ + G I +
Sbjct: 550 IQRSKSKVSWGG*TTS*DSYHS*LPLASLFSYCCARISLSNGTMIVK 690
>TC211815
Length = 704
Score = 45.4 bits (106), Expect(2) = 5e-09
Identities = 39/142 (27%), Positives = 67/142 (46%), Gaps = 2/142 (1%)
Frame = +2
Query: 2221 TPFSLVYGMEAVLPVEVEIPSL--RVLMEVDLSEAEWVQNRYDQLNLIEEKRMAALCHGQ 2278
TPF L+YG+ +LP+EV L EV EA + L+LI++ R +
Sbjct: 248 TPF*LIYGISVMLPIEVGEVFL*RHYFAEV*NKEALQI-----DLDLIKQVREDTVIMT* 412
Query: 2279 LYQKRMKQAFDKKVRPREFKEGDLVLKKIFSFQPDSRGKWAPNYEGPYVVKKAFSGGAMT 2338
+++RM + F+ K+ G + S + R K+ N EGP+ ++ GA
Sbjct: 413 AFKQRMTRCFNSKLPSTV*GRGPSMEGIQRSLEVLVRSKFTTN*EGPFKIRHNSKNGAYK 592
Query: 2339 LQTMDGEELPRPVNTDTVKKYF 2360
L+ + G+ + R N+ +K Y+
Sbjct: 593 LEELSGKVVLRIWNSMHLKVYY 658
Score = 35.4 bits (80), Expect(2) = 5e-09
Identities = 18/51 (35%), Positives = 26/51 (50%)
Frame = +3
Query: 2143 DNGTNLNNKMMKELCDDFKIEHHNSSPYRPQMNGAVEAANKNIKRIVQKMV 2193
DNG N+ + E I+H +S PQ N EAANK I ++K++
Sbjct: 3 DNGLQFTNRKLNEFPSGLNIKHRVTSVKHPQTNRRAEAANKVILGDLKKLL 155
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.327 0.140 0.434
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 105,654,890
Number of Sequences: 63676
Number of extensions: 1621135
Number of successful extensions: 16637
Number of sequences better than 10.0: 225
Number of HSP's better than 10.0 without gapping: 12884
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 15215
length of query: 2360
length of database: 12,639,632
effective HSP length: 113
effective length of query: 2247
effective length of database: 5,444,244
effective search space: 12233216268
effective search space used: 12233216268
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 67 (30.4 bits)
Medicago: description of AC146971.7


