

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146866.2 + phase: 0
(504 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
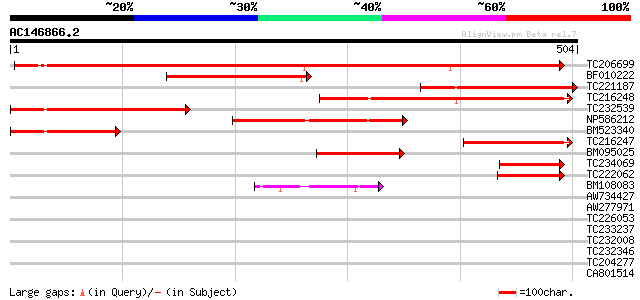
Score E
Sequences producing significant alignments: (bits) Value
TC206699 UP|Q7XA53 (Q7XA53) Sucrose transporter, complete 537 e-153
BF010222 similar to GP|28172870|em sucrose transporter 4 protein... 224 5e-59
TC221187 similar to UP|Q84RQ3 (Q84RQ3) Sucrose transporter 4 pro... 221 6e-58
TC216248 similar to UP|O04077 (O04077) Sucrose transport protein... 213 2e-55
TC232539 similar to UP|Q7XA53 (Q7XA53) Sucrose transporter, part... 209 2e-54
NP586212 hypothetical protein [Glycine max] 140 1e-33
BM523340 115 5e-26
TC216247 similar to UP|O04077 (O04077) Sucrose transport protein... 115 6e-26
BM095025 107 2e-23
TC234069 similar to UP|Q8LPM4 (Q8LPM4) Sucrose transporter 2, pa... 87 1e-17
TC222062 similar to UP|Q8LPM4 (Q8LPM4) Sucrose transporter 2, pa... 87 1e-17
BM108083 52 8e-07
AW734427 36 0.035
AW277971 similar to GP|22947842|gb 20S proteasome alpha 6 subuni... 33 0.30
TC226053 similar to UP|Q9SHY3 (Q9SHY3) F1E22.9 (At1g65720/F1E22_... 33 0.30
TC233237 33 0.39
TC232008 weakly similar to UP|Q9FIH3 (Q9FIH3) Serine/threonine p... 33 0.39
TC232346 similar to UP|Q96469 (Q96469) Calcium dependent protein... 32 0.51
TC204277 similar to UP|Q8LF69 (Q8LF69) 20S proteasome subunit PA... 32 0.67
CA801514 homologue to GP|7649151|gb|A sucrose transport protein ... 32 0.67
>TC206699 UP|Q7XA53 (Q7XA53) Sucrose transporter, complete
Length = 2053
Score = 537 bits (1384), Expect = e-153
Identities = 273/499 (54%), Positives = 347/499 (68%), Gaps = 10/499 (2%)
Frame = +1
Query: 5 TTTNPHRSRTRSSTSTSTSRPVQPVQPRTPLRQLLRVASVASGIQFGWALQLSLLTPYVQ 64
T N + + T + S P P++P +PLR+++ VAS+A+G+QFGWALQLSLLTPYVQ
Sbjct: 34 TKQNHNNNNTLTKPSLHVESP--PLEP-SPLRKIIVVASIAAGVQFGWALQLSLLTPYVQ 204
Query: 65 QLGIPHKWASIIWLCGPVSGLFVQPLVGHLSDRCSSRFGRRRPFILVGAASIVVAVVIIG 124
LGIPH WA+ IWLCGP+SG+ VQP+VG+ SDRC+SRFGRRRPFI G+ ++ +AV +IG
Sbjct: 205 LLGIPHTWAAYIWLCGPISGMLVQPIVGYHSDRCTSRFGRRRPFIAAGSFAVAIAVFLIG 384
Query: 125 YAADIGYLIGDDITQNYRPFAIVVFVIGFWILDVANNVTQGPCRALLADLTCNDARRTRV 184
YAAD+G+ GDD+++ RP AI +FV+GFWILDVANN+ QGPCRALL DL + ++TR
Sbjct: 385 YAADLGHSFGDDLSKKVRPRAIGIFVVGFWILDVANNMLQGPCRALLGDLCAGNHQKTRN 564
Query: 185 ANAYFSLFMAVGNILGYATGSYSGWYKIFTFTLTPACSISCANLKSAFFLDVAFIVVTTY 244
ANA+FS FMAVGN+LGYA G+YS Y +F FT T AC + CANLKS FFL +A +
Sbjct: 565 ANAFFSFFMAVGNVLGYAAGAYSKLYHVFPFTKTTACDVYCANLKSCFFLSIALLTTLAT 744
Query: 245 LSIVSAHEVPLSSSGA--GESGSAEEAFMWELFGTFKYFSMPVWIVLSVTALTWIGWFPF 302
++V EVPLS A + +LFG F+ P+WI+L VT L WI WFPF
Sbjct: 745 AALVYVKEVPLSPEKAVIDSDDNGGMPCFGQLFGAFRELKRPMWILLLVTCLNWIAWFPF 924
Query: 303 NLFDTDWMGREIYGGDPEGGLIYDTGVRMGALGLLLNSVVLAVTSLLMERLCR-KRGAGF 361
LFDTDWMGRE+Y G G YD GVR GALGL+LNSVVL TSL +E L R G
Sbjct: 925 LLFDTDWMGREVYEGTVGEGKAYDRGVRAGALGLMLNSVVLGATSLGVEVLARGVGGVKR 1104
Query: 362 VWGISNIFMAICF-IAMLVLTYAANSIGYV------SKGQPPPTGIVIAALAIFTILGFP 414
+WGI N +A+C + +LV A +S Y + PPP + ALA+F++LG P
Sbjct: 1105LWGIVNFLLAVCLAMTVLVTKMAQHSRQYTLLPNAHQEPLPPPAAVKAGALALFSLLGIP 1284
Query: 415 MAITYSVPYALISTHIEPLGLGQGLSMGVLNLAIVVPQIVVSLGSGPWDQLFGGGNSPAF 474
+AITYS+P+AL S G GQGLS+GVLNLAIV+PQ+VVS+ SGPWD LFGGGN PAF
Sbjct: 1285LAITYSIPFALASIFSSTSGAGQGLSLGVLNLAIVIPQMVVSVISGPWDALFGGGNLPAF 1464
Query: 475 AVAAVAALLSGLLALLAIP 493
V AVAA SG+L+++ +P
Sbjct: 1465VVGAVAAAASGILSIILLP 1521
>BF010222 similar to GP|28172870|em sucrose transporter 4 protein {Lotus
japonicus}, partial (26%)
Length = 414
Score = 224 bits (572), Expect = 5e-59
Identities = 109/136 (80%), Positives = 121/136 (88%), Gaps = 7/136 (5%)
Frame = +1
Query: 140 NYRPFAIVVFVIGFWILDVANNVTQGPCRALLADLTCNDARRTRVANAYFSLFMAVGNIL 199
+YRP AI VF++GFWILDVANNVTQGPCRALL DLT D RRTRVANAY+SLFMA+GNIL
Sbjct: 7 DYRPAAITVFIVGFWILDVANNVTQGPCRALLGDLTSKDPRRTRVANAYYSLFMAIGNIL 186
Query: 200 GYATGSYSGWYKIFTFTLTPACSISCANLKSAFFLDVAFIVVTTYLSIVSAHEVPLSSS- 258
GYATGSYSGWYKIFTF L+PAC+ISCANLKSAFFLD+AFI VTTY+SI++AHEVPL+SS
Sbjct: 187 GYATGSYSGWYKIFTFALSPACTISCANLKSAFFLDIAFIAVTTYISIMAAHEVPLNSSE 366
Query: 259 ------GAGESGSAEE 268
GAGESGSAEE
Sbjct: 367 AAHAEAGAGESGSAEE 414
>TC221187 similar to UP|Q84RQ3 (Q84RQ3) Sucrose transporter 4 protein,
partial (26%)
Length = 977
Score = 221 bits (563), Expect = 6e-58
Identities = 111/139 (79%), Positives = 125/139 (89%)
Frame = +2
Query: 366 SNIFMAICFIAMLVLTYAANSIGYVSKGQPPPTGIVIAALAIFTILGFPMAITYSVPYAL 425
SNI M +CF+AMLV+TY AN++GY+ K PP TGIVIAAL IFTILGFP+AITYSVPYAL
Sbjct: 2 SNIMMTVCFLAMLVVTYVANNMGYIGKDLPP-TGIVIAALIIFTILGFPLAITYSVPYAL 178
Query: 426 ISTHIEPLGLGQGLSMGVLNLAIVVPQIVVSLGSGPWDQLFGGGNSPAFAVAAVAALLSG 485
ISTHIE LGLGQGLSMGVLNLAIVVPQI+VSLGSGPWDQLFGGGNSPAFAVAAV+AL+SG
Sbjct: 179 ISTHIESLGLGQGLSMGVLNLAIVVPQIIVSLGSGPWDQLFGGGNSPAFAVAAVSALISG 358
Query: 486 LLALLAIPRTRTQKPRVRI 504
L+A+LAIPR+ QK R +
Sbjct: 359 LIAVLAIPRSGAQKARSHV 415
>TC216248 similar to UP|O04077 (O04077) Sucrose transport protein, partial
(41%)
Length = 1234
Score = 213 bits (541), Expect = 2e-55
Identities = 113/228 (49%), Positives = 150/228 (65%), Gaps = 3/228 (1%)
Frame = +1
Query: 276 GTFKYFSMPVWIVLSVTALTWIGWFPFNLFDTDWMGREIYGGDPEGGLIYDTGVRMGALG 335
G K P+W+++ VTA+ W+GWFP+ LFDTDWMGRE+YGG G Y GVR+G+LG
Sbjct: 1 GALKELKRPMWMLMLVTAVNWVGWFPYFLFDTDWMGREVYGGTA-GEDAYAKGVRVGSLG 177
Query: 336 LLLNSVVLAVTSLLMERLCRK-RGAGFVWGISNIFMAICFIAMLVLTYAANSIGYVSKGQ 394
L++N+VVL SL +E L + G +W I N +AI F +V+T A ++
Sbjct: 178 LMVNAVVLGFMSLAVEPLGKMVGGVKRLWAIVNFILAIGFGMTVVITKVAEHQRRMNPAA 357
Query: 395 P--PPTGIVIAALAIFTILGFPMAITYSVPYALISTHIEPLGLGQGLSMGVLNLAIVVPQ 452
P G+V+ ++ F +LG P+AIT+SVP+AL S + G GQGLS+GVLNLAIVVPQ
Sbjct: 358 VGHPSEGVVVGSMVFFGVLGVPLAITFSVPFALASIYCSASGAGQGLSLGVLNLAIVVPQ 537
Query: 453 IVVSLGSGPWDQLFGGGNSPAFAVAAVAALLSGLLALLAIPRTRTQKP 500
+VVS SGPWD LFGGGN PAF V A AA LS ++A++ +P T KP
Sbjct: 538 MVVSALSGPWDSLFGGGNLPAFMVGAAAAALSAIMAIVLLP---TPKP 672
>TC232539 similar to UP|Q7XA53 (Q7XA53) Sucrose transporter, partial (25%)
Length = 591
Score = 209 bits (532), Expect = 2e-54
Identities = 104/160 (65%), Positives = 124/160 (77%)
Frame = +1
Query: 1 MPNPTTTNPHRSRTRSSTSTSTSRPVQPVQPRTPLRQLLRVASVASGIQFGWALQLSLLT 60
M P+ T P+ S+TST+ S P Q +PLR++ VAS+A+GIQFGWALQLSLLT
Sbjct: 100 MEAPSPTKPNDPTKPSNTSTTLSLEAGPAQA-SPLRKMFAVASIAAGIQFGWALQLSLLT 276
Query: 61 PYVQQLGIPHKWASIIWLCGPVSGLFVQPLVGHLSDRCSSRFGRRRPFILVGAASIVVAV 120
PYVQ LG+PH AS IWLCGP+SGL VQP+VG+ SD C+SRFGRRRPFIL GA ++ VAV
Sbjct: 277 PYVQLLGVPHAAASFIWLCGPISGLVVQPIVGYYSDHCTSRFGRRRPFILGGALAVAVAV 456
Query: 121 VIIGYAADIGYLIGDDITQNYRPFAIVVFVIGFWILDVAN 160
+IGYAADIGY GDDI++ RP A+ VFVIGFWILDVAN
Sbjct: 457 FLIGYAADIGYAAGDDISKTTRPRAVGVFVIGFWILDVAN 576
>NP586212 hypothetical protein [Glycine max]
Length = 470
Score = 140 bits (353), Expect = 1e-33
Identities = 66/155 (42%), Positives = 99/155 (63%)
Frame = +2
Query: 199 LGYATGSYSGWYKIFTFTLTPACSISCANLKSAFFLDVAFIVVTTYLSIVSAHEVPLSSS 258
LGYA GSY G +K+F FT T AC + CANLKS FF + ++ ++++ + + +
Sbjct: 2 LGYAAGSYKGLHKMFPFTETKACDVFCANLKSCFFFSILLLLFLATVALLYVKDKQVEAR 181
Query: 259 GAGESGSAEEAFMWELFGTFKYFSMPVWIVLSVTALTWIGWFPFNLFDTDWMGREIYGGD 318
++ + + ++LF K P+W+++ VTA+ W+GWFP+ LFDTDWMGRE+YGG
Sbjct: 182 ALDDA--TQPSCFFQLFSALKELKRPMWMLMLVTAVNWVGWFPYFLFDTDWMGREVYGGQ 355
Query: 319 PEGGLIYDTGVRMGALGLLLNSVVLAVTSLLMERL 353
G Y GVR+G+LGL++N+VVL SL +E L
Sbjct: 356 -VGEDAYANGVRVGSLGLMVNAVVLGFMSLAVEPL 457
>BM523340
Length = 429
Score = 115 bits (288), Expect = 5e-26
Identities = 57/98 (58%), Positives = 72/98 (73%)
Frame = +2
Query: 1 MPNPTTTNPHRSRTRSSTSTSTSRPVQPVQPRTPLRQLLRVASVASGIQFGWALQLSLLT 60
M P+ T P+ S+TS++ S P + +PLR++ VAS+A+GIQFGWALQLSLLT
Sbjct: 137 MEAPSLTKPNDPTKPSNTSSTLSLEGGPGEA-SPLRKMFAVASIAAGIQFGWALQLSLLT 313
Query: 61 PYVQQLGIPHKWASIIWLCGPVSGLFVQPLVGHLSDRC 98
PYVQ LG+PH AS IWLCGP+SGL VQP+VG+ SD C
Sbjct: 314 PYVQLLGVPHAAASFIWLCGPISGLVVQPIVGYYSDHC 427
>TC216247 similar to UP|O04077 (O04077) Sucrose transport protein, partial
(17%)
Length = 930
Score = 115 bits (287), Expect = 6e-26
Identities = 58/97 (59%), Positives = 72/97 (73%)
Frame = +2
Query: 404 ALAIFTILGFPMAITYSVPYALISTHIEPLGLGQGLSMGVLNLAIVVPQIVVSLGSGPWD 463
++ F +LG P+AIT+SVP+AL S + G GQGLS+GVLNLAIVVPQ+VVS SGPWD
Sbjct: 2 SMVFFGVLGVPLAITFSVPFALASIYCSASGAGQGLSLGVLNLAIVVPQMVVSTLSGPWD 181
Query: 464 QLFGGGNSPAFAVAAVAALLSGLLALLAIPRTRTQKP 500
LFGGGN PAF V A AA LS ++A++ +P T KP
Sbjct: 182 ALFGGGNLPAFMVGAAAAALSAIMAIVLLP---TPKP 283
>BM095025
Length = 335
Score = 107 bits (266), Expect = 2e-23
Identities = 50/79 (63%), Positives = 57/79 (71%)
Frame = +3
Query: 273 ELFGTFKYFSMPVWIVLSVTALTWIGWFPFNLFDTDWMGREIYGGDPEGGLIYDTGVRMG 332
+LFG F+ P+WI+L VT L WI WFPF LFDTDWMGRE+Y G G YD GVR G
Sbjct: 99 QLFGAFRELKRPMWILLLVTCLNWIAWFPFLLFDTDWMGREVYEGTVGEGKAYDRGVRAG 278
Query: 333 ALGLLLNSVVLAVTSLLME 351
ALGL+LNSVVL TSL +E
Sbjct: 279 ALGLMLNSVVLGATSLGVE 335
>TC234069 similar to UP|Q8LPM4 (Q8LPM4) Sucrose transporter 2, partial (12%)
Length = 612
Score = 87.4 bits (215), Expect = 1e-17
Identities = 38/58 (65%), Positives = 50/58 (85%)
Frame = +3
Query: 436 GQGLSMGVLNLAIVVPQIVVSLGSGPWDQLFGGGNSPAFAVAAVAALLSGLLALLAIP 493
GQGL++GVLNLAIV+PQ+++SLGSGPWD LFGGGN PAF +A++ AL G++A L +P
Sbjct: 15 GQGLAIGVLNLAIVIPQMIISLGSGPWDALFGGGNIPAFVLASLCALAGGVIATLKLP 188
>TC222062 similar to UP|Q8LPM4 (Q8LPM4) Sucrose transporter 2, partial (13%)
Length = 428
Score = 87.4 bits (215), Expect = 1e-17
Identities = 40/60 (66%), Positives = 50/60 (82%)
Frame = +1
Query: 434 GLGQGLSMGVLNLAIVVPQIVVSLGSGPWDQLFGGGNSPAFAVAAVAALLSGLLALLAIP 493
G GQGL++GVLNLAIVVPQ+++SLGSGPWD LFGGGN PAF +A+V AL ++A L +P
Sbjct: 10 GGGQGLAIGVLNLAIVVPQMIISLGSGPWDALFGGGNIPAFVLASVCALAGAVIATLKLP 189
>BM108083
Length = 611
Score = 51.6 bits (122), Expect = 8e-07
Identities = 34/122 (27%), Positives = 54/122 (43%), Gaps = 7/122 (5%)
Frame = +3
Query: 218 TPACSISCANLKSAFFLDVAFI-----VVTTYLSIVSAHEVPLSSSGAGESGSAEEAFMW 272
T +C CANLKS FF + + V Y+ +PL + + + +
Sbjct: 3 TSSCRF-CANLKSCFFFSILLLLFLATVALLYVKDKQVEAIPLDDA-------TQPSCFF 158
Query: 273 ELFGTFKYFSMPVWIVLSVTALTWIGWFPFNLFD--TDWMGREIYGGDPEGGLIYDTGVR 330
+LFG K P+W+++ VTA+ W+G F T W+ E+YGG + G+
Sbjct: 159 QLFGALKELKRPMWMLMLVTAVNWVGVGFLTSFSTLTGWV-LEVYGGTAGEXRLTPKGIA 335
Query: 331 MG 332
G
Sbjct: 336 RG 341
Score = 39.3 bits (90), Expect = 0.004
Identities = 28/78 (35%), Positives = 39/78 (49%), Gaps = 4/78 (5%)
Frame = +2
Query: 294 LTWIGWFPFNLFDTDWMGREIYGGDPEGGLIYDTG-VRMGALGLLLNSVVLAVTSLLMER 352
L W GWFP+ LFDTDWMG + G Y G +LGL++N+V++ + L
Sbjct: 227 LGW-GWFPYFLFDTDWMGP*GLRWNCWGXTPYAXGNCAWDSLGLMVNAVLVWASCSLAVE 403
Query: 353 LCRKRGAG---FVWGISN 367
+R G +W I N
Sbjct: 404 PVGERXLGELKKLWAIVN 457
>AW734427
Length = 377
Score = 36.2 bits (82), Expect = 0.035
Identities = 17/22 (77%), Positives = 20/22 (90%)
Frame = -3
Query: 417 ITYSVPYALISTHIEPLGLGQG 438
ITY++PYALIS HI+ LGLGQG
Sbjct: 66 ITYNIPYALIS-HIQSLGLGQG 4
>AW277971 similar to GP|22947842|gb 20S proteasome alpha 6 subunit {Nicotiana
benthamiana}, partial (72%)
Length = 598
Score = 33.1 bits (74), Expect = 0.30
Identities = 15/28 (53%), Positives = 17/28 (60%)
Frame = +3
Query: 6 TTNPHRSRTRSSTSTSTSRPVQPVQPRT 33
T N HR+R RSS ST+TS P P T
Sbjct: 156 TPNSHRTRRRSSRSTTTSASASPASPST 239
>TC226053 similar to UP|Q9SHY3 (Q9SHY3) F1E22.9 (At1g65720/F1E22_13), partial
(26%)
Length = 1335
Score = 33.1 bits (74), Expect = 0.30
Identities = 17/41 (41%), Positives = 26/41 (62%), Gaps = 4/41 (9%)
Frame = +2
Query: 2 PNPTTTNPHRSRTRS----STSTSTSRPVQPVQPRTPLRQL 38
P P+TT+P ++TRS S +TS RP++P+ P P +L
Sbjct: 479 PPPSTTSPPTTKTRSRAPRSLATSRFRPLRPLPPSPPQLRL 601
>TC233237
Length = 420
Score = 32.7 bits (73), Expect = 0.39
Identities = 14/17 (82%), Positives = 17/17 (99%)
Frame = +1
Query: 437 QGLSMGVLNLAIVVPQI 453
+GLS+GVLNLAIVVPQ+
Sbjct: 274 EGLSLGVLNLAIVVPQV 324
>TC232008 weakly similar to UP|Q9FIH3 (Q9FIH3) Serine/threonine protein
kinase-like protein (At5g42440), partial (29%)
Length = 618
Score = 32.7 bits (73), Expect = 0.39
Identities = 17/45 (37%), Positives = 24/45 (52%)
Frame = +2
Query: 2 PNPTTTNPHRSRTRSSTSTSTSRPVQPVQPRTPLRQLLRVASVAS 46
P P TT+P S + + S S++RPV P P +P + S AS
Sbjct: 350 PAPPTTSPLTSSSAMAASASSTRPVSPTVPPSPSKNSPPTPSRAS 484
>TC232346 similar to UP|Q96469 (Q96469) Calcium dependent protein kinase ,
partial (5%)
Length = 521
Score = 32.3 bits (72), Expect = 0.51
Identities = 15/19 (78%), Positives = 18/19 (93%)
Frame = +3
Query: 437 QGLSMGVLNLAIVVPQIVV 455
+GLS+GVLNLAIVVPQ V+
Sbjct: 135 EGLSLGVLNLAIVVPQPVL 191
>TC204277 similar to UP|Q8LF69 (Q8LF69) 20S proteasome subunit PAF1, partial
(92%)
Length = 1153
Score = 32.0 bits (71), Expect = 0.67
Identities = 15/28 (53%), Positives = 17/28 (60%)
Frame = +1
Query: 6 TTNPHRSRTRSSTSTSTSRPVQPVQPRT 33
T N HR+R RSS ST+TS P P T
Sbjct: 232 TPNSHRTRRRSSRSTTTSASPSPASPPT 315
>CA801514 homologue to GP|7649151|gb|A sucrose transport protein {Euphorbia
esula}, partial (3%)
Length = 409
Score = 32.0 bits (71), Expect = 0.67
Identities = 14/16 (87%), Positives = 16/16 (99%)
Frame = +2
Query: 438 GLSMGVLNLAIVVPQI 453
GLS+GVLNLAIVVPQ+
Sbjct: 212 GLSLGVLNLAIVVPQV 259
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.325 0.140 0.437
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 23,962,165
Number of Sequences: 63676
Number of extensions: 364334
Number of successful extensions: 3080
Number of sequences better than 10.0: 64
Number of HSP's better than 10.0 without gapping: 3004
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3061
length of query: 504
length of database: 12,639,632
effective HSP length: 101
effective length of query: 403
effective length of database: 6,208,356
effective search space: 2501967468
effective search space used: 2501967468
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 61 (28.1 bits)
Medicago: description of AC146866.2


