

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146866.12 + phase: 0
(273 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
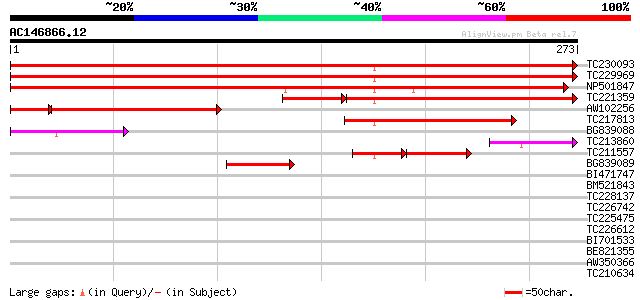
TC230093 similar to UP|Q8LGT9 (Q8LGT9) Phosphoglycerate mutase-l... 462 e-131
TC229969 UP|Q8LGT9 (Q8LGT9) Phosphoglycerate mutase-like protein... 447 e-126
NP501847 unknown [Glycine max] 310 6e-85
TC221359 UP|Q8LGT9 (Q8LGT9) Phosphoglycerate mutase-like protein... 179 1e-54
AW102256 145 2e-39
TC217813 similar to GB|BAB11607.1|10178227|AB025640 ZW10-like pr... 110 5e-25
BG839088 similar to GP|6520148|dbj| ZW10 {Arabidopsis thaliana},... 79 2e-15
TC213860 similar to UP|Q8LGT8 (Q8LGT8) Phosphoglycerate mutase-l... 56 2e-08
TC211557 similar to GB|BAB11607.1|10178227|AB025640 ZW10-like pr... 56 2e-08
BG839089 similar to GP|10178227|dbj ZW10-like protein {Arabidops... 47 9e-06
BI471747 34 0.062
BM521843 31 0.53
TC228137 similar to UP|Q8S902 (Q8S902) Syringolide-induced prote... 30 0.90
TC226742 similar to UP|ERO2_ARATH (Q7X9I4) Endoplasmic oxidoredu... 29 2.0
TC225475 UP|O22124 (O22124) Proton pyrophosphatase , partial (18%) 29 2.6
TC226612 similar to UP|Q8H0V8 (Q8H0V8) Spliceosome associated pr... 29 2.6
BI701533 28 5.8
BE821355 28 5.8
AW350366 similar to GP|25082966|gb| spliceosome associated prote... 27 9.9
TC210634 homologue to UP|Q6PV95 (Q6PV95) Beta-carotene hydroxyla... 27 9.9
>TC230093 similar to UP|Q8LGT9 (Q8LGT9) Phosphoglycerate mutase-like protein,
partial (96%)
Length = 1196
Score = 462 bits (1188), Expect = e-131
Identities = 216/274 (78%), Positives = 246/274 (88%), Gaps = 1/274 (0%)
Frame = +1
Query: 1 MDTAPGQSLYPLHHSKTIHLVRHAQGVHNVEGEKNHDAYLSYDFFDANLTPLGWQQVENL 60
MD A QSLYPLH KT+HLVRHAQGVHNV GEKNHDAY SY+FFDA+LT LGW+QV NL
Sbjct: 91 MDAAASQSLYPLHRCKTVHLVRHAQGVHNVAGEKNHDAYNSYEFFDAHLTSLGWEQVNNL 270
Query: 61 QKHVKAIGLSKKIELVVVSPLLRTMQTAVGVFGGEANTDGVNKPPLMIENVGHSDHPAVS 120
+KHVKA GLSK IELVV+SPLLRTMQTAVGVFGGEA TDG+ +PPLM ENVGHSDHPAVS
Sbjct: 271 RKHVKASGLSKNIELVVISPLLRTMQTAVGVFGGEAYTDGIGEPPLMTENVGHSDHPAVS 450
Query: 121 SLNCPPFVAVELCREQMGLHPCDKRRTVSEYRHMFPGIDFSLIETDDDTWWKPE-REKKE 179
S+NCPPF+AVELCREQ+G+HPCDKRRT+SEYR+MFP IDFSLIE+D+D W+ + REK +
Sbjct: 451 SMNCPPFIAVELCREQIGVHPCDKRRTISEYRNMFPAIDFSLIESDEDILWESDVREKTD 630
Query: 180 EVTGRGLKFLEWLCTRKEKEIAVVTHSSFLFNTLSAFGNDCHPNIKTEMCAHFANCELRS 239
EV+ RGLKFLEWL TR+EKEIAVVTHSSFLFNTL AFGNDCHP+IK+E+C HFANCELRS
Sbjct: 631 EVSARGLKFLEWLWTREEKEIAVVTHSSFLFNTLRAFGNDCHPDIKSEICTHFANCELRS 810
Query: 240 MVIVDKCMIGSNNSTTNYPGKIPHGPDLPSDATD 273
+VI+D+ MIGS+ STTNYPGKIP GPD+PS+A D
Sbjct: 811 IVIIDRGMIGSSESTTNYPGKIPRGPDVPSEAAD 912
>TC229969 UP|Q8LGT9 (Q8LGT9) Phosphoglycerate mutase-like protein, complete
Length = 1133
Score = 447 bits (1151), Expect = e-126
Identities = 209/274 (76%), Positives = 238/274 (86%), Gaps = 1/274 (0%)
Frame = +1
Query: 1 MDTAPGQSLYPLHHSKTIHLVRHAQGVHNVEGEKNHDAYLSYDFFDANLTPLGWQQVENL 60
MDTA GQSLYPLH KT+HLVRHAQG HNVEGEKN +AY SYD FDANLTPLGW+QV+NL
Sbjct: 91 MDTAAGQSLYPLHRCKTLHLVRHAQGFHNVEGEKNFEAYKSYDLFDANLTPLGWKQVDNL 270
Query: 61 QKHVKAIGLSKKIELVVVSPLLRTMQTAVGVFGGEANTDGVNKPPLMIENVGHSDHPAVS 120
++HVKA GLSK+IELV+VSPLLRTMQTAVGVFGG+ TDG+N PPLM +NVG S PA+S
Sbjct: 271 RQHVKASGLSKRIELVIVSPLLRTMQTAVGVFGGQPYTDGINVPPLMNDNVGDSGRPAIS 450
Query: 121 SLNCPPFVAVELCREQMGLHPCDKRRTVSEYRHMFPGIDFSLIETDDDTWWKPE-REKKE 179
SLN PPF+AVELCRE +G+HPCDKRR +++YRHMFP IDFSLIE D+D WKP+ REK E
Sbjct: 451 SLNAPPFIAVELCREHLGVHPCDKRRNITDYRHMFPAIDFSLIENDEDILWKPDIREKNE 630
Query: 180 EVTGRGLKFLEWLCTRKEKEIAVVTHSSFLFNTLSAFGNDCHPNIKTEMCAHFANCELRS 239
EV RGLKFLEWL TRKEKEIAVVTHS FLF++LSAFGNDCHPN+K E+C HFANCELRS
Sbjct: 631 EVAARGLKFLEWLWTRKEKEIAVVTHSGFLFHSLSAFGNDCHPNVKNEICTHFANCELRS 810
Query: 240 MVIVDKCMIGSNNSTTNYPGKIPHGPDLPSDATD 273
MVI+D+ MIGS+ S+TNYPGK+P G DLPSD D
Sbjct: 811 MVIIDRGMIGSDESSTNYPGKVPDGLDLPSDVAD 912
>NP501847 unknown [Glycine max]
Length = 942
Score = 310 bits (793), Expect = 6e-85
Identities = 165/272 (60%), Positives = 195/272 (71%), Gaps = 3/272 (1%)
Frame = +1
Query: 1 MDTAPGQSLYPLHHSKTIHLVRHAQGVHNVEGEKNHDAYLSYDFFDANLTPLGWQQVENL 60
MDTA GQS +PLH KT+HLVRHAQG HNVEGEKN +AY SYD FDANLTPLGW QV+NL
Sbjct: 1 MDTAAGQSPHPLHRCKTLHLVRHAQGFHNVEGEKNFEAYKSYDLFDANLTPLGWNQVDNL 180
Query: 61 QKHVKAIGLSKKIELVVVSPLLRTMQTAVGVFGGEANTDGVNKPPLMIENVGHSDHPAVS 120
++HVKA GLSKKIELV+VSPLLRTMQTAVGVFGGEA TDG+N PPLM +NVG S PA+S
Sbjct: 181 REHVKASGLSKKIELVIVSPLLRTMQTAVGVFGGEAYTDGINVPPLMNDNVGDSRRPAIS 360
Query: 121 SLNCPPFVAVE-LCREQMGLHPCDKRRTVSEYRHMFPGIDFSLIETDDDTWWKPE-REKK 178
SLN PPF + L R G+ C +++ ++ + F +T P REK
Sbjct: 361 SLNVPPFNSSRALPRTFWGVSLCKEKKHHCLPTYVSQLLIFHCYKTMPTFCGNPPIREKN 540
Query: 179 EEVTGRGLKFLEWLC-TRKEKEIAVVTHSSFLFNTLSAFGNDCHPNIKTEMCAHFANCEL 237
+G + C RK+KE AVVTH FLF++L A GNDCHPN+K E+C HFANCEL
Sbjct: 541 CRSCCQGTEIFGNGCGHRKKKEKAVVTHRGFLFHSLRALGNDCHPNVKNEICTHFANCEL 720
Query: 238 RSMVIVDKCMIGSNNSTTNYPGKIPHGPDLPS 269
RSMVI+DK +IGSN S+TNY GKIP+G PS
Sbjct: 721 RSMVIIDKGVIGSNESSTNYTGKIPYGRPCPS 816
>TC221359 UP|Q8LGT9 (Q8LGT9) Phosphoglycerate mutase-like protein, partial
(52%)
Length = 1195
Score = 179 bits (454), Expect(2) = 1e-54
Identities = 83/112 (74%), Positives = 95/112 (84%), Gaps = 1/112 (0%)
Frame = +3
Query: 163 IETDDDTWWKPE-REKKEEVTGRGLKFLEWLCTRKEKEIAVVTHSSFLFNTLSAFGNDCH 221
IE D+D WKP+ REK EEV RGLKFLEWL TRKEKEIAVVTHS FLF++LSAFGNDCH
Sbjct: 723 IENDEDILWKPDIREKNEEVAARGLKFLEWLWTRKEKEIAVVTHSGFLFHSLSAFGNDCH 902
Query: 222 PNIKTEMCAHFANCELRSMVIVDKCMIGSNNSTTNYPGKIPHGPDLPSDATD 273
PN+K ++C HFANCELRSMVI+D+ MIGS+ S+TNYPGK+P G DLPSD D
Sbjct: 903 PNVKNKICTHFANCELRSMVIIDRGMIGSDESSTNYPGKVPDGLDLPSDVAD 1058
Score = 51.6 bits (122), Expect(2) = 1e-54
Identities = 21/31 (67%), Positives = 25/31 (79%)
Frame = +1
Query: 132 LCREQMGLHPCDKRRTVSEYRHMFPGIDFSL 162
L R G+HPCDKRR +++YRHMFP IDFSL
Sbjct: 634 LFRVIQGVHPCDKRRNITDYRHMFPAIDFSL 726
>AW102256
Length = 408
Score = 145 bits (365), Expect(2) = 2e-39
Identities = 69/82 (84%), Positives = 75/82 (91%)
Frame = +1
Query: 21 VRHAQGVHNVEGEKNHDAYLSYDFFDANLTPLGWQQVENLQKHVKAIGLSKKIELVVVSP 80
VRHAQG HNVEGEKN +AY SYD FDANLTPLGW QV+NL++HVKA GLSKKIELV+VSP
Sbjct: 160 VRHAQGFHNVEGEKNFEAYKSYDLFDANLTPLGWNQVDNLREHVKASGLSKKIELVIVSP 339
Query: 81 LLRTMQTAVGVFGGEANTDGVN 102
LLRTMQTAVGVFGGEA TDG+N
Sbjct: 340 LLRTMQTAVGVFGGEAYTDGIN 405
Score = 35.0 bits (79), Expect(2) = 2e-39
Identities = 15/21 (71%), Positives = 17/21 (80%)
Frame = +3
Query: 1 MDTAPGQSLYPLHHSKTIHLV 21
MDTA GQS +PLH KT+HLV
Sbjct: 66 MDTAAGQSPHPLHRCKTLHLV 128
>TC217813 similar to GB|BAB11607.1|10178227|AB025640 ZW10-like protein
{Arabidopsis thaliana;} , partial (30%)
Length = 863
Score = 110 bits (276), Expect = 5e-25
Identities = 54/84 (64%), Positives = 65/84 (77%), Gaps = 1/84 (1%)
Frame = -1
Query: 162 LIETDDDTWWKPE-REKKEEVTGRGLKFLEWLCTRKEKEIAVVTHSSFLFNTLSAFGNDC 220
LI++++DTWWK + RE KEE+ RG KF+ WL TRKEKEIA+VTH + L +TLSAFGN
Sbjct: 863 LIDSNEDTWWKADVRETKEELAARGRKFMNWLGTRKEKEIAIVTHRALLLHTLSAFGNYS 684
Query: 221 HPNIKTEMCAHFANCELRSMVIVD 244
HP K E+ FANCELRSMVIVD
Sbjct: 683 HPLEKKELSKPFANCELRSMVIVD 612
>BG839088 similar to GP|6520148|dbj| ZW10 {Arabidopsis thaliana}, partial
(16%)
Length = 691
Score = 79.0 bits (193), Expect = 2e-15
Identities = 41/91 (45%), Positives = 50/91 (54%), Gaps = 34/91 (37%)
Frame = +2
Query: 1 MDTAPGQSLYPLHHSKTIHLV----------------------------------RHAQG 26
MD G SL+PLH KTIHLV RHAQG
Sbjct: 128 MDCVAGSSLFPLHRCKTIHLVGNIHSPLFNSPHTQHNYLIRINHCFVHSPILLPVRHAQG 307
Query: 27 VHNVEGEKNHDAYLSYDFFDANLTPLGWQQV 57
+HNVEG+KN++AY++ D+FDA+LTPLGWQQV
Sbjct: 308 IHNVEGDKNYNAYINPDYFDAHLTPLGWQQV 400
>TC213860 similar to UP|Q8LGT8 (Q8LGT8) Phosphoglycerate mutase-like protein,
partial (15%)
Length = 690
Score = 56.2 bits (134), Expect = 2e-08
Identities = 33/68 (48%), Positives = 34/68 (49%), Gaps = 26/68 (38%)
Frame = +2
Query: 232 FANCELRSMVIVDK--------------------------CMIGSNNSTTNYPGKIPHGP 265
FANCELRSMVIVD+ MIG STTNYPGKIP G
Sbjct: 230 FANCELRSMVIVDRG*VCENIHKKKTGFFLS*SRFVCFFSSMIGLEQSTTNYPGKIPSGL 409
Query: 266 DLPSDATD 273
DLPSD D
Sbjct: 410 DLPSDVAD 433
>TC211557 similar to GB|BAB11607.1|10178227|AB025640 ZW10-like protein
{Arabidopsis thaliana;} , partial (20%)
Length = 740
Score = 56.2 bits (134), Expect = 2e-08
Identities = 24/31 (77%), Positives = 29/31 (93%)
Frame = +2
Query: 192 LCTRKEKEIAVVTHSSFLFNTLSAFGNDCHP 222
L T+KEKEIA+VTHS FLF+TL+AFG+DCHP
Sbjct: 644 LWTQKEKEIAIVTHSGFLFHTLNAFGSDCHP 736
Score = 42.0 bits (97), Expect = 3e-04
Identities = 17/27 (62%), Positives = 20/27 (73%), Gaps = 1/27 (3%)
Frame = +2
Query: 166 DDDTWWKPE-REKKEEVTGRGLKFLEW 191
D+DTWWK RE KEE+ RG+KFL W
Sbjct: 2 DEDTWWKANVRETKEELAARGMKFLNW 82
>BG839089 similar to GP|10178227|dbj ZW10-like protein {Arabidopsis
thaliana}, partial (12%)
Length = 616
Score = 47.0 bits (110), Expect = 9e-06
Identities = 21/33 (63%), Positives = 25/33 (75%)
Frame = -1
Query: 105 PLMIENVGHSDHPAVSSLNCPPFVAVELCREQM 137
PLM+ N G+S A+SSLN PP VAVELCRE +
Sbjct: 601 PLMVANAGNSYRAAISSLNSPPVVAVELCREHL 503
>BI471747
Length = 421
Score = 34.3 bits (77), Expect = 0.062
Identities = 14/23 (60%), Positives = 19/23 (81%)
Frame = +3
Query: 16 KTIHLVRHAQGVHNVEGEKNHDA 38
KT VRH QG+HNVEG+K+++A
Sbjct: 351 KTRIQVRHGQGIHNVEGDKDYNA 419
>BM521843
Length = 428
Score = 31.2 bits (69), Expect = 0.53
Identities = 24/86 (27%), Positives = 37/86 (42%)
Frame = +1
Query: 8 SLYPLHHSKTIHLVRHAQGVHNVEGEKNHDAYLSYDFFDANLTPLGWQQVENLQKHVKAI 67
S P+ +K + LVRH Q N EG + S LT G Q E ++ +
Sbjct: 160 SFPPIRAAKRVVLVRHGQSTWNAEGRIQGSSNFSV------LTKKGESQAETSRQML--- 312
Query: 68 GLSKKIELVVVSPLLRTMQTAVGVFG 93
+ + SPL R+ +TA ++G
Sbjct: 313 -IDDHFDACFASPLARSKRTAEIIWG 387
>TC228137 similar to UP|Q8S902 (Q8S902) Syringolide-induced protein 19-1-5,
partial (89%)
Length = 1115
Score = 30.4 bits (67), Expect = 0.90
Identities = 12/24 (50%), Positives = 16/24 (66%)
Frame = +2
Query: 29 NVEGEKNHDAYLSYDFFDANLTPL 52
N+ GEKN + SY F+D N+ PL
Sbjct: 914 NISGEKNKATHSSYIFYDQNVIPL 985
>TC226742 similar to UP|ERO2_ARATH (Q7X9I4) Endoplasmic oxidoreductin 2
precursor , partial (48%)
Length = 984
Score = 29.3 bits (64), Expect = 2.0
Identities = 16/43 (37%), Positives = 21/43 (48%), Gaps = 7/43 (16%)
Frame = +2
Query: 116 HPAVSSLNCPPFVAVELCREQ-------MGLHPCDKRRTVSEY 151
H AV S NCP + + ELC+E+ GLH S+Y
Sbjct: 2 HQAVYSENCPKYPSQELCQEEKILYKLISGLHSSISIHIASDY 130
>TC225475 UP|O22124 (O22124) Proton pyrophosphatase , partial (18%)
Length = 739
Score = 28.9 bits (63), Expect = 2.6
Identities = 22/84 (26%), Positives = 41/84 (48%), Gaps = 11/84 (13%)
Frame = +1
Query: 168 DTWWKPEREKKEEVTGR--GLKFLEWLCTRKEKEIAV---------VTHSSFLFNTLSAF 216
DTWW ++ K+ + R + FL+W KEK + + +TH S F+ F
Sbjct: 385 DTWWPTLQDLKDSIEKRASSINFLQW--QNKEKPVPLPVPSLFC*SLTHLSCQFH---CF 549
Query: 217 GNDCHPNIKTEMCAHFANCELRSM 240
N P+++T++ F +C +++
Sbjct: 550 *NLSIPHLETKV--*FVSCSFKTL 615
>TC226612 similar to UP|Q8H0V8 (Q8H0V8) Spliceosome associated protein-like
(At4g21660), partial (17%)
Length = 745
Score = 28.9 bits (63), Expect = 2.6
Identities = 14/38 (36%), Positives = 17/38 (43%)
Frame = -2
Query: 105 PLMIENVGHSDHPAVSSLNCPPFVAVELCREQMGLHPC 142
P N H ++SSL CPPF+A L PC
Sbjct: 408 PAKETN*AHPKT*SLSSLTCPPFLASFFSSSSRSLQPC 295
>BI701533
Length = 421
Score = 27.7 bits (60), Expect = 5.8
Identities = 12/21 (57%), Positives = 13/21 (61%)
Frame = -2
Query: 10 YPLHHSKTIHLVRHAQGVHNV 30
YP HHS+ IHL AQ V V
Sbjct: 132 YPNHHSRMIHLFSSAQSVMKV 70
>BE821355
Length = 596
Score = 27.7 bits (60), Expect = 5.8
Identities = 11/31 (35%), Positives = 19/31 (60%)
Frame = -3
Query: 225 KTEMCAHFANCELRSMVIVDKCMIGSNNSTT 255
+ E+C H CEL+ + ++ K M+ SN T+
Sbjct: 159 RCEVCGHCLKCELKILAMLLKSMLFSNIGTS 67
>AW350366 similar to GP|25082966|gb| spliceosome associated protein - like
{Arabidopsis thaliana}, partial (11%)
Length = 698
Score = 26.9 bits (58), Expect = 9.9
Identities = 13/38 (34%), Positives = 16/38 (41%)
Frame = +2
Query: 105 PLMIENVGHSDHPAVSSLNCPPFVAVELCREQMGLHPC 142
P N H ++ SL CPPF+A L PC
Sbjct: 299 PAKETN*AHPKT*SLCSLTCPPFLASFFSSSSRSLQPC 412
>TC210634 homologue to UP|Q6PV95 (Q6PV95) Beta-carotene hydroxylase, partial
(52%)
Length = 689
Score = 26.9 bits (58), Expect = 9.9
Identities = 10/25 (40%), Positives = 17/25 (68%)
Frame = -1
Query: 183 GRGLKFLEWLCTRKEKEIAVVTHSS 207
G+GL+FL +CT+++ + VV S
Sbjct: 170 GKGLRFLGSICTKRQSTVCVVCDRS 96
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.318 0.135 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,980,187
Number of Sequences: 63676
Number of extensions: 205956
Number of successful extensions: 1006
Number of sequences better than 10.0: 40
Number of HSP's better than 10.0 without gapping: 991
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 997
length of query: 273
length of database: 12,639,632
effective HSP length: 96
effective length of query: 177
effective length of database: 6,526,736
effective search space: 1155232272
effective search space used: 1155232272
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 58 (26.9 bits)
Medicago: description of AC146866.12


