

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146940.4 + phase: 1 /pseudo
(674 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
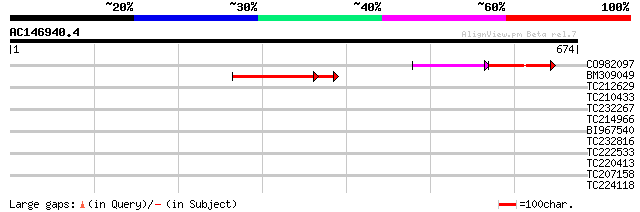
Score E
Sequences producing significant alignments: (bits) Value
CO982097 117 2e-35
BM309049 107 5e-25
TC212629 39 0.006
TC210433 similar to UP|Q9LRP0 (Q9LRP0) Emb|CAB39405.1, partial (... 39 0.007
TC232267 similar to GB|AAF68117.1|7715599|AC010793 F20B17.17 {Ar... 33 0.31
TC214966 similar to UP|Q6R5L6 (Q6R5L6) Sadtomato protein (Fragme... 30 0.33
BI967540 33 0.54
TC232816 33 0.54
TC222533 similar to UP|Q9SKV9 (Q9SKV9) F5J5.11, partial (11%) 31 2.0
TC220413 similar to UP|Q9LF51 (Q9LF51) Glutamine-rich protein, p... 30 3.5
TC207158 29 5.9
TC224118 similar to UP|Q940Z3 (Q940Z3) At2g36480/F1O11.11 (Expre... 29 7.7
>CO982097
Length = 805
Score = 117 bits (293), Expect(2) = 2e-35
Identities = 57/81 (70%), Positives = 70/81 (86%), Gaps = 1/81 (1%)
Frame = -2
Query: 570 KRNRLEHQRLNDLVFVRYNLMLENRNN-KIRNYDPINDELLDDHHDNWVLEDSPPFLTVE 628
KRNRLEHQ+LNDLV+VRYNL L+ RN +++NYDPIN E LDDH NWV+++SPPFLT E
Sbjct: 531 KRNRLEHQKLNDLVYVRYNLKLQQRNPLRLQNYDPINFETLDDH-SNWVVKESPPFLTNE 355
Query: 629 ELESLRNDLANMTIQPISNDI 649
E++ LRNDL NM+IQPIS+DI
Sbjct: 354 EIDVLRNDLTNMSIQPISDDI 292
Score = 51.2 bits (121), Expect(2) = 2e-35
Identities = 40/92 (43%), Positives = 51/92 (54%)
Frame = -1
Query: 479 YRCH*KVCS**S*AASKVKYRDKYI*KF*GRLWKEICYRSSKFTISR*MVRTLRVSSTTF 538
+ CH*KVCS S* A ++ I*K R W+ CY +++ +R*MV L + T
Sbjct: 802 FGCH*KVCSWRS*FAI*FGK*NENI*KCRVRFWEASCYT*TEYGDAR*MVGILWMWYTKL 623
Query: 539 AKIGDSGSKSNL*LFWLREKLECV*AYSLKKK 570
K G S SK NL F L +LE +*AYS K K
Sbjct: 622 TKAGYSCSKPNLQCFKL*TELEYI*AYSFK*K 527
>BM309049
Length = 430
Score = 107 bits (267), Expect(2) = 5e-25
Identities = 55/102 (53%), Positives = 70/102 (67%)
Frame = +2
Query: 266 VLVSLCNSLH*LDVSRHWKIT*S*RGNSTCHKCYQVCI*SLLSIVFDEEIYSWKRDTLSC 325
+LVS+C ++H* DV H++I S TC K YQV + SLL +FDE+IY WKR T S
Sbjct: 17 ILVSMCYTVH*FDVG*HYEIRGSK*DCVTCFKNYQVYLQSLLFFIFDEKIYRWKRYTWSS 196
Query: 326 SNSLCH*FHCFAEYFVSEKCT*SHGNISRMDNFCLCKRCQGQ 367
SNS+CH*FHC A+Y S +CT SHGNI R+D LC+R G+
Sbjct: 197 SNSVCH*FHCLAKYIGS*RCTKSHGNI*RLDKLSLCQRI*GK 322
Score = 25.8 bits (55), Expect(2) = 5e-25
Identities = 11/29 (37%), Positives = 18/29 (61%)
Frame = +3
Query: 362 KRCQGQTICGTSLEH*LLDCLC*HSETHR 390
K + + CGT+L +L+ +C*+ E HR
Sbjct: 306 KESKAKKNCGTNLRLQVLEKMC*YCEAHR 392
>TC212629
Length = 792
Score = 39.3 bits (90), Expect = 0.006
Identities = 21/44 (47%), Positives = 27/44 (60%)
Frame = +3
Query: 569 KKRNRLEHQRLNDLVFVRYNLMLENRNNKIRNYDPINDELLDDH 612
KKRNR EH+RL++LV+V N L R N DPI+ +D H
Sbjct: 195 KKRNRHEHKRLHNLVYVECNQALVKRYNYRDEIDPISLNDIDIH 326
>TC210433 similar to UP|Q9LRP0 (Q9LRP0) Emb|CAB39405.1, partial (10%)
Length = 748
Score = 38.9 bits (89), Expect = 0.007
Identities = 20/59 (33%), Positives = 37/59 (61%), Gaps = 8/59 (13%)
Frame = +3
Query: 569 KKRNRLEHQRLNDLVFVRYNLML-------ENRNNKIRNYDPI-NDELLDDHHDNWVLE 619
K++NRL ++LND+++V YNL L +R++K+ + D + + LLDD W+++
Sbjct: 306 KRQNRLSQKKLNDIIYVHYNLRLRECQLRKRSRDSKLSSVDNVLQEHLLDD----WIVD 470
>TC232267 similar to GB|AAF68117.1|7715599|AC010793 F20B17.17 {Arabidopsis
thaliana;} , partial (35%)
Length = 851
Score = 33.5 bits (75), Expect = 0.31
Identities = 13/25 (52%), Positives = 19/25 (76%)
Frame = +1
Query: 567 LKKKRNRLEHQRLNDLVFVRYNLML 591
L +KRN+++ + NDLV+V YNL L
Sbjct: 322 LSEKRNKIDRETFNDLVYVNYNLKL 396
>TC214966 similar to UP|Q6R5L6 (Q6R5L6) Sadtomato protein (Fragment), partial
(88%)
Length = 1176
Score = 29.6 bits (65), Expect(2) = 0.33
Identities = 19/67 (28%), Positives = 25/67 (36%), Gaps = 10/67 (14%)
Frame = +3
Query: 311 FDEEIYSWK--RDTLSCSNSLCH*FHCFAEYFVSEK--------CT*SHGNISRMDNFCL 360
F E++Y W+ +C LC HCF K HG N C+
Sbjct: 270 FQEQVYVWESHHPA*ACGR*LCWNSHCFLYVIGRSKPQRV*F*VSRQHHGGTILCPNQCV 449
Query: 361 CKRCQGQ 367
C+RC Q
Sbjct: 450 CERCGQQ 470
Score = 22.3 bits (46), Expect(2) = 0.33
Identities = 11/23 (47%), Positives = 13/23 (55%), Gaps = 1/23 (4%)
Frame = +2
Query: 252 CCCW*VIGERVPWPVL-VSLCNS 273
CCCW G + WP L +L NS
Sbjct: 119 CCCW--SGWLMLWPTLPTNLINS 181
>BI967540
Length = 692
Score = 32.7 bits (73), Expect = 0.54
Identities = 21/61 (34%), Positives = 35/61 (56%), Gaps = 4/61 (6%)
Frame = -1
Query: 569 KKRNRLEHQRLNDLVFVRYNLMLEN--RNNKIRNYDPINDELLDDHHDNW--VLEDSPPF 624
+KRNR+E ++ ++LVFV NL L+ + + ++ I DE+ D W +E SPP
Sbjct: 425 RKRNRVELEKFSELVFVHSNLWLQTIFKRREAKSEPIIFDEI--DCSCEWPSEIESSPPL 252
Query: 625 L 625
+
Sbjct: 251 M 249
>TC232816
Length = 768
Score = 32.7 bits (73), Expect = 0.54
Identities = 18/55 (32%), Positives = 30/55 (53%), Gaps = 2/55 (3%)
Frame = +2
Query: 293 STCHKCYQVCI*SLLSIVFDEEIYSWKRD-TLS-CSNSLCH*FHCFAEYFVSEKC 345
S CH+C+ C +LS VF ++YS K++ LS N + + + F +Y + C
Sbjct: 197 SLCHRCWLKCKSKILSTVFSRKLYSRKKEKNLSWVGNGVVYNYSAF*KYEIYFSC 361
>TC222533 similar to UP|Q9SKV9 (Q9SKV9) F5J5.11, partial (11%)
Length = 596
Score = 30.8 bits (68), Expect = 2.0
Identities = 44/170 (25%), Positives = 68/170 (39%)
Frame = +2
Query: 242 SDSDR*CCKLCCCW*VIGERVPWPVLVSLCNSLH*LDVSRHWKIT*S*RGNSTCHKCYQV 301
S +R* +L *V+G +L SLC+SL+* D R+W+ + N + V
Sbjct: 2 SSCNR*WEQLYFSG*VVGGEKETYLLDSLCSSLY*FDA*RYWEASLDKEDN*KGN*SSWV 181
Query: 302 CI*SLLSIVFDEEIYSWKRDTLSCSNSLCH*FHCFAEYFVSEKCT*SHGNISRMDNFCLC 361
+ + F E+ Y + +C +CH + E * MD
Sbjct: 182 YLCPF*YLKFVEKFYKQEGIGETCYY*ICHFLSNLGKASQRESQY*KDVYF**MDLEQAI 361
Query: 362 KRCQGQTICGTSLEH*LLDCLC*HSETHRTTCTCVASRGQ*R*TCYGFSL 411
G+ C S L+ HS +H +TC +S G *+ T +G L
Sbjct: 362 *GA*GKRSCKGSAHAFFLE*CGLHS*SHGSTCESASSCGW*KETSHGLYL 511
>TC220413 similar to UP|Q9LF51 (Q9LF51) Glutamine-rich protein, partial (31%)
Length = 761
Score = 30.0 bits (66), Expect = 3.5
Identities = 10/16 (62%), Positives = 10/16 (62%)
Frame = -1
Query: 240 CCSDSDR*CCKLCCCW 255
CCS S CC CCCW
Sbjct: 323 CCSRSCCCCCCCCCCW 276
>TC207158
Length = 1004
Score = 29.3 bits (64), Expect = 5.9
Identities = 14/37 (37%), Positives = 20/37 (53%)
Frame = +1
Query: 337 AEYFVSEKCT*SHGNISRMDNFCLCKRCQGQTICGTS 373
A +S K T* H N+ +MDN C+R Q + +S
Sbjct: 58 ARSLMSSK*T*GHANLGKMDNGLSCQRDQESRLASSS 168
>TC224118 similar to UP|Q940Z3 (Q940Z3) At2g36480/F1O11.11 (Expressed
protein), partial (46%)
Length = 891
Score = 28.9 bits (63), Expect = 7.7
Identities = 16/42 (38%), Positives = 21/42 (49%)
Frame = +3
Query: 356 DNFCLCKRCQGQTICGTSLEH*LLDCLC*HSETHRTTCTCVA 397
DN+ L C+G++ CG E CLC*HS + C A
Sbjct: 714 DNYQLDYHCRGESSCG---EGHCSHCLC*HSRGELSLQYCAA 830
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.366 0.162 0.645
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 38,033,374
Number of Sequences: 63676
Number of extensions: 661973
Number of successful extensions: 11530
Number of sequences better than 10.0: 25
Number of HSP's better than 10.0 without gapping: 4750
Number of HSP's successfully gapped in prelim test: 447
Number of HSP's that attempted gapping in prelim test: 5911
Number of HSP's gapped (non-prelim): 6094
length of query: 674
length of database: 12,639,632
effective HSP length: 104
effective length of query: 570
effective length of database: 6,017,328
effective search space: 3429876960
effective search space used: 3429876960
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.6 bits)
S2: 62 (28.5 bits)
Medicago: description of AC146940.4


