

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146941.2 - phase: 0 /pseudo
(240 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
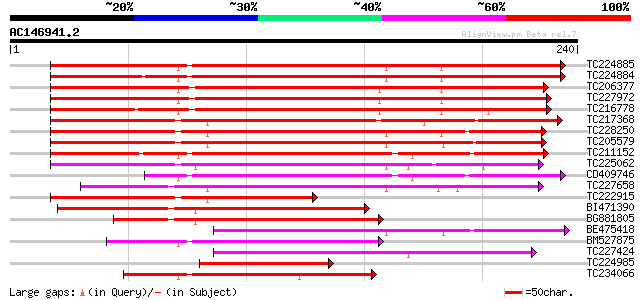
TC224885 similar to UP|Q93YI3 (Q93YI3) Calcium dependent calmodu... 251 2e-67
TC224884 similar to UP|Q93YI3 (Q93YI3) Calcium dependent calmodu... 239 8e-64
TC206377 similar to UP|O81390 (O81390) Calcium-dependent protein... 212 1e-55
TC227972 similar to UP|Q9AR92 (Q9AR92) Protein kinase , partial... 208 2e-54
TC216778 UP|O24431 (O24431) Calmodulin-like domain protein kinas... 200 5e-52
TC217368 similar to UP|Q93YF4 (Q93YF4) Calcium-dependent protein... 197 5e-51
TC228250 UP|Q84P29 (Q84P29) Seed calcium dependent protein kinas... 191 3e-49
TC205579 homologue to UP|Q43676 (Q43676) Calcium dependent prote... 185 1e-47
TC211152 UP|Q84P28 (Q84P28) Seed calcium dependent protein kinas... 184 2e-47
TC225062 similar to UP|Q8W4I7 (Q8W4I7) Calcium-dependent protein... 150 7e-37
CD409746 similar to PIR|T08873|T08 calcium-dependent protein kin... 131 3e-31
TC227658 similar to UP|O48565 (O48565) Calcium-dependent protein... 123 7e-29
TC222915 homologue to UP|Q84P29 (Q84P29) Seed calcium dependent ... 119 1e-27
BI471390 similar to GP|28416563|gb At1g18890 {Arabidopsis thalia... 113 7e-26
BG881805 94 4e-20
BE475418 similar to PIR|T06126|T06 calcium-dependent protein kin... 86 2e-17
BM527875 similar to PIR|T46189|T46 calcium-dependent protein kin... 83 1e-16
TC227424 UP|Q39890 (Q39890) Calmodulin, complete 77 5e-15
TC224985 homologue to UP|Q93YI3 (Q93YI3) Calcium dependent calmo... 77 5e-15
TC234066 similar to UP|Q8LDS1 (Q8LDS1) Calcium-dependent protein... 77 9e-15
>TC224885 similar to UP|Q93YI3 (Q93YI3) Calcium dependent calmodulin
independent protein kinase (Fragment), partial (92%)
Length = 1484
Score = 251 bits (641), Expect = 2e-67
Identities = 141/228 (61%), Positives = 166/228 (71%), Gaps = 10/228 (4%)
Frame = +2
Query: 18 WPSISPSAKDLVRKMLNSDPKQRISAYEVLNHPWIKEDGEAPDTPLDNAVLNR------- 70
WPSIS SAKDLV+KML +DPK+R+SA EVLNHPW++ DG+APD PLD AVL R
Sbjct: 497 WPSISSSAKDLVKKMLRADPKERLSAVEVLNHPWMRVDGDAPDKPLDIAVLTRMKQFRAM 676
Query: 71 NSLEQ*TDSRKLL*SCLSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQ 130
N L++ + K++ LSEEEI+GLK+MFK MDTDNSGTIT EELK GL K GT+LSE +
Sbjct: 677 NKLKK--VALKVIAENLSEEEIIGLKEMFKSMDTDNSGTITFEELKAGLPKLGTKLSESE 850
Query: 131 VKQLMEAADADGNGIIDYDEFITASQQL--CTRTD*TEKNMFILPSNTSIKIT-TELEQA 187
V+QLMEAAD DGNG IDY EFITA+ + R D K + S IT ELE A
Sbjct: 851 VRQLMEAADVDGNGTIDYIEFITATMHMNRMEREDHLYKAFEYFDKDKSGYITMEELESA 1030
Query: 188 LHEYNMHDGRYIKEIISEVDADNDGRINYDEFVAMMRKGNPEAHTKKR 235
L +YNM D + IKEII+EVDADNDGRINYDEFVAMMRKGNP+ +R
Sbjct: 1031LKKYNMGDEKTIKEIIAEVDADNDGRINYDEFVAMMRKGNPDLVNNRR 1174
>TC224884 similar to UP|Q93YI3 (Q93YI3) Calcium dependent calmodulin
independent protein kinase (Fragment), partial (82%)
Length = 1302
Score = 239 bits (610), Expect = 8e-64
Identities = 138/228 (60%), Positives = 163/228 (70%), Gaps = 10/228 (4%)
Frame = +1
Query: 18 WPSISPSAKDLVRKMLNSDPKQRISAYEVLNHPWIKEDGEAPDTPLDNAVLNR------- 70
WPSIS SAKDLV+KML +DPKQR+SA EVL+HPW++EDG A D PLD AVL+R
Sbjct: 343 WPSISSSAKDLVKKMLRADPKQRLSAVEVLDHPWMREDG-ASDKPLDVAVLSRMKQFRAM 519
Query: 71 NSLEQ*TDSRKLL*SCLSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQ 130
N L++ + K++ LSEEEI+GLK+MFK MDTDNSGTIT EELK GL K GT++SE +
Sbjct: 520 NKLKK--VALKVIAENLSEEEIIGLKEMFKSMDTDNSGTITFEELKAGLPKLGTKVSESE 693
Query: 131 VKQLMEAADADGNGIIDYDEFITASQQL--CTRTD*TEKNMFILPSNTSIKIT-TELEQA 187
V+QLMEAAD DGNG IDY EFITA+ + R D K + S IT ELE
Sbjct: 694 VRQLMEAADVDGNGTIDYIEFITATMHMNRMEREDHLYKAFEYFDKDRSGYITMEELEST 873
Query: 188 LHEYNMHDGRYIKEIISEVDADNDGRINYDEFVAMMRKGNPEAHTKKR 235
L +YNM D + IKEII EVD DNDGRINYDEFVAMMRKG P+ T +R
Sbjct: 874 LKKYNMGDEKTIKEIIVEVDTDNDGRINYDEFVAMMRKGKPDLVTNRR 1017
>TC206377 similar to UP|O81390 (O81390) Calcium-dependent protein kinase ,
partial (74%)
Length = 1644
Score = 212 bits (539), Expect = 1e-55
Identities = 124/221 (56%), Positives = 152/221 (68%), Gaps = 10/221 (4%)
Frame = +3
Query: 18 WPSISPSAKDLVRKMLNSDPKQRISAYEVLNHPWIKEDGEAPDTPLDNAVLNR------- 70
WPSIS SAKDLVRKML DP +RI++ +VL HPW++E G+A D P+D+AVL+R
Sbjct: 567 WPSISDSAKDLVRKMLTQDPNKRITSSQVLEHPWMREGGDASDKPIDSAVLSRMKQFRAM 746
Query: 71 NSLEQ*TDSRKLL*SCLSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQ 130
N L++ + K++ LSEEEI GLK MF MDTDNSGTIT EELK GL + G++LSE +
Sbjct: 747 NKLKKL--ALKVIAENLSEEEIKGLKAMFANMDTDNSGTITYEELKTGLARIGSKLSEAE 920
Query: 131 VKQLMEAADADGNGIIDYDEFITAS--QQLCTRTD*TEKNMFILPSNTSIKIT-TELEQA 187
VKQLM+AAD DGNG IDY EFI+A+ + R + K + S IT ELE A
Sbjct: 921 VKQLMDAADVDGNGSIDYLEFISATMHRHRLERDEHLYKAFQYFDKDNSGYITRDELEIA 1100
Query: 188 LHEYNMHDGRYIKEIISEVDADNDGRINYDEFVAMMRKGNP 228
+ + M D IKEIISEVDADNDGRINY+EF AMMR G P
Sbjct: 1101MTQNGMGDEATIKEIISEVDADNDGRINYEEFCAMMRSGMP 1223
>TC227972 similar to UP|Q9AR92 (Q9AR92) Protein kinase , partial (85%)
Length = 2010
Score = 208 bits (529), Expect = 2e-54
Identities = 121/222 (54%), Positives = 151/222 (67%), Gaps = 10/222 (4%)
Frame = +3
Query: 18 WPSISPSAKDLVRKMLNSDPKQRISAYEVLNHPWIKEDGEAPDTPLDNAVLNR------- 70
WPSIS SAKDLVRKML DPK+RI+A +VL HPW+KE G A D P+++AVL+R
Sbjct: 1077 WPSISNSAKDLVRKMLIKDPKKRITAAQVLEHPWLKEGGNASDKPINSAVLSRMKQFRAM 1256
Query: 71 NSLEQ*TDSRKLL*SCLSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQ 130
N L++ + K++ LSEEEI GLK MF +DTDNSGTIT EEL+ GL + G++L+E +
Sbjct: 1257 NKLKKL--ALKVIAENLSEEEIQGLKAMFTNIDTDNSGTITYEELRAGLQRLGSKLTETE 1430
Query: 131 VKQLMEAADADGNGIIDYDEFITAS--QQLCTRTD*TEKNMFILPSNTSIKIT-TELEQA 187
V+QLM+AAD DGNG IDY EFITA+ + R + K + S IT ELE A
Sbjct: 1431 VRQLMDAADVDGNGTIDYIEFITATMHRHRLERDEHLYKAFQYFDKDGSGYITRDELEIA 1610
Query: 188 LHEYNMHDGRYIKEIISEVDADNDGRINYDEFVAMMRKGNPE 229
+ EY M D I+EIISEVD DNDGRINY+EF MMR G +
Sbjct: 1611 MKEYGMGDEATIREIISEVDTDNDGRINYEEFCTMMRSGTQQ 1736
>TC216778 UP|O24431 (O24431) Calmodulin-like domain protein kinase isoenzyme
gamma , complete
Length = 2377
Score = 200 bits (508), Expect = 5e-52
Identities = 123/229 (53%), Positives = 151/229 (65%), Gaps = 17/229 (7%)
Frame = +1
Query: 18 WPSISPSAKDLVRKMLNSDPKQRISAYEVLNHPWIKEDGEAPDTPLDNAVLNR------- 70
WP+IS +AKDLVRKML DPK+RI++ +VL HPWIK DG A D P+D+AVL+R
Sbjct: 1363 WPNISNNAKDLVRKMLIQDPKKRITSAQVLEHPWIK-DGNASDKPIDSAVLSRMKQFRAM 1539
Query: 71 NSLEQ*TDSRKLL*SCLSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQ 130
N L++ + K++ +S EEI GLK MF MDTD SGTIT EELK GL + G++L+E +
Sbjct: 1540 NKLKKL--ALKVIAENMSAEEIQGLKAMFTNMDTDKSGTITYEELKSGLHRLGSKLTEAE 1713
Query: 131 VKQLMEAADADGNGIIDYDEFITAS--QQLCTRTD*TEKNMFILPSNTSIKIT-TELEQA 187
VKQLMEAAD DGNG IDY EFITA+ + R D K + S IT ELE A
Sbjct: 1714 VKQLMEAADVDGNGSIDYIEFITATMHRHKLERDDQLFKAFQYFDKDNSGFITRDELESA 1893
Query: 188 LHEYNMHDGRYIKE-------IISEVDADNDGRINYDEFVAMMRKGNPE 229
+ EY M D IKE IISEVD D+DGRINY+EF AMM+ GN +
Sbjct: 1894 MKEYGMGDDATIKEIISEVDTIISEVDTDHDGRINYEEFSAMMKSGNQQ 2040
>TC217368 similar to UP|Q93YF4 (Q93YF4) Calcium-dependent protein kinase 2,
partial (66%)
Length = 1349
Score = 197 bits (500), Expect = 5e-51
Identities = 118/229 (51%), Positives = 150/229 (64%), Gaps = 12/229 (5%)
Frame = +1
Query: 18 WPSISPSAKDLVRKMLNSDPKQRISAYEVLNHPWIKEDGEAPDTPLDNAVLNRNSLEQ*T 77
WPSIS SAKDLVRKML DP++R++A++VL HPWI+ DG APD PLD+AVL+R L+Q +
Sbjct: 442 WPSISESAKDLVRKMLVRDPRRRLTAHQVLCHPWIQVDGVAPDKPLDSAVLSR--LKQFS 615
Query: 78 DSRKL-------L*SCLSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQ 130
KL + LSEEEI GLK+MFK +D DNSG IT EELK GL + G L E +
Sbjct: 616 AMNKLKKMALIIIAESLSEEEIAGLKEMFKMIDADNSGQITFEELKAGLKRVGANLKESE 795
Query: 131 VKQLMEAADADGNGIIDYDEFITASQQLCTRTD*TEKNMFILPS-----NTSIKITTELE 185
+ LM+AAD D +G IDY EF+ A+ L E N+F S + EL+
Sbjct: 796 IYDLMQAADVDNSGTIDYGEFLAAT--LHRNKIEREDNLFAAFSYFDKDGSGYITQEELQ 969
Query: 186 QALHEYNMHDGRYIKEIISEVDADNDGRINYDEFVAMMRKGNPEAHTKK 234
QA E+ + D R ++EII E+D DNDGRI+Y+EFVAMM+KGN A KK
Sbjct: 970 QACDEFGIKDVR-LEEIIKEIDEDNDGRIDYNEFVAMMQKGNLPAVGKK 1113
>TC228250 UP|Q84P29 (Q84P29) Seed calcium dependent protein kinase a, complete
Length = 1843
Score = 191 bits (484), Expect = 3e-49
Identities = 111/220 (50%), Positives = 147/220 (66%), Gaps = 10/220 (4%)
Frame = +3
Query: 18 WPSISPSAKDLVRKMLNSDPKQRISAYEVLNHPWIKEDGEAPDTPLDNAVLNRNSLEQ*T 77
WPSIS SAKDL+RKML+ +PK R++A+EVL HPWI +D APD PLD+AVL+R L+Q +
Sbjct: 894 WPSISDSAKDLIRKMLDQNPKTRLTAHEVLRHPWIVDDNIAPDKPLDSAVLSR--LKQFS 1067
Query: 78 DSRKL-------L*SCLSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQ 130
KL + LSEEEI GLK++FK +DTDNSGTIT +ELK GL + G+ L E +
Sbjct: 1068 AMNKLKKMALRVIAERLSEEEIGGLKELFKMIDTDNSGTITFDELKDGLKRVGSELMESE 1247
Query: 131 VKQLMEAADADGNGIIDYDEFITASQQL--CTRTD*TEKNMFILPSNTSIKIT-TELEQA 187
+K LM+AAD D +G IDY EFI A+ L R + + S IT E++QA
Sbjct: 1248 IKDLMDAADIDKSGTIDYGEFIAATVHLNKLEREENLVSAFSYFDKDGSGYITLDEIQQA 1427
Query: 188 LHEYNMHDGRYIKEIISEVDADNDGRINYDEFVAMMRKGN 227
++ + D +I ++I E+D DNDG+I+Y EF AMMRKGN
Sbjct: 1428 CKDFGL-DDIHIDDMIKEIDQDNDGQIDYGEFAAMMRKGN 1544
>TC205579 homologue to UP|Q43676 (Q43676) Calcium dependent protein kinase ,
partial (52%)
Length = 1481
Score = 185 bits (470), Expect = 1e-47
Identities = 112/220 (50%), Positives = 144/220 (64%), Gaps = 10/220 (4%)
Frame = +1
Query: 18 WPSISPSAKDLVRKMLNSDPKQRISAYEVLNHPWIKEDGEAPDTPLDNAVLNRNSLEQ*T 77
WP IS SAKDL+RKML S P +R++A++VL HPWI E+G APD LD AVL+R L+Q +
Sbjct: 40 WPLISDSAKDLIRKMLCSRPSERLTAHQVLCHPWICENGVAPDRSLDPAVLSR--LKQFS 213
Query: 78 DSRKL-------L*SCLSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQ 130
KL + LSEEEI GL++MF+ MDTDNSG IT +ELK GL + G+ L + +
Sbjct: 214 AMNKLKKMALRVIAESLSEEEIAGLREMFQAMDTDNSGAITFDELKAGLRRYGSTLKDIE 393
Query: 131 VKQLMEAADADGNGIIDYDEFITASQQL--CTRTD*TEKNMFILPSNTSIKITT-ELEQA 187
++ LMEAAD D +G IDY EFI A+ L R + + S IT EL+QA
Sbjct: 394 IRDLMEAADVDKSGTIDYGEFIAATVHLNKLEREEHLIAAFQYFDKDGSGYITVDELQQA 573
Query: 188 LHEYNMHDGRYIKEIISEVDADNDGRINYDEFVAMMRKGN 227
E NM D ++++II EVD DNDGRI+Y EF AMM+KGN
Sbjct: 574 CAEQNMTDA-FLEDIIREVDQDNDGRIDYGEFAAMMQKGN 690
>TC211152 UP|Q84P28 (Q84P28) Seed calcium dependent protein kinase b, complete
Length = 1740
Score = 184 bits (468), Expect = 2e-47
Identities = 108/223 (48%), Positives = 149/223 (66%), Gaps = 12/223 (5%)
Frame = +1
Query: 18 WPSISPSAKDLVRKMLNSDPKQRISAYEVLNHPWIKEDGEAPDTPLDNAVLNR------- 70
WPSIS +AK+LV++ML+ DPK+RISA+EVL +PW+ +D APD PLD+AVL R
Sbjct: 799 WPSISENAKELVKQMLDRDPKKRISAHEVLCNPWVVDD-IAPDKPLDSAVLTRLKHFSAM 975
Query: 71 NSLEQ*TDSRKLL*SCLSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQ 130
N L++ + +++ LSEEEI GLK++FK +DTDNSGTIT EELK GL G+ L E +
Sbjct: 976 NKLKK--MALRVIAERLSEEEIGGLKELFKMIDTDNSGTITFEELKEGLKSVGSNLMESE 1149
Query: 131 VKQLMEAADADGNGIIDYDEFITASQQLCTRTD*TEKNM-----FILPSNTSIKITTELE 185
+K LMEAAD D NG IDY EF+ A+ L E+N+ + + EL+
Sbjct: 1150 IKSLMEAADIDNNGSIDYGEFLAATLHLNKME--REENLVAAFAYFDKDGSGYITIDELQ 1323
Query: 186 QALHEYNMHDGRYIKEIISEVDADNDGRINYDEFVAMMRKGNP 228
QA ++++ D ++ E+I E+D DNDGRI+Y EF AMM+KG+P
Sbjct: 1324 QACKDFSLGD-VHLDEMIKEIDQDNDGRIDYAEFAAMMKKGDP 1449
>TC225062 similar to UP|Q8W4I7 (Q8W4I7) Calcium-dependent protein kinase,
partial (60%)
Length = 1701
Score = 150 bits (378), Expect = 7e-37
Identities = 95/221 (42%), Positives = 130/221 (57%), Gaps = 12/221 (5%)
Frame = +1
Query: 18 WPSISPSAKDLVRKMLNSDPKQRISAYEVLNHPWIKEDGEAPDTPLDNAVLNRNSLEQ*T 77
WPSIS SAK LVR+ML DPK R++A +VL HPWI+ +AP+ PL + V +R L+Q +
Sbjct: 196 WPSISESAKSLVRQMLEPDPKLRLTAKQVLEHPWIQNAKKAPNVPLGDVVKSR--LKQFS 369
Query: 78 -------DSRKLL*SCLSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQ 130
+ +++ LS EE+ +K MFK MD DN G ++IEELK G G+ L++ +
Sbjct: 370 MMNRFKRKALRVIADFLSNEEVEDIKDMFKKMDNDNDGIVSIEELKAGFRNFGSLLADSE 549
Query: 131 VKQLMEAADADGNGIIDYDEFITASQQL--CTRTD*TEK--NMFILPSNTSIKITTELEQ 186
V+ L+EA D++G G +DY EF+ S L D K + F N I+ EL
Sbjct: 550 VQLLIEAVDSNGKGTLDYGEFVAVSLHLRRMANDDHLHKAFSYFDKDGNGYIE-PDELRN 726
Query: 187 ALHEYNMHDGRYI-KEIISEVDADNDGRINYDEFVAMMRKG 226
AL E D + +I EVD D DGRI+YDEFVAMM+ G
Sbjct: 727 ALMEDGADDCTDVANDIFLEVDTDKDGRISYDEFVAMMKTG 849
>CD409746 similar to PIR|T08873|T08 calcium-dependent protein kinase (EC
2.7.1.-) beta - soybean, partial (37%)
Length = 688
Score = 131 bits (329), Expect = 3e-31
Identities = 82/190 (43%), Positives = 111/190 (58%), Gaps = 12/190 (6%)
Frame = -3
Query: 58 APDTPLDNAVLNR-------NSLEQ*TDSRKLL*SCLSEEEIMGLKQMFKGMDTDNSGTI 110
APD PLD AVL R N L++ + +++ LSEEEI GLK++FK +D DNSGTI
Sbjct: 680 APDKPLDPAVLTRLKHFSTMNKLQK--MALRIIAERLSEEEIGGLKELFKMIDEDNSGTI 507
Query: 111 TIEELK*GLVKQGTRLSEQQVKQLMEAADADGNGIIDYDEFITASQQLCTRTD*TEKNM- 169
T EELK L G L E ++K LMEAAD D NG IDY EF+ + L E+N+
Sbjct: 506 TFEELKDSLKSVGCDLIESEIKFLMEAADIDNNGTIDYGEFLAGTLHLNKME--REENLV 333
Query: 170 ----FILPSNTSIKITTELEQALHEYNMHDGRYIKEIISEVDADNDGRINYDEFVAMMRK 225
+ + E++QA ++ + ++ EII+E+D DNDGRINY EF AMMRK
Sbjct: 332 AAFAYFDKDGSGYITIEEIQQACKDFGL-GNLHLDEIINEIDQDNDGRINYAEFAAMMRK 156
Query: 226 GNPEAHTKKR 235
G P+ ++
Sbjct: 155 GGPDVGRSRK 126
>TC227658 similar to UP|O48565 (O48565) Calcium-dependent protein kinase,
partial (43%)
Length = 1107
Score = 123 bits (309), Expect = 7e-29
Identities = 82/207 (39%), Positives = 115/207 (54%), Gaps = 11/207 (5%)
Frame = +2
Query: 31 KMLNSDPKQRISAYEVLNHPWIKEDGEAPDTPLDNAVLNRNSLEQ*TDSRKL-------L 83
KML+ DPK+R++A +VL+HPW++ +AP+ L V R L+Q + KL +
Sbjct: 14 KMLDPDPKRRLTAQDVLDHPWLQNAKKAPNVSLGETV--RARLKQFSVMNKLKKRALRVI 187
Query: 84 *SCLSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQVKQLMEAADADGN 143
L+ EE GLK+ F+ MDT+N G I I+EL+ GL K G ++ E V+ LMEA D DG+
Sbjct: 188 AEHLTVEEAAGLKEGFQVMDTNNRGKINIDELRVGLHKLGHQVPESDVQALMEAGDVDGD 367
Query: 144 GIIDYDEFITASQQL--CTRTD*TEKNMFILPSNTSIKI-TTELEQAL-HEYNMHDGRYI 199
G +DY EF+ S L + K N S I EL AL + + + I
Sbjct: 368 GHLDYGEFVAISVHLRKMGNDEHLRKAFQFFDQNKSEYIEIEELRSALSDDLDTNSEEVI 547
Query: 200 KEIISEVDADNDGRINYDEFVAMMRKG 226
I+ +VD D DGRI+YDEF MM+ G
Sbjct: 548 SAIMHDVDTDKDGRISYDEFSTMMKAG 628
>TC222915 homologue to UP|Q84P29 (Q84P29) Seed calcium dependent protein
kinase a, partial (45%)
Length = 688
Score = 119 bits (298), Expect = 1e-27
Identities = 65/120 (54%), Positives = 86/120 (71%), Gaps = 7/120 (5%)
Frame = +1
Query: 18 WPSISPSAKDLVRKMLNSDPKQRISAYEVLNHPWIKEDGEAPDTPLDNAVLNRNSLEQ*T 77
WPSIS SAKDL+RKML+ +PK R++A++VL HPWI +D APD PLD+AVL+R L+Q +
Sbjct: 334 WPSISDSAKDLIRKMLDRNPKTRVTAHQVLCHPWIVDDNIAPDKPLDSAVLSR--LKQFS 507
Query: 78 DSRKL-------L*SCLSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQ 130
KL + LSEEEI GLK++F+ +D DNSGTIT +ELK GL + G+ L E +
Sbjct: 508 AMNKLKKMALRVIAERLSEEEIGGLKELFRMIDADNSGTITFDELKEGLKRVGSELMESE 687
Score = 27.7 bits (60), Expect = 4.9
Identities = 11/31 (35%), Positives = 18/31 (57%)
Frame = +1
Query: 199 IKEIISEVDADNDGRINYDEFVAMMRKGNPE 229
+KE+ +DADN G I +DE +++ E
Sbjct: 580 LKELFRMIDADNSGTITFDELKEGLKRVGSE 672
>BI471390 similar to GP|28416563|gb At1g18890 {Arabidopsis thaliana}, partial
(25%)
Length = 421
Score = 113 bits (283), Expect = 7e-26
Identities = 63/139 (45%), Positives = 90/139 (64%), Gaps = 7/139 (5%)
Frame = +3
Query: 21 ISPSAKDLVRKMLNSDPKQRISAYEVLNHPWIKEDGEAPDTPLDNAVLNRNSLEQ*T--- 77
IS SAK LVR+ML DPK+R++A +VL H W++ +A + PL + V R L+Q +
Sbjct: 6 ISDSAKSLVRQMLEPDPKKRLTAEQVLEHSWLQNAKKASNVPLGDIV--RTRLKQFSVMN 179
Query: 78 ----DSRKLL*SCLSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQVKQ 133
+ +++ LS EE+ +K MF MDTD G +T EELK GL K G++L+E ++K
Sbjct: 180 RFKKRALRVIAEHLSVEEVEIIKDMFTLMDTDKDGKVTYEELKVGLRKVGSQLAEPEIKM 359
Query: 134 LMEAADADGNGIIDYDEFI 152
LME AD DGNG++DY EF+
Sbjct: 360 LMEVADVDGNGVLDYGEFV 416
Score = 28.1 bits (61), Expect = 3.7
Identities = 10/27 (37%), Positives = 18/27 (66%)
Frame = +3
Query: 199 IKEIISEVDADNDGRINYDEFVAMMRK 225
IK++ + +D D DG++ Y+E +RK
Sbjct: 243 IKDMFTLMDTDKDGKVTYEELKVGLRK 323
>BG881805
Length = 401
Score = 94.4 bits (233), Expect = 4e-20
Identities = 51/121 (42%), Positives = 76/121 (62%), Gaps = 7/121 (5%)
Frame = +1
Query: 45 EVLNHPWIKEDGEAPDTPLDNAVLNRNSLEQ*T-------DSRKLL*SCLSEEEIMGLKQ 97
+VL HPW++ +AP+ PL + V R+ L+Q + + +++ LS EE+ +K
Sbjct: 4 QVLEHPWLQNAKKAPNVPLGDIV--RSRLKQFSVMNRFKKKALRVIADHLSVEEVEIIKD 177
Query: 98 MFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQVKQLMEAADADGNGIIDYDEFITASQQ 157
MF MDTD G +T EELK GL K G++L+E ++K LME AD DGNG++DY EF+ +
Sbjct: 178 MFTLMDTDKDGRVTFEELKAGLRKVGSQLAEPEIKMLMEVADVDGNGVLDYGEFVAVTIH 357
Query: 158 L 158
L
Sbjct: 358 L 360
>BE475418 similar to PIR|T06126|T06 calcium-dependent protein kinase (EC
2.7.1.-) CPK5 - Arabidopsis thaliana, partial (25%)
Length = 523
Score = 85.5 bits (210), Expect = 2e-17
Identities = 60/154 (38%), Positives = 86/154 (54%), Gaps = 3/154 (1%)
Frame = +2
Query: 87 LSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQVKQLMEAADADGNGII 146
LSEE+I GL QMF+ MDTDNSG +T +ELK GL++ G+ L + + LMEAAD D +G I
Sbjct: 26 LSEEDIAGLIQMFQAMDTDNSGALTFDELKAGLIRYGSTLKDI*IHDLMEAADVDKSGTI 205
Query: 147 DYDEFITASQQL-CTRTD*TEKNMFILPSNTSIKITT--ELEQALHEYNMHDGRYIKEII 203
Y EFI A+ L +* F N T +L+QA ++NM D ++ II
Sbjct: 206 HYGEFIAATFHLNKLDRE*HLIATFQYSDNDGRGYITXDKLQQACAQHNMTD-TSLEYII 382
Query: 204 SEVDADNDGRINYDEFVAMMRKGNPEAHTKKRHD 237
+ + +I+Y++F A + K P K H+
Sbjct: 383 Q*IYQNYYTKIDYNKFSAFIHKYPPHIINKTIHN 484
>BM527875 similar to PIR|T46189|T46 calcium-dependent protein kinase -
Arabidopsis thaliana, partial (25%)
Length = 426
Score = 83.2 bits (204), Expect = 1e-16
Identities = 47/124 (37%), Positives = 75/124 (59%), Gaps = 7/124 (5%)
Frame = +1
Query: 42 SAYEVLNHPWIKEDGEAPDTPLDNAVLNR-------NSLEQ*TDSRKLL*SCLSEEEIMG 94
+A EVL+HPW++ + +AP+ L V +R N L++ + +++ LS EE G
Sbjct: 1 TAQEVLDHPWLQNEKKAPNVSLGETVRSRLMQFSVMNKLKK--RALRVIAEYLSLEEAAG 174
Query: 95 LKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQVKQLMEAADADGNGIIDYDEFITA 154
+K+ F+ MDT N G I+++EL+ GL K G ++ + V+ LM+A D D +G IDY EF+
Sbjct: 175 IKEGFQLMDTSNKGKISVDELRVGLHKLGHQIPDGDVQILMDAGDVDNDGYIDYGEFVAI 354
Query: 155 SQQL 158
S L
Sbjct: 355 SIHL 366
Score = 31.6 bits (70), Expect = 0.34
Identities = 21/57 (36%), Positives = 33/57 (57%), Gaps = 5/57 (8%)
Frame = +1
Query: 183 ELEQALHE--YNMHDGRYIKEIISEVDADNDGRINYDEFVAM---MRKGNPEAHTKK 234
EL LH+ + + DG ++ ++ D DNDG I+Y EFVA+ +RK + + H K
Sbjct: 232 ELRVGLHKLGHQIPDGD-VQILMDAGDVDNDGYIDYGEFVAISIHLRKIDNDEHLHK 399
>TC227424 UP|Q39890 (Q39890) Calmodulin, complete
Length = 1424
Score = 77.4 bits (189), Expect = 5e-15
Identities = 45/142 (31%), Positives = 74/142 (51%), Gaps = 5/142 (3%)
Frame = +3
Query: 87 LSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQVKQLMEAADADGNGII 146
LSEE+I+ K+ F D D G IT+EEL + +E++++ ++ DADGNG I
Sbjct: 669 LSEEQIVDFKEAFGLFDKDGDGCITVEELATVIRSLDQNPTEEELQDMISEVDADGNGTI 848
Query: 147 DYDEFITASQQLCTRTD*TEK-----NMFILPSNTSIKITTELEQALHEYNMHDGRYIKE 201
++DEF++ + TD E+ +F N I + ++ +++
Sbjct: 849 EFDEFLSLMAKKVKDTDAEEELKEAFKVFDKDQNGYISASELRHVMINLGEKLTDEEVEQ 1028
Query: 202 IISEVDADNDGRINYDEFVAMM 223
+I E D D DG++NY+EFV MM
Sbjct: 1029MIKEADLDGDGQVNYEEFVKMM 1094
Score = 38.9 bits (89), Expect = 0.002
Identities = 32/103 (31%), Positives = 48/103 (46%), Gaps = 3/103 (2%)
Frame = +3
Query: 126 LSEQQVKQLMEAA---DADGNGIIDYDEFITASQQLCTRTD*TEKNMFILPSNTSIKITT 182
LSE+Q+ EA D DG+G I +E T + L ++N T
Sbjct: 669 LSEEQIVDFKEAFGLFDKDGDGCITVEELATVIRSL-------DQN------------PT 791
Query: 183 ELEQALHEYNMHDGRYIKEIISEVDADNDGRINYDEFVAMMRK 225
E E ++++ISEVDAD +G I +DEF+++M K
Sbjct: 792 EEE-------------LQDMISEVDADGNGTIEFDEFLSLMAK 881
>TC224985 homologue to UP|Q93YI3 (Q93YI3) Calcium dependent calmodulin
independent protein kinase (Fragment), partial (14%)
Length = 852
Score = 77.4 bits (189), Expect = 5e-15
Identities = 39/57 (68%), Positives = 47/57 (82%)
Frame = +1
Query: 81 KLL*SCLSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQVKQLMEA 137
K++ LSEEEI+GLK+MFK MD DNSGTIT EELK GL K GT++SE +V+QLMEA
Sbjct: 67 KVIAEXLSEEEIIGLKEMFKSMDIDNSGTITFEELKAGLPKLGTKVSESEVRQLMEA 237
>TC234066 similar to UP|Q8LDS1 (Q8LDS1) Calcium-dependent protein kinase,
partial (28%)
Length = 445
Score = 76.6 bits (187), Expect = 9e-15
Identities = 41/115 (35%), Positives = 73/115 (62%), Gaps = 8/115 (6%)
Frame = +1
Query: 49 HPWIKEDGEAPDTPLDNAVLNR-------NSLEQ*TDSRKLL*SCLSEEEIMGLKQMFKG 101
HPW++E GEA + P+D +VLN + L+Q + + L S L+E E+ LK F
Sbjct: 10 HPWVREGGEALEIPIDISVLNNMRQFVKYSRLKQ--FALRALASTLNEGELSDLKDQFDA 183
Query: 102 MDTDNSGTITIEELK*GLVK-QGTRLSEQQVKQLMEAADADGNGIIDYDEFITAS 155
+D D +G+I++EE++ L K Q +L E +V ++++A D++ +G++D+ EF+ A+
Sbjct: 184 IDVDKNGSISLEEMRQALAKDQPWKLKESRVLEILQAIDSNTDGLVDFTEFVAAT 348
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.323 0.137 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,813,613
Number of Sequences: 63676
Number of extensions: 80301
Number of successful extensions: 690
Number of sequences better than 10.0: 215
Number of HSP's better than 10.0 without gapping: 571
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 614
length of query: 240
length of database: 12,639,632
effective HSP length: 94
effective length of query: 146
effective length of database: 6,654,088
effective search space: 971496848
effective search space used: 971496848
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 57 (26.6 bits)
Medicago: description of AC146941.2


