

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
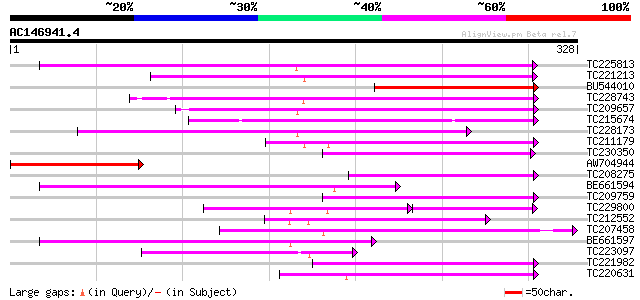
Query= AC146941.4 + phase: 0
(328 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC225813 149 2e-36
TC221213 weakly similar to UP|Q9C9F2 (Q9C9F2) MtN21-like protein... 139 1e-33
BU544010 similar to GP|22795261|gb| nodulin-like protein {Oryza ... 139 2e-33
TC228743 weakly similar to UP|Q9FL09 (Q9FL09) Nodulin-like prote... 115 3e-26
TC209657 weakly similar to UP|Q94JU2 (Q94JU2) AT3g28050/MMG15_6,... 115 4e-26
TC215674 weakly similar to UP|Q9SUD5 (Q9SUD5) Medicago nodulin N... 112 2e-25
TC228173 similar to UP|Q9SUF0 (Q9SUF0) Nodulin-like protein, par... 110 7e-25
TC211179 weakly similar to UP|Q9C9F2 (Q9C9F2) MtN21-like protein... 100 2e-21
TC230350 weakly similar to UP|Q9C9F2 (Q9C9F2) MtN21-like protein... 98 4e-21
AW704944 91 5e-19
TC208275 similar to UP|Q84TV1 (Q84TV1) Nodulin-like protein, par... 91 5e-19
BE661594 similar to GP|5262203|emb nodulin-like protein {Arabido... 89 4e-18
TC209759 88 5e-18
TC229800 similar to UP|O24091 (O24091) MtN21 protein, partial (53%) 53 2e-16
TC212552 similar to UP|Q9FGG3 (Q9FGG3) Nodulin-like protein, par... 80 1e-15
TC207458 similar to UP|Q9SUF0 (Q9SUF0) Nodulin-like protein, par... 79 3e-15
BE661597 similar to GP|5262203|emb nodulin-like protein {Arabido... 79 4e-15
TC223097 77 1e-14
TC221982 similar to UP|O81326 (O81326) F3D13.3 protein, partial ... 77 1e-14
TC220631 weakly similar to UP|O80638 (O80638) Nodulin-like prote... 76 2e-14
>TC225813
Length = 1551
Score = 149 bits (376), Expect = 2e-36
Identities = 86/301 (28%), Positives = 145/301 (47%), Gaps = 13/301 (4%)
Frame = +1
Query: 18 SMIFVQLIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAPLAIYFERGQPKNFSCEV 77
+M+ +Q G ++SR L G YR ++A + + P A + E+ + +
Sbjct: 157 AMLALQFGYAGFHVVSRAALNMGISKLVFPVYRNIIAFLLLLPFAYFLEKKERPAITLNF 336
Query: 78 LTKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMENLNIHTWN 137
L + FL VG++ Y G+ +TS T+A N +P TFL A++ R+E + ++ +
Sbjct: 337 LLQFFLLALVGITANQGFYLLGLDNTSPTFASAIQNSVPAITFLMAVILRIEQVRLNRKD 516
Query: 138 GRAKCVGAILCVAGTLAARLYKGKEFY------------IAQYHSFHSVA-AHKTQMLRG 184
G AK G I CVAG LYKG Y + ++ + S+ A G
Sbjct: 517 GIAKVAGTIFCVAGATVITLYKGPTIYSPSPPLQSESSVVVEFGTLSSLGDAKGKNWTLG 696
Query: 185 TLFLIGACFSYSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVDSSKEAWRLE 244
L+LIG C S+SAW +Q +++ +P R C IQ VI V+ +AW +
Sbjct: 697 CLYLIGHCLSWSAWLVLQAPVLKKYPARLSVTSYTCFFGLIQFLVIALIVERDAQAWIFQ 876
Query: 245 WNLQLITILYSGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAMILGEPLTV 304
++ TILY+G +++ F +Q W + GP + +++ P+ + VA ++ LGE +
Sbjct: 877 SGGEVFTILYAGVVASGIAFAVQIWCIDRGGPVFVAVYQPVQTLVVAIMASLALGEEFYL 1056
Query: 305 G 305
G
Sbjct: 1057G 1059
>TC221213 weakly similar to UP|Q9C9F2 (Q9C9F2) MtN21-like protein;
91922-89607, partial (37%)
Length = 941
Score = 139 bits (351), Expect = 1e-33
Identities = 77/232 (33%), Positives = 119/232 (51%), Gaps = 8/232 (3%)
Frame = +1
Query: 82 FLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMENLNIHTWNGRAK 141
FL G G S+ Y + TSAT+A NLIP TF+ A+ F ME LN+ T G+AK
Sbjct: 1 FLCGLFGGSLAQNFYLQALALTSATFASAMSNLIPGITFILAVCFGMERLNLRTAAGKAK 180
Query: 142 CVGAILCVAGTLAARLYKGKEFYIAQYH--------SFHSVAAHKTQMLRGTLFLIGACF 193
VG ++ + G + KG +H H+ +A L G+L + +
Sbjct: 181 IVGTLIGIGGAMVLTFVKGVHIEFGSFHLNLLHPQNGTHAHSATGAHTLLGSLCALASGI 360
Query: 194 SYSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVDSSKEAWRLEWNLQLITIL 253
SY+ W +Q K+ E +P Y L + ++ S V+ CV+ WRL WN++L+T
Sbjct: 361 SYALWLIIQAKMSESYPRPYSSTALMSLWGSLLSIVLALCVERDWSQWRLGWNIKLLTAA 540
Query: 254 YSGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAMILGEPLTVG 305
Y+G + + + + SW + ++GP + S+F+PL LV A A + IL E L +G
Sbjct: 541 YTGIVVSGEMVDVISWCVHMRGPLFASVFSPLMLVTEALAGSTILNEKLHLG 696
>BU544010 similar to GP|22795261|gb| nodulin-like protein {Oryza sativa
(japonica cultivar-group)}, partial (8%)
Length = 554
Score = 139 bits (349), Expect = 2e-33
Identities = 61/95 (64%), Positives = 80/95 (84%)
Frame = -3
Query: 212 RYWGIMLQCVMAAIQSAVIGACVDSSKEAWRLEWNLQLITILYSGALSTAAVFCLQSWAM 271
R+WG M+ C++AAIQ+A+IG +DSSK +WR EW+LQLITI+YSGAL+TAA FC SWA+
Sbjct: 552 RFWGTMIACILAAIQAAIIGXXLDSSKASWRXEWDLQLITIVYSGALATAASFCFLSWAI 373
Query: 272 TIKGPTYPSMFNPLALVFVAFAEAMILGEPLTVGT 306
IKGP+YP MFNPLAL+FVA +EA++ G+P+ V T
Sbjct: 372 KIKGPSYPPMFNPLALIFVAISEAIVXGQPIGVET 268
>TC228743 weakly similar to UP|Q9FL09 (Q9FL09) Nodulin-like protein, partial
(40%)
Length = 1383
Score = 115 bits (288), Expect = 3e-26
Identities = 73/244 (29%), Positives = 121/244 (48%), Gaps = 7/244 (2%)
Frame = +3
Query: 70 PKNFSCEVLTKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRME 129
P FS +L+KI L G +G S +L Y GIR +S T + NL P TF+ A++ RME
Sbjct: 15 PLTFS--ILSKIALLGVIGCSS-QILGYAGIRYSSPTLSSAISNLTPAFTFVLAVICRME 185
Query: 130 NLNIHTWNGRAKCVGAILCVAGTLAARLYKGKEFYIAQY-------HSFHSVAAHKTQML 182
+ + +AK +G+I+ V G YKG+ IA S + + +
Sbjct: 186 KIAVKRRTTQAKILGSIISVLGAFVVTFYKGQSVIIADNSPSIQLPQSNGILTSVDRNWV 365
Query: 183 RGTLFLIGACFSYSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVDSSKEAWR 242
G L L + WF QV++++ FP + + AAI ++++G + + AW+
Sbjct: 366 IGGLLLTACNILLTVWFVYQVEILKEFPDELSMVFFYNLCAAIVASIVGLLGEKNSSAWK 545
Query: 243 LEWNLQLITILYSGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAMILGEPL 302
+ ++ LI+I +G + + +W + +KGP Y +MF PL++V M LG+ L
Sbjct: 546 IRPDISLISIACTGIFNKFLSTAIYAWGIHLKGPVYVAMFKPLSIVIAVAMGVMFLGDSL 725
Query: 303 TVGT 306
VG+
Sbjct: 726 YVGS 737
>TC209657 weakly similar to UP|Q94JU2 (Q94JU2) AT3g28050/MMG15_6, partial
(31%)
Length = 1014
Score = 115 bits (287), Expect = 4e-26
Identities = 68/213 (31%), Positives = 105/213 (48%), Gaps = 3/213 (1%)
Frame = +1
Query: 97 YYGIRDTSATYALNFLNLIPICTFLTAILFRMENLNIHTWNGRAKCVGAILCVAGTLAAR 156
YYG SAT + + LNL+P TF+ A+LFRME L+ + AK +G I+ +AG
Sbjct: 10 YYG----SATLSTSILNLVPGFTFILAVLFRMEKLDWRKLSSLAKLLGTIVSIAGAFIVT 177
Query: 157 LYKGKEFYI---AQYHSFHSVAAHKTQMLRGTLFLIGACFSYSAWFFMQVKLVEVFPLRY 213
LYKG + + S + + + + LFL C SA+ +Q +++ +P
Sbjct: 178 LYKGPALLMGVSSANTSQQPLLSEDSNWILAGLFLAADCVMASAYIIVQASILKKYPAEL 357
Query: 214 WGIMLQCVMAAIQSAVIGACVDSSKEAWRLEWNLQLITILYSGALSTAAVFCLQSWAMTI 273
+ C AIQSAV V+ AW LE L+L+ +LYSG +A + W +
Sbjct: 358 IVVFFYCFFVAIQSAVTCLVVERDISAWSLEPKLRLLAVLYSGVFGSAFQVGIICWCLHQ 537
Query: 274 KGPTYPSMFNPLALVFVAFAEAMILGEPLTVGT 306
GP + SMF PL ++ + LG+ +G+
Sbjct: 538 TGPVFVSMFKPLGILISVVLGVLFLGDAFYLGS 636
>TC215674 weakly similar to UP|Q9SUD5 (Q9SUD5) Medicago nodulin N21-like
protein, partial (28%)
Length = 1577
Score = 112 bits (281), Expect = 2e-25
Identities = 68/204 (33%), Positives = 103/204 (50%), Gaps = 1/204 (0%)
Frame = +3
Query: 104 SATYALNFLNLIPICTFLTAILFRMENLNIHTWNGRAKCVGAILCVAGTLAARLYKGKEF 163
S+T A NLIP TF+ A + E ++I + AK +G + CVAG L L KG++
Sbjct: 513 SSTAATAMSNLIPALTFVIAAIAGFEKVDI-SLRSTAKILGTVCCVAGALTMALVKGQKL 689
Query: 164 YIAQY-HSFHSVAAHKTQMLRGTLFLIGACFSYSAWFFMQVKLVEVFPLRYWGIMLQCVM 222
++ S H + L G L L+ + +S W +QV + P C+
Sbjct: 690 LHTEFLPSIHLTGSQGDDWLLGCLLLLASSVFWSCWMILQVPITSCCPDHLLSTFWMCLF 869
Query: 223 AAIQSAVIGACVDSSKEAWRLEWNLQLITILYSGALSTAAVFCLQSWAMTIKGPTYPSMF 282
+ IQ+A+ +S +AW L+ LQ+ LY+G + A F +QSW ++ +GP Y +MF
Sbjct: 870 STIQAALFALLSESDLQAWILQSPLQISCSLYAG-IGIAVSFFIQSWCISERGPLYCAMF 1046
Query: 283 NPLALVFVAFAEAMILGEPLTVGT 306
NPLA V A A L E + VG+
Sbjct: 1047NPLATVITALISATFLEEEVYVGS 1118
>TC228173 similar to UP|Q9SUF0 (Q9SUF0) Nodulin-like protein, partial (54%)
Length = 904
Score = 110 bits (276), Expect = 7e-25
Identities = 61/238 (25%), Positives = 117/238 (48%), Gaps = 10/238 (4%)
Frame = +2
Query: 40 GTFIFALTSYRVLVAAVCVAPLAIYFERGQPKNFSCEVLTKIFLNGFVGMSMVMVLYYYG 99
G + L+ R AA+ + PLA+ ER + + +I GF+ + LY+ G
Sbjct: 191 GMGFWILSRXRHAAAAIIIVPLALVLERKIRPLMTLPIFLRIVALGFLEPVLDQNLYHMG 370
Query: 100 IRDTSATYALNFLNLIPICTFLTAILFRMENLNIHTWNGRAKCVGAILCVAGTLAARLYK 159
++ TS T+A +N++P TF+ A++FR+E +N+ ++ AK +G ++ V+G + LYK
Sbjct: 371 MKMTSTTFASATVNVLPAITFVMALIFRLEKVNLRKFHSVAKVIGTLITVSGAMVMTLYK 550
Query: 160 GKEFYI----------AQYHSFHSVAAHKTQMLRGTLFLIGACFSYSAWFFMQVKLVEVF 209
G F I + S + + GT++LI +C S++ +F +Q ++ +
Sbjct: 551 GPAFQIIKGGGAISNHSNSSSTSTTEPSDQHWIVGTVYLISSCASWAGFFILQSFTLKKY 730
Query: 210 PLRYWGIMLQCVMAAIQSAVIGACVDSSKEAWRLEWNLQLITILYSGALSTAAVFCLQ 267
P CVM I+ ++ + W + W+ +L+ +YSG + + + +Q
Sbjct: 731 PAELSLTAWICVMGIIEGSIASLIFERDFSVWAIGWDSRLLACVYSGVICSGMAYYVQ 904
>TC211179 weakly similar to UP|Q9C9F2 (Q9C9F2) MtN21-like protein;
91922-89607, partial (30%)
Length = 771
Score = 99.8 bits (247), Expect = 2e-21
Identities = 54/168 (32%), Positives = 86/168 (51%), Gaps = 10/168 (5%)
Frame = +2
Query: 149 VAGTLAARLYKGKEFYIAQYH-------SFHSVAAHKTQMLR---GTLFLIGACFSYSAW 198
++G + KG E + +H + H V H T L G L + + SY+ W
Sbjct: 11 ISGAMLLTFIKGPEVKMLSFHVNLFNHRNGHVVHPHATSGLMTIFGALASVASNVSYAMW 190
Query: 199 FFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVDSSKEAWRLEWNLQLITILYSGAL 258
+Q K+ E +P Y L +M A+ S CV+ WRL WN++L+T+ Y+G +
Sbjct: 191 LIIQAKMSERYPCPYSSTALMSLMGAVLSISFAFCVERDLSQWRLGWNIRLLTVAYAGIV 370
Query: 259 STAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAMILGEPLTVGT 306
+ + + SW + +GP + S+F+PL LV VAFA + IL E L +G+
Sbjct: 371 VSGVMVAVISWCVRTRGPLFVSIFSPLMLVVVAFAGSTILDEKLYLGS 514
>TC230350 weakly similar to UP|Q9C9F2 (Q9C9F2) MtN21-like protein;
91922-89607, partial (29%)
Length = 1057
Score = 98.2 bits (243), Expect = 4e-21
Identities = 42/123 (34%), Positives = 71/123 (57%)
Frame = +3
Query: 182 LRGTLFLIGACFSYSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVDSSKEAW 241
L G + + +CFS++ W +Q K+ + +P Y L AIQ+ G C + W
Sbjct: 72 LLGAICSLASCFSFALWLTIQAKMSKEYPCHYSSTALMSTAGAIQATAFGFCFERDLTQW 251
Query: 242 RLEWNLQLITILYSGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAMILGEP 301
+L WN++L+ + YSG +++ V + +W + ++GP + S+FNPL LV VA A +++L E
Sbjct: 252 KLGWNIRLLAVAYSGIVASGIVVIITAWCIQMRGPLFASVFNPLMLVLVAIASSLMLNEN 431
Query: 302 LTV 304
L V
Sbjct: 432 LYV 440
>AW704944
Length = 291
Score = 91.3 bits (225), Expect = 5e-19
Identities = 43/77 (55%), Positives = 58/77 (74%)
Frame = +2
Query: 1 MNFNDVKEWFVLSSILWSMIFVQLIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAP 60
+ N +K+WF S + SM+ VQL VTG Q+LSR+ILV+G+FI AL +YR +VAA+CVAP
Sbjct: 50 IRMNTLKKWFTSSQAILSMVLVQLFVTGLQLLSRVILVKGSFIGALITYRHIVAAICVAP 229
Query: 61 LAIYFERGQPKNFSCEV 77
A+YFERG K F+ +V
Sbjct: 230 FALYFERGLTKKFTWKV 280
>TC208275 similar to UP|Q84TV1 (Q84TV1) Nodulin-like protein, partial (34%)
Length = 732
Score = 91.3 bits (225), Expect = 5e-19
Identities = 45/110 (40%), Positives = 58/110 (51%)
Frame = +2
Query: 197 AWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVDSSKEAWRLEWNLQLITILYSG 256
AWF +Q + + FP Y L C MA+ Q +I CVD AW L ++L + LY+G
Sbjct: 2 AWFIIQKDISKTFPAPYTSTGLMCFMASFQCVIIAVCVDHRASAWSLHNAMRLSSALYAG 181
Query: 257 ALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAMILGEPLTVGT 306
T +CL SW + KGP Y S+F PL LV A +L E L VGT
Sbjct: 182 IFCTGLAYCLMSWTIERKGPLYVSVFTPLQLVLTAILSWALLREKLYVGT 331
>BE661594 similar to GP|5262203|emb nodulin-like protein {Arabidopsis
thaliana}, partial (41%)
Length = 786
Score = 88.6 bits (218), Expect = 4e-18
Identities = 58/215 (26%), Positives = 104/215 (47%), Gaps = 6/215 (2%)
Frame = +2
Query: 18 SMIFVQLIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAPLAIYFERGQPKNFSCEV 77
+M+ +Q +G I++ + G + L+ YR +VA + +AP A ER + V
Sbjct: 131 AMMSLQFGYSGMYIITMVSFKHGMSHWVLSVYRHIVATLIMAPFAFVLERKIRPKMTLPV 310
Query: 78 LTKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMENLNIHTWN 137
++ GF+ + LY G+++TS T+A +N++P TF+ A++ R+E +N+
Sbjct: 311 FLRLAALGFLEPVLDQNLYNMGMKNTSTTFASATVNVMPAITFIMALICRLETVNLRKIP 490
Query: 138 GRAKCVGAILCVAGTLAARLYKGKEF-YIAQYHSFHSVAAHKTQMLRGTL-----FLIGA 191
AK VG + V+G + LYKG +I + H + + TQ L LI
Sbjct: 491 SVAKVVGTAVTVSGAMVMTLYKGPALQFIKGQAATHHESGNSTQPX*TKLGAWHSRLIAX 670
Query: 192 CFSYSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQ 226
C ++++F +Q E P +W C +A ++
Sbjct: 671 CGGWASFFILQSFTFEKSPQNFWAQAWNCFLAFLR 775
>TC209759
Length = 899
Score = 88.2 bits (217), Expect = 5e-18
Identities = 44/125 (35%), Positives = 69/125 (55%)
Frame = +1
Query: 182 LRGTLFLIGACFSYSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVDSSKEAW 241
L G + +I + ++ WF +Q + + +P Y C+MA+IQ I + + AW
Sbjct: 121 LLGPVAVIVSALVWAVWFIVQKNMSKSYPAPYTSTFYMCLMASIQCVDIALAAEHNVSAW 300
Query: 242 RLEWNLQLITILYSGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAMILGEP 301
L ++L + LY+G +STA + L +W + KGP Y S+F+PL LV +A A +L E
Sbjct: 301 SLHSTIRLTSALYAGTISTALAYVLLAWTIERKGPLYVSVFSPLLLVIIAVASWALLHEQ 480
Query: 302 LTVGT 306
L VGT
Sbjct: 481 LYVGT 495
>TC229800 similar to UP|O24091 (O24091) MtN21 protein, partial (53%)
Length = 1303
Score = 52.8 bits (125), Expect(2) = 2e-16
Identities = 20/72 (27%), Positives = 43/72 (58%)
Frame = +2
Query: 234 VDSSKEAWRLEWNLQLITILYSGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFA 293
V+ + WR+ W++ L+ Y+G ++++ + +Q + +KGP + + F+PL ++ VA
Sbjct: 404 VEHNPSVWRIGWDVSLLAAAYAGIVTSSISYYVQGLVIKMKGPVFATAFSPLMMIIVAIM 583
Query: 294 EAMILGEPLTVG 305
+ IL E + +G
Sbjct: 584 GSFILAEQIYLG 619
Score = 50.4 bits (119), Expect(2) = 2e-16
Identities = 36/129 (27%), Positives = 62/129 (47%), Gaps = 8/129 (6%)
Frame = +1
Query: 113 NLIPICTFLTAILFRMENLNIHTWNGRAKCVGAILCVAGTLAARLYKGK--EFYIAQYHS 170
N++P TF+ A+ RME ++I AK VG ++ VAG + LY+G E A++
Sbjct: 16 NMLPAMTFVMAVFCRMEKIDIKKVRCIAKIVGTLVTVAGAMLMTLYRGPIVEMVWAKHPH 195
Query: 171 FHSVAAHKTQML-----RGTLFLIGACFSYSAWFFMQVKLVEVFPLRYWGI-MLQCVMAA 224
+ A T L G FLI A ++++ F +Q K ++ + + L C +
Sbjct: 196 NKTNATTTTGSLDKDWFLGCTFLIIATLAWASLFVLQAKAIQTYKNHQLSLTSLVCFIGT 375
Query: 225 IQSAVIGAC 233
+Q+ + C
Sbjct: 376 LQAIAVYFC 402
>TC212552 similar to UP|Q9FGG3 (Q9FGG3) Nodulin-like protein, partial (28%)
Length = 426
Score = 80.5 bits (197), Expect = 1e-15
Identities = 39/139 (28%), Positives = 69/139 (49%), Gaps = 8/139 (5%)
Frame = +3
Query: 148 CVAGTLAARLYKG------KEFYIAQYHSF--HSVAAHKTQMLRGTLFLIGACFSYSAWF 199
C+AG YKG F++ YH H A ++G ++ + + W
Sbjct: 3 CLAGAATFAFYKGPSLKFLSHFHLLDYHKSIQHQGHAQSGAWIKGCFLMLLSNTFFGLWL 182
Query: 200 FMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVDSSKEAWRLEWNLQLITILYSGALS 259
+Q +++ +P + +QC +++IQS VI V+ E W+L WN++L+ +LY G +
Sbjct: 183 VLQTFIIKRYPSKLLFTTIQCFLSSIQSFVIALAVERDIEQWKLGWNVRLLAVLYCGIMV 362
Query: 260 TAAVFCLQSWAMTIKGPTY 278
T + LQ+W + KGP +
Sbjct: 363 TGVSYYLQTWVIEKKGPVF 419
>TC207458 similar to UP|Q9SUF0 (Q9SUF0) Nodulin-like protein, partial (47%)
Length = 1006
Score = 79.0 bits (193), Expect = 3e-15
Identities = 48/213 (22%), Positives = 101/213 (46%), Gaps = 6/213 (2%)
Frame = +2
Query: 122 TAILFRMENLNIHTWNGRAKCVGAILCVAGTLAARLYKGKEF-YIAQYHSFHSVAAHKTQ 180
+A++ R+E +N+ + AK VG + V+G + LYKG +I + H + + TQ
Sbjct: 14 SALICRLETVNLRKIHSVAKVVGTAVTVSGAMVMTLYKGPALQFIKGQAATHHESGNSTQ 193
Query: 181 -----MLRGTLFLIGACFSYSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVD 235
+ GT+ LI +C ++++F +Q ++++P C + + A+ +
Sbjct: 194 PSEQNWVLGTVELIASCGGWASFFILQSFTLKMYPAELSVTAWICFLGIFEGAIATLIFE 373
Query: 236 SSKEAWRLEWNLQLITILYSGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEA 295
W + + +L+ +YSG + + + +Q +GP + + F+PL ++ A +
Sbjct: 374 RDMSVWSIGMDSRLLACVYSGVVCSGMAYYVQGVVTRERGPVFVTSFSPLCMIITAALGS 553
Query: 296 MILGEPLTVGTYEYFSFLPLIITIFISSLYCYV 328
++L E + +G+ + I +S LY V
Sbjct: 554 IVLAEQVYLGSV-------IGAIIIVSGLYTVV 631
>BE661597 similar to GP|5262203|emb nodulin-like protein {Arabidopsis
thaliana}, partial (44%)
Length = 849
Score = 78.6 bits (192), Expect = 4e-15
Identities = 53/197 (26%), Positives = 95/197 (47%), Gaps = 2/197 (1%)
Frame = +3
Query: 18 SMIFVQLIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAPLAIYFERGQPKNFSCEV 77
+M+ +Q +G I++ + G + L+ YR +VA + +AP A ER + V
Sbjct: 135 AMMSLQFGYSGMYIITMVSFKHGMSHWVLSVYRHIVATLIMAPFAFVLERKIRPKMTLPV 314
Query: 78 LTKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMENLNIHTWN 137
++ GF+ + LY G+++TS T+A +N++P TF+ A++ R+E +N+
Sbjct: 315 FLRLAALGFLEPVLDQNLYNMGMKNTSTTFASATVNVMPAITFIMALICRLETVNLRKIP 494
Query: 138 GRAKCVGAILCVAGTLAARLYKGK--EFYIAQYHSFHSVAAHKTQMLRGTLFLIGACFSY 195
AK VG + V+G + LYKG +F Q + H + TQ T L +
Sbjct: 495 SVAKVVGTAVTVSGAMGMTLYKGPALQFIKGQAATHH*KVGNSTQPF*QTGVLGTSRAHC 674
Query: 196 SAWFFMQVKLVEVFPLR 212
W+ +++ F L+
Sbjct: 675 KLWWVASFFILQSFTLK 725
>TC223097
Length = 651
Score = 77.0 bits (188), Expect = 1e-14
Identities = 46/134 (34%), Positives = 70/134 (51%), Gaps = 9/134 (6%)
Frame = +2
Query: 77 VLTKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMENLNIHTW 136
++ +I + G++ +LY+ G++ +SAT A NL+P TF+ A+LFR ENL I
Sbjct: 251 LMIQILFSSLTGVTGNQMLYFVGLKYSSATIACALTNLLPAFTFILAVLFRQENLGIKKR 430
Query: 137 NGRAKCVGAILCVAGTLAARLYKGKEFYIAQYHSFH---------SVAAHKTQMLRGTLF 187
G AK G ILCV+G L Y GK + Q S H + ++ K M G L
Sbjct: 431 AGLAKVFGTILCVSGALLLSFYHGKTIGLGQ-SSIHWRYAEKMEGTSSSGKGNMFLGPLV 607
Query: 188 LIGACFSYSAWFFM 201
+I + ++AWF +
Sbjct: 608 VILSTLVWAAWFII 649
>TC221982 similar to UP|O81326 (O81326) F3D13.3 protein, partial (5%)
Length = 679
Score = 76.6 bits (187), Expect = 1e-14
Identities = 36/131 (27%), Positives = 65/131 (49%)
Frame = +1
Query: 176 AHKTQMLRGTLFLIGACFSYSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVD 235
A + + G LF A S +AW Q +++ + + + C+ IQSA++ V
Sbjct: 13 AETSNWVIGGLFFATASISLAAWNITQAAILKGYSSQLTILAYYCLFGTIQSAILSLIVV 192
Query: 236 SSKEAWRLEWNLQLITILYSGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEA 295
W++ ++ LI + YS + + F + +W + KGP + S+F P+ + AF+
Sbjct: 193 RDPNDWKISPDIDLIAVFYSAVVGSVVTFSVNTWCIKKKGPVFVSLFKPVGIAIAAFSTV 372
Query: 296 MILGEPLTVGT 306
+ LGE L VG+
Sbjct: 373 VFLGETLHVGS 405
>TC220631 weakly similar to UP|O80638 (O80638) Nodulin-like protein
(At2g39510), partial (31%)
Length = 797
Score = 75.9 bits (185), Expect = 2e-14
Identities = 44/157 (28%), Positives = 77/157 (49%), Gaps = 7/157 (4%)
Frame = +1
Query: 157 LYKGKEFYIAQYHSFHSVAAHKTQMLRGTLFLIGACF------SYSAWFFMQVKLVEVFP 210
LYKG IA + H G+ +LIGACF +SA++ +Q + +P
Sbjct: 16 LYKGPVVNIAGSSASHVGQPENVNDPSGSHWLIGACFLLIGCAGFSAFYILQAITLRKYP 195
Query: 211 LRYWGIMLQCVMAAIQSAVIGACVD-SSKEAWRLEWNLQLITILYSGALSTAAVFCLQSW 269
C + A+QS+ + ++ +S + W L W+ +L+ YSG +++A F +Q
Sbjct: 196 AEMSLATWVCFVGALQSSAVSFFMERNSPDVWSLAWDSRLVAYAYSGIVTSAIQFYVQGM 375
Query: 270 AMTIKGPTYPSMFNPLALVFVAFAEAMILGEPLTVGT 306
+ GP + + FNPL ++ V ++L E L +G+
Sbjct: 376 VIKTTGPVFVTAFNPLRMIIVTALACIVLSEKLHLGS 486
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.332 0.141 0.447
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,091,154
Number of Sequences: 63676
Number of extensions: 253003
Number of successful extensions: 1795
Number of sequences better than 10.0: 94
Number of HSP's better than 10.0 without gapping: 1777
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1785
length of query: 328
length of database: 12,639,632
effective HSP length: 98
effective length of query: 230
effective length of database: 6,399,384
effective search space: 1471858320
effective search space used: 1471858320
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (22.0 bits)
S2: 59 (27.3 bits)
Medicago: description of AC146941.4


