

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
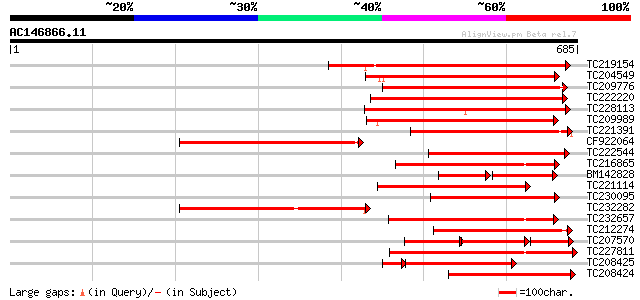
Query= AC146866.11 - phase: 0 /pseudo
(685 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC219154 similar to PIR|T05754|T05754 S-receptor kinase M4I22.1... 334 9e-92
TC204549 similar to UP|Q9M0X5 (Q9M0X5) Receptor protein kinase-l... 311 8e-85
TC209776 similar to UP|O65470 (O65470) Serine/threonine kinase-l... 290 1e-78
TC222220 weakly similar to UP|Q9ZP16 (Q9ZP16) Receptor-like prot... 286 2e-77
TC228113 similar to UP|Q9XED4 (Q9XED4) Receptor-like protein kin... 285 6e-77
TC209989 similar to UP|Q9XED4 (Q9XED4) Receptor-like protein kin... 273 2e-73
TC221391 similar to UP|O81906 (O81906) Serine/threonine kinase-l... 265 4e-71
CF922064 243 3e-64
TC222544 weakly similar to UP|O23743 (O23743) SFR1 protein precu... 230 2e-60
TC216865 similar to UP|Q9LPG0 (Q9LPG0) T3F20.24 protein, partial... 226 3e-59
BM142828 similar to GP|836954|gb|A receptor protein kinase {Ipom... 143 9e-59
TC221114 similar to UP|Q8RZ15 (Q8RZ15) Receptor kinase-like prot... 217 2e-56
TC230095 weakly similar to UP|Q9XED4 (Q9XED4) Receptor-like prot... 216 3e-56
TC232282 similar to UP|Q84XK3 (Q84XK3) S-related kinase 8 (Fragm... 215 6e-56
TC232657 weakly similar to UP|O22580 (O22580) Receptor-like seri... 212 5e-55
TC212274 similar to PIR|T05754|T05754 S-receptor kinase M4I22.1... 211 1e-54
TC207570 weakly similar to UP|O65470 (O65470) Serine/threonine k... 98 2e-53
TC227811 similar to PIR|JQ1674|JQ1674 protein kinase TMK1 , rece... 201 7e-52
TC208425 serine/threonine kinase-related protein [Glycine max] 182 2e-51
TC208424 weakly similar to UP|O65466 (O65466) Serine/threonine k... 199 3e-51
>TC219154 similar to PIR|T05754|T05754 S-receptor kinase M4I22.110 precursor
- Arabidopsis thaliana {Arabidopsis thaliana;} ,
partial (31%)
Length = 1363
Score = 334 bits (856), Expect = 9e-92
Identities = 181/332 (54%), Positives = 216/332 (64%), Gaps = 40/332 (12%)
Frame = +1
Query: 386 SCTAYTYLDPNGAVSGCSLWFNDLIDLRL-SQSSEGDDLYIRVD---------------- 428
SC AYT + +GA SGC +WF DL+D++L S + G L+IR+
Sbjct: 1 SCMAYTNYNISGAGSGCVMWFGDLLDIKLYSVAESGRRLHIRLPPSELESIKSKKSSKII 180
Query: 429 -----------------------RDSNFGKKERDGGEHEDFDLPFFDLATIIKATDNFST 465
D + KK D + +D D+P FD+ TI ATDNF
Sbjct: 181 IGTSVAAPLGVVLAICFIYRRNIADKSKTKKSIDR-QLQDVDVPLFDMLTITAATDNFLL 357
Query: 466 NNKLGEGGFGPVYKATLQDGHVIAVKRLSGNSEQGSKEFKNEVILCVKLQHRNLVKVLGC 525
NNK+GEGGFGPVYK L G IAVKRLS S QG EF EV L KLQHRNLVK+LGC
Sbjct: 358 NNKIGEGGFGPVYKGKLVGGQEIAVKRLSSLSGQGITEFITEVKLIAKLQHRNLVKLLGC 537
Query: 526 CIEGDEKLLIYEYMPNKSLDSFLFDPTQSKLLSWSMRLNILNAIARGIQYLHQDSRLRII 585
CI+G EKLL+YEY+ N SL+SF+FD +SKLL W R NI+ IARG+ YLHQDSRLRII
Sbjct: 538 CIKGQEKLLVYEYVVNGSLNSFIFDQIKSKLLDWPRRFNIILGIARGLLYLHQDSRLRII 717
Query: 586 HRDLKASNILLDNEMDPKISDFGMARMCGGDQIEGKTRRIVGTYGYMAPEYVIHGLFSIK 645
HRDLKASN+LLD +++PKISDFGMAR GGDQ EG T R+VGTYGYMAPEY G FSIK
Sbjct: 718 HRDLKASNVLLDEKLNPKISDFGMARAFGGDQTEGNTNRVVGTYGYMAPEYAFDGNFSIK 897
Query: 646 SDVFSFGVLLLETISGKKNRTLTYHEHDHNLI 677
SDVFSFG+LLLE + G KN++L + NL+
Sbjct: 898 SDVFSFGILLLEIVCGIKNKSLCHENQTLNLV 993
>TC204549 similar to UP|Q9M0X5 (Q9M0X5) Receptor protein kinase-like protein,
partial (54%)
Length = 2032
Score = 311 bits (796), Expect = 8e-85
Identities = 154/242 (63%), Positives = 188/242 (77%), Gaps = 8/242 (3%)
Frame = +3
Query: 431 SNFGKKERDGGEHED---FDLPF-----FDLATIIKATDNFSTNNKLGEGGFGPVYKATL 482
S +K++ G E +D+P FD +TI AT+ FS +NKLGEGGFG VYK TL
Sbjct: 708 SRRARKKQQGSVKEGKTAYDIPTVDSLQFDFSTIEAATNKFSADNKLGEGGFGEVYKGTL 887
Query: 483 QDGHVIAVKRLSGNSEQGSKEFKNEVILCVKLQHRNLVKVLGCCIEGDEKLLIYEYMPNK 542
G V+AVKRLS +S QG +EFKNEV++ KLQHRNLV++LG C++G+EK+L+YEY+PNK
Sbjct: 888 SSGQVVAVKRLSKSSGQGGEEFKNEVVVVAKLQHRNLVRLLGFCLQGEEKILVYEYVPNK 1067
Query: 543 SLDSFLFDPTQSKLLSWSMRLNILNAIARGIQYLHQDSRLRIIHRDLKASNILLDNEMDP 602
SLD LFDP + + L W R I+ IARGIQYLH+DSRLRIIHRDLKASNILLD +M+P
Sbjct: 1068SLDYILFDPEKQRELDWGRRYKIIGGIARGIQYLHEDSRLRIIHRDLKASNILLDGDMNP 1247
Query: 603 KISDFGMARMCGGDQIEGKTRRIVGTYGYMAPEYVIHGLFSIKSDVFSFGVLLLETISGK 662
KISDFGMAR+ G DQ +G T RIVGTYGYMAPEY +HG FS+KSDV+SFGVLL+E +SGK
Sbjct: 1248KISDFGMARIFGVDQTQGNTSRIVGTYGYMAPEYAMHGEFSVKSDVYSFGVLLMEILSGK 1427
Query: 663 KN 664
KN
Sbjct: 1428KN 1433
>TC209776 similar to UP|O65470 (O65470) Serine/threonine kinase-like protein
, partial (47%)
Length = 1305
Score = 290 bits (742), Expect = 1e-78
Identities = 143/223 (64%), Positives = 174/223 (77%)
Frame = +2
Query: 451 FDLATIIKATDNFSTNNKLGEGGFGPVYKATLQDGHVIAVKRLSGNSEQGSKEFKNEVIL 510
FD +TI AT FS NKLGEGGFG VYK L G +AVKRLS S QG +EFKNEV +
Sbjct: 44 FDFSTIEAATQKFSEANKLGEGGFGEVYKGLLPSGQEVAVKRLSKISGQGGEEFKNEVEI 223
Query: 511 CVKLQHRNLVKVLGCCIEGDEKLLIYEYMPNKSLDSFLFDPTQSKLLSWSMRLNILNAIA 570
KLQHRNLV++LG C+EG+EK+L+YE++ NKSLD LFDP + K L W+ R I+ IA
Sbjct: 224 VAKLQHRNLVRLLGFCLEGEEKILVYEFVANKSLDYILFDPEKQKSLDWTRRYKIVEGIA 403
Query: 571 RGIQYLHQDSRLRIIHRDLKASNILLDNEMDPKISDFGMARMCGGDQIEGKTRRIVGTYG 630
RGIQYLH+DSRL+IIHRDLKASN+LLD +M+PKISDFGMAR+ G DQ + T RIVGTYG
Sbjct: 404 RGIQYLHEDSRLKIIHRDLKASNVLLDGDMNPKISDFGMARIFGVDQTQANTNRIVGTYG 583
Query: 631 YMAPEYVIHGLFSIKSDVFSFGVLLLETISGKKNRTLTYHEHD 673
YM+PEY +HG +S KSDV+SFGVL+LE ISGK+N +++E D
Sbjct: 584 YMSPEYAMHGEYSAKSDVYSFGVLILEIISGKRNS--SFYETD 706
>TC222220 weakly similar to UP|Q9ZP16 (Q9ZP16) Receptor-like protein kinase,
RLK3 precursor, partial (35%)
Length = 1304
Score = 286 bits (733), Expect = 2e-77
Identities = 142/238 (59%), Positives = 177/238 (73%)
Frame = +3
Query: 436 KERDGGEHEDFDLPFFDLATIIKATDNFSTNNKLGEGGFGPVYKATLQDGHVIAVKRLSG 495
+E G E + F L TI AT+ FS ++GEGGFG VYK L DG IAVK+LS
Sbjct: 144 RENFGEESATLESLQFGLVTIEAATNKFSYEKRIGEGGFGVVYKGVLPDGREIAVKKLSK 323
Query: 496 NSEQGSKEFKNEVILCVKLQHRNLVKVLGCCIEGDEKLLIYEYMPNKSLDSFLFDPTQSK 555
+S QG+ EFKNE++L KLQHRNLV +LG C+E EK+LIYE++ NKSLD FLFD +SK
Sbjct: 324 SSGQGANEFKNEILLIAKLQHRNLVTLLGFCLEEHEKMLIYEFVSNKSLDYFLFDSHRSK 503
Query: 556 LLSWSMRLNILNAIARGIQYLHQDSRLRIIHRDLKASNILLDNEMDPKISDFGMARMCGG 615
L+WS R I+ IA+GI YLH+ SRL++IHRDLK SN+LLD+ M+PKISDFGMAR+
Sbjct: 504 QLNWSERYKIIEGIAQGISYLHEHSRLKVIHRDLKPSNVLLDSNMNPKISDFGMARIVAI 683
Query: 616 DQIEGKTRRIVGTYGYMAPEYVIHGLFSIKSDVFSFGVLLLETISGKKNRTLTYHEHD 673
DQ++GKT RIVGTYGYM+PEY +HG FS KSDVFSFGV++LE IS K+N + +HD
Sbjct: 684 DQLQGKTNRIVGTYGYMSPEYAMHGQFSEKSDVFSFGVIVLEIISAKRNTRSVFSDHD 857
>TC228113 similar to UP|Q9XED4 (Q9XED4) Receptor-like protein kinase homolog
RK20-1, partial (53%)
Length = 1492
Score = 285 bits (728), Expect = 6e-77
Identities = 152/285 (53%), Positives = 187/285 (65%), Gaps = 36/285 (12%)
Frame = +1
Query: 429 RDSNFGKKERDGGEHEDFDLPFFDLATIIKATDNFSTNNKLGEGGFGPVYKATLQDGHVI 488
R S+ K+ D E E F+ TI AT++FS +NKLG+GGFG VY+ L DG +I
Sbjct: 52 RKSSLVKQHEDDDEIEIAQSLQFNFDTIRVATEDFSDSNKLGQGGFGAVYRGRLSDGQMI 231
Query: 489 AVKRLSGNSEQGSKEFKNEVILCVKLQHRNLVKVLGCCIEGDEKLLIYEYMPNKSLDSFL 548
AVKRLS S QG EFKNEV+L KLQHRNLV++LG C+EG E+LLIYEY+PNKSLD F+
Sbjct: 232 AVKRLSRESSQGDTEFKNEVLLVAKLQHRNLVRLLGFCLEGKERLLIYEYVPNKSLDYFI 411
Query: 549 F------------------------------------DPTQSKLLSWSMRLNILNAIARG 572
F DPT+ L+W MR I+ +ARG
Sbjct: 412 FGRS**VHN*TVALYFTSGSGQRLNIHIPLKLMAVYADPTKKAQLNWEMRYKIITGVARG 591
Query: 573 IQYLHQDSRLRIIHRDLKASNILLDNEMDPKISDFGMARMCGGDQIEGKTRRIVGTYGYM 632
+ YLH+DS LRIIHRDLKASNILL+ EM+PKI+DFGMAR+ DQ + T RIVGTYGYM
Sbjct: 592 LLYLHEDSHLRIIHRDLKASNILLNEEMNPKIADFGMARLVLMDQTQANTNRIVGTYGYM 771
Query: 633 APEYVIHGLFSIKSDVFSFGVLLLETISGKKNRTLTYHEHDHNLI 677
APEY +HG FS+KSDVFSFGVL+LE ISG KN + + E+ +L+
Sbjct: 772 APEYAMHGQFSMKSDVFSFGVLVLEIISGHKNSGIRHGENVEDLL 906
>TC209989 similar to UP|Q9XED4 (Q9XED4) Receptor-like protein kinase homolog
RK20-1, partial (32%)
Length = 738
Score = 273 bits (697), Expect = 2e-73
Identities = 140/237 (59%), Positives = 174/237 (73%), Gaps = 5/237 (2%)
Frame = +3
Query: 432 NFGKKERDGGE-----HEDFDLPFFDLATIIKATDNFSTNNKLGEGGFGPVYKATLQDGH 486
N G+ + D GE D L F+ ATI AT+NFS NKLG+GGFG VYK TL DG
Sbjct: 27 NEGEGDDDEGELANDIKTDDQLLQFEFATIKFATNNFSDANKLGQGGFGIVYKGTLSDGQ 206
Query: 487 VIAVKRLSGNSEQGSKEFKNEVILCVKLQHRNLVKVLGCCIEGDEKLLIYEYMPNKSLDS 546
IA+KRLS NS QG EFKNE++L +LQHRNLV++LG C E+LLIYE++PNKSLD
Sbjct: 207 EIAIKRLSINSNQGETEFKNEILLTGRLQHRNLVRLLGFCFARRERLLIYEFVPNKSLDY 386
Query: 547 FLFDPTQSKLLSWSMRLNILNAIARGIQYLHQDSRLRIIHRDLKASNILLDNEMDPKISD 606
F+FDP + L+ +R I+ IARG+ YLH+DSRL ++HRDLK SNILLD E++PKISD
Sbjct: 387 FIFDPNKRVNLN*EIRYKIIRGIARGLLYLHEDSRLNVVHRDLKTSNILLDGELNPKISD 566
Query: 607 FGMARMCGGDQIEGKTRRIVGTYGYMAPEYVIHGLFSIKSDVFSFGVLLLETISGKK 663
FGMAR+ +Q E T IVGT+GYMAPEY+ HG FSIKSDVFSFGV++LE + G++
Sbjct: 567 FGMARLFEINQTEASTTTIVGTFGYMAPEYIKHGQFSIKSDVFSFGVMILEIVCGQR 737
>TC221391 similar to UP|O81906 (O81906) Serine/threonine kinase-like protein
, partial (25%)
Length = 642
Score = 265 bits (678), Expect = 4e-71
Identities = 130/199 (65%), Positives = 158/199 (79%), Gaps = 3/199 (1%)
Frame = +2
Query: 485 GHVIAVKRLSGNSEQGSKEFKNEVILCVKLQHRNLVKVLGCCIEGDEKLLIYEYMPNKSL 544
G +AVKRLS S QG +EFKNE++L KLQHRNLV++LGCCI+G+EK+L+YEY+PNKSL
Sbjct: 2 GEEVAVKRLSRKSSQGLEEFKNEMVLIAKLQHRNLVRLLGCCIQGEEKILVYEYLPNKSL 181
Query: 545 DSFLFDPTQSKLLSWSMRLNILNAIARGIQYLHQDSRLRIIHRDLKASNILLDNEMDPKI 604
D FLFDP + L W+ R I+ IARG+ YLH+DSRLRIIHRDLKASNILLD M+PKI
Sbjct: 182 DCFLFDPVKQTQLDWAKRFEIIEGIARGLLYLHRDSRLRIIHRDLKASNILLDESMNPKI 361
Query: 605 SDFGMARMCGGDQIEGKTRRIVGTYGYMAPEYVIHGLFSIKSDVFSFGVLLLETISGKKN 664
SDFG+AR+ GG+Q E T R+VGTYGYM+PEY + GLFSIKSDV+SFGVLLLE +SG+KN
Sbjct: 362 SDFGLARIFGGNQNEANTNRVVGTYGYMSPEYAMEGLFSIKSDVYSFGVLLLEIMSGRKN 541
Query: 665 RTLTYHEHDHNLI---WHV 680
T D +LI WH+
Sbjct: 542 -TSFRDTDDSSLIGYAWHL 595
>CF922064
Length = 740
Score = 243 bits (619), Expect = 3e-64
Identities = 109/223 (48%), Positives = 148/223 (65%), Gaps = 1/223 (0%)
Frame = -1
Query: 206 IWKGSTKICRSGPWNPLS-SGVVGMKPNPLYDYKVVNNEDEVYYQFVLRNSSVTSIAVLN 264
+WKG+T+ RSGPW+ + SG+ + + +Y +V+N+DE Y + L + S+ S V+N
Sbjct: 740 MWKGTTQYYRSGPWDGIGFSGIPSVSSDSNTNYTIVSNKDEFYITYSLIDKSLISRVVMN 561
Query: 265 QTLLIRQRLVYVPESKIWSVYQIMPSDTCEYYNVCGANAQCTIDGSPMCQCLPGFKPKSP 324
QT RQRL + +S+ W V +P+D C+ YN+CGA C I +P C+CL GFKPKSP
Sbjct: 560 QTRYARQRLAWNIDSQTWRVSSELPTDFCDQYNICGAFGICVIGQAPACKCLDGFKPKSP 381
Query: 325 QQWNSMDWTQGCVRGGNWSCGIKNRDGFQKFVRMKLPDTTNSWINLNMTLQDCKTKCLQN 384
+ W M W QGCV WSC K RDGF KF +K+PDT SW+N NMTL +CK KC +N
Sbjct: 380 RNWTQMSWNQGCVHNQTWSCRKKGRDGFNKFSNVKVPDTRRSWVNANMTLDECKNKCWEN 201
Query: 385 CSCTAYTYLDPNGAVSGCSLWFNDLIDLRLSQSSEGDDLYIRV 427
CSCTAY D G SGC++WF+DL+D+RL ++ G DLYIR+
Sbjct: 200 CSCTAYANSDIKGGGSGCAIWFSDLLDIRLMPNA-GQDLYIRL 75
>TC222544 weakly similar to UP|O23743 (O23743) SFR1 protein precursor ,
partial (27%)
Length = 861
Score = 230 bits (586), Expect = 2e-60
Identities = 111/171 (64%), Positives = 136/171 (78%), Gaps = 1/171 (0%)
Frame = +3
Query: 507 EVILCVKLQHRNLVKVLGCCIEGDEKLLIYEYMPNKSLDSFLFDPTQSKLLSWSMRLNIL 566
EV++ KLQHRNLV++LGCCIE DE++L+YE+MPNKSLDSFLFDP Q K+L W R NI+
Sbjct: 3 EVVVISKLQHRNLVRLLGCCIERDEQMLVYEFMPNKSLDSFLFDPLQRKILDWKKRFNII 182
Query: 567 NAIARGIQYLHQDSRLRIIHRDLKASNILLDNEMDPKISDFGMARMC-GGDQIEGKTRRI 625
IARGI YLH+DSRLRIIHRDLKASNILLD+EM PKISDFG+AR+ GD E T+R+
Sbjct: 183 EGIARGILYLHRDSRLRIIHRDLKASNILLDDEMHPKISDFGLARIVRSGDDDEANTKRV 362
Query: 626 VGTYGYMAPEYVIHGLFSIKSDVFSFGVLLLETISGKKNRTLTYHEHDHNL 676
VGTYGYM PEY + G+FS KSDV+ VLLLE +SG++N + +E +L
Sbjct: 363 VGTYGYMPPEYAMEGIFSEKSDVYXXXVLLLEIVSGRRNTSFYNNEQSLSL 515
>TC216865 similar to UP|Q9LPG0 (Q9LPG0) T3F20.24 protein, partial (28%)
Length = 1358
Score = 226 bits (576), Expect = 3e-59
Identities = 108/199 (54%), Positives = 147/199 (73%), Gaps = 1/199 (0%)
Frame = +1
Query: 467 NKLGEGGFGPVYKATLQDGHVIAVKRLSGNSEQGSKEFKNEVILCVKLQHRNLVKVLGCC 526
NK+GEGGFGPVYK DG +IAVK+LS S QG++EF NE+ + LQH +LVK+ GCC
Sbjct: 7 NKIGEGGFGPVYKGCFSDGTLIAVKQLSSKSRQGNREFLNEIGMISALQHPHLVKLYGCC 186
Query: 527 IEGDEKLLIYEYMPNKSLDSFLFDPTQSKL-LSWSMRLNILNAIARGIQYLHQDSRLRII 585
+EGD+ LL+YEYM N SL LF + ++ L W+ R I IARG+ YLH++SRL+I+
Sbjct: 187 VEGDQLLLVYEYMENNSLARALFGAEEHQIKLDWTTRYKICVGIARGLAYLHEESRLKIV 366
Query: 586 HRDLKASNILLDNEMDPKISDFGMARMCGGDQIEGKTRRIVGTYGYMAPEYVIHGLFSIK 645
HRD+KA+N+LLD +++PKISDFG+A++ D T RI GT+GYMAPEY +HG + K
Sbjct: 367 HRDIKATNVLLDQDLNPKISDFGLAKLDEEDNTHIST-RIAGTFGYMAPEYAMHGYLTDK 543
Query: 646 SDVFSFGVLLLETISGKKN 664
+DV+SFG++ LE I+G+ N
Sbjct: 544 ADVYSFGIVALEIINGRSN 600
>BM142828 similar to GP|836954|gb|A receptor protein kinase {Ipomoea
trifida}, partial (16%)
Length = 432
Score = 143 bits (361), Expect(2) = 9e-59
Identities = 69/79 (87%), Positives = 73/79 (92%)
Frame = +3
Query: 584 IIHRDLKASNILLDNEMDPKISDFGMARMCGGDQIEGKTRRIVGTYGYMAPEYVIHGLFS 643
IIHRDLKASNILLDN M+PKISDFG+A+MCGGDQ+EG T RIVGTYGYMAPEY I GLFS
Sbjct: 195 IIHRDLKASNILLDNNMNPKISDFGLAKMCGGDQVEGNTNRIVGTYGYMAPEYAIDGLFS 374
Query: 644 IKSDVFSFGVLLLETISGK 662
IKSDVFSFGVLLLE ISGK
Sbjct: 375 IKSDVFSFGVLLLEIISGK 431
Score = 102 bits (255), Expect(2) = 9e-59
Identities = 47/63 (74%), Positives = 54/63 (85%)
Frame = +1
Query: 519 LVKVLGCCIEGDEKLLIYEYMPNKSLDSFLFDPTQSKLLSWSMRLNILNAIARGIQYLHQ 578
LVKVLGCC+ +EK+L+YEYMPN+SLDSF+FDP QSKLL W R NIL AIARG+ YLHQ
Sbjct: 1 LVKVLGCCV*REEKMLLYEYMPNRSLDSFIFDPAQSKLLDWPTRFNILCAIARGLLYLHQ 180
Query: 579 DSR 581
DSR
Sbjct: 181 DSR 189
>TC221114 similar to UP|Q8RZ15 (Q8RZ15) Receptor kinase-like protein, partial
(29%)
Length = 946
Score = 217 bits (552), Expect = 2e-56
Identities = 106/185 (57%), Positives = 131/185 (70%)
Frame = +3
Query: 445 DFDLPFFDLATIIKATDNFSTNNKLGEGGFGPVYKATLQDGHVIAVKRLSGNSEQGSKEF 504
++++ F I AT NFS NKLG+GGFGPVYK L DG IA+KRLS S QG EF
Sbjct: 390 NYEMQIFSFPIIAAATGNFSVANKLGQGGFGPVYKGVLPDGQEIAIKRLSSRSGQGLVEF 569
Query: 505 KNEVILCVKLQHRNLVKVLGCCIEGDEKLLIYEYMPNKSLDSFLFDPTQSKLLSWSMRLN 564
KNE L KLQH NLV++ G CI+ +E +LIYEY+PNKSLD LFD + + + W R N
Sbjct: 570 KNEAELVAKLQHTNLVRLSGLCIQNEENILIYEYLPNKSLDFHLFDSKRREKIVWEKRFN 749
Query: 565 ILNAIARGIQYLHQDSRLRIIHRDLKASNILLDNEMDPKISDFGMARMCGGDQIEGKTRR 624
I+ IA G+ YLH SRL++IHRDLKA NILLD EM+PKISDFGMA + + +E KT+R
Sbjct: 750 IIEGIAHGLIYLHHFSRLKVIHRDLKAGNILLDYEMNPKISDFGMAVILDSEVVEVKTKR 929
Query: 625 IVGTY 629
+VGTY
Sbjct: 930 VVGTY 944
>TC230095 weakly similar to UP|Q9XED4 (Q9XED4) Receptor-like protein kinase
homolog RK20-1, partial (30%)
Length = 819
Score = 216 bits (549), Expect = 3e-56
Identities = 103/156 (66%), Positives = 127/156 (81%)
Frame = +3
Query: 509 ILCVKLQHRNLVKVLGCCIEGDEKLLIYEYMPNKSLDSFLFDPTQSKLLSWSMRLNILNA 568
+L KLQH+NLV+++G C E EK+LIYEY+PNKSLD FLFD + + L+WS R I+
Sbjct: 6 LLIAKLQHKNLVRLVGFCQEDREKILIYEYVPNKSLDHFLFDSQKHRQLTWSERFKIVKG 185
Query: 569 IARGIQYLHQDSRLRIIHRDLKASNILLDNEMDPKISDFGMARMCGGDQIEGKTRRIVGT 628
IARGI YLH+DSRL+IIHRD+K SN+LLD ++PKISDFGMARM DQI+G T R+VGT
Sbjct: 186 IARGILYLHEDSRLKIIHRDIKPSNVLLDYGINPKISDFGMARMVATDQIQGCTNRVVGT 365
Query: 629 YGYMAPEYVIHGLFSIKSDVFSFGVLLLETISGKKN 664
YGYM+PEY +HG FS KSDVFSFGV++L+ ISGKKN
Sbjct: 366 YGYMSPEYAMHGQFSEKSDVFSFGVMVLDIISGKKN 473
>TC232282 similar to UP|Q84XK3 (Q84XK3) S-related kinase 8 (Fragment),
partial (10%)
Length = 1030
Score = 215 bits (547), Expect = 6e-56
Identities = 107/236 (45%), Positives = 152/236 (64%), Gaps = 5/236 (2%)
Frame = +3
Query: 206 IWKGSTKICRSGPWNPLS-SGVVGMKPNPLYDYKVVNNEDEVYYQFVLRNSSVTSIAVLN 264
+ KG+ K R GPWN L SG NP+Y YK V+N++E+YY++ L+N+S+ S V+N
Sbjct: 3 LMKGNKKYQRVGPWNGLQFSGGRPKINNPVYLYKFVSNKEEIYYEWTLKNASLLSKLVVN 182
Query: 265 QTLLIRQRLVYVPESKIWSVYQIMPSDTCEYYNVCGANAQCTIDGSPMCQCLPGFKPKSP 324
QT R R V+ +K W Y P D C++Y +CGAN C+ PMC+CL G+KP+SP
Sbjct: 183 QTAQDRSRYVWSETTKSWGFYSTRPEDPCDHYGICGANEYCSPSVLPMCECLKGYKPESP 362
Query: 325 QQWNSMDWTQGCVRGGNWSCGIKNRDGFQKFVRMKLPDTTNSWINLNMTLQDCKTKCLQN 384
++WNSMD TQGCV SC DGF R+K+PDT ++++ ++ L+ CKTKCL++
Sbjct: 363 EKWNSMDRTQGCVLKHPLSC---KDDGFAPLDRLKVPDTKRTYVDESIDLEQCKTKCLKD 533
Query: 385 CSCTAYTYLDPNGAVSGCSLWFNDLIDLRLSQSSE-GDDLYIRV---DRDSNFGKK 436
CSC AYT + +GA SGC +WF +L D++L E G LYIR+ + +SN+ KK
Sbjct: 534 CSCMAYTNTNISGAGSGCVMWFGELFDIKLFPDRESGQRLYIRLPPSELESNWHKK 701
>TC232657 weakly similar to UP|O22580 (O22580) Receptor-like serine/threonine
kinase, partial (35%)
Length = 1208
Score = 212 bits (539), Expect = 5e-55
Identities = 106/206 (51%), Positives = 144/206 (69%)
Frame = +3
Query: 458 KATDNFSTNNKLGEGGFGPVYKATLQDGHVIAVKRLSGNSEQGSKEFKNEVILCVKLQHR 517
KAT+ F+ NKLG+GG G VYK + DG+ +A+KRLS N+ Q ++ F EV L + H+
Sbjct: 195 KATNYFNEANKLGQGGSGSVYKGVMPDGNTVAIKRLSYNTTQWAEHFFTEVNLISGIHHK 374
Query: 518 NLVKVLGCCIEGDEKLLIYEYMPNKSLDSFLFDPTQSKLLSWSMRLNILNAIARGIQYLH 577
NLVK+LGC I G E LL+YEY+PN+SL S+ L+W MR I+ IA G+ YLH
Sbjct: 375 NLVKLLGCSITGPESLLVYEYVPNQSLHDHFSVRRTSQPLTWEMRQKIILGIAEGMAYLH 554
Query: 578 QDSRLRIIHRDLKASNILLDNEMDPKISDFGMARMCGGDQIEGKTRRIVGTYGYMAPEYV 637
++S +RIIHRD+K SNILL+ + PKI+DFG+AR+ D+ T I GT GYMAPEY+
Sbjct: 555 EESHVRIIHRDIKLSNILLEEDFTPKIADFGLARLFPEDKSHIST-AIAGTLGYMAPEYI 731
Query: 638 IHGLFSIKSDVFSFGVLLLETISGKK 663
+ G + K+DV+SFGVL++E +SGKK
Sbjct: 732 VRGKLTEKADVYSFGVLVIEIVSGKK 809
>TC212274 similar to PIR|T05754|T05754 S-receptor kinase M4I22.110 precursor
- Arabidopsis thaliana {Arabidopsis thaliana;} ,
partial (20%)
Length = 553
Score = 211 bits (536), Expect = 1e-54
Identities = 101/168 (60%), Positives = 131/168 (77%)
Frame = +2
Query: 513 KLQHRNLVKVLGCCIEGDEKLLIYEYMPNKSLDSFLFDPTQSKLLSWSMRLNILNAIARG 572
KLQHRNLV++LG CI+ +E+LL+YE+M +KSLDSF+FD +S LL R I+N +ARG
Sbjct: 5 KLQHRNLVRLLGYCIQSEERLLVYEFMASKSLDSFIFDENKSMLLDRPTRSLIINGVARG 184
Query: 573 IQYLHQDSRLRIIHRDLKASNILLDNEMDPKISDFGMARMCGGDQIEGKTRRIVGTYGYM 632
+ YLHQDSR I+HRDLKA N+LLD+EM+PKISD G+AR GG++IE T+ +VGTYGY+
Sbjct: 185 LLYLHQDSRHTIVHRDLKAGNVLLDSEMNPKISDSGLARSFGGNEIEATTKHVVGTYGYL 364
Query: 633 APEYVIHGLFSIKSDVFSFGVLLLETISGKKNRTLTYHEHDHNLIWHV 680
PEY+I G +S KSDVFSFGVL+LE +SGK+N+ H NL+ HV
Sbjct: 365 PPEYIIDGAYSTKSDVFSFGVLILEIVSGKRNKGFC---HQDNLLAHV 499
>TC207570 weakly similar to UP|O65470 (O65470) Serine/threonine kinase-like
protein , partial (30%)
Length = 1285
Score = 98.2 bits (243), Expect(3) = 2e-53
Identities = 49/82 (59%), Positives = 60/82 (72%)
Frame = +1
Query: 547 FLFDPTQSKLLSWSMRLNILNAIARGIQYLHQDSRLRIIHRDLKASNILLDNEMDPKISD 606
F D + LL W RLNI+ IA+G+ YLH+ SRL++IHRDLKASNILLD+EM+ KISD
Sbjct: 295 FKTDSARKDLLDWEKRLNIIGGIAQGLLYLHKYSRLKVIHRDLKASNILLDHEMNAKISD 474
Query: 607 FGMARMCGGDQIEGKTRRIVGT 628
FGMAR+ G E T R+VGT
Sbjct: 475 FGMARIFGVRVSEENTNRVVGT 540
Score = 84.7 bits (208), Expect(3) = 2e-53
Identities = 41/72 (56%), Positives = 52/72 (71%)
Frame = +2
Query: 478 YKATLQDGHVIAVKRLSGNSEQGSKEFKNEVILCVKLQHRNLVKVLGCCIEGDEKLLIYE 537
++ L D +A+KRLS +S QG EF NE L KLQH NLVK+LG CI+ DE++L+YE
Sbjct: 2 HEGNLSDQQEVAIKRLSKSSGQGLIEFTNEAKLMAKLQHTNLVKLLGFCIQRDERILVYE 181
Query: 538 YMPNKSLDSFLF 549
YM NKSLD +LF
Sbjct: 182 YMSNKSLDFYLF 217
Score = 66.6 bits (161), Expect(3) = 2e-53
Identities = 32/52 (61%), Positives = 40/52 (76%)
Frame = +2
Query: 630 GYMAPEYVIHGLFSIKSDVFSFGVLLLETISGKKNRTLTYHEHDHNLIWHVS 681
GYMAPEY + G+ SIK+DVFSFGVLLLE +S KKN + + +H NLI +VS
Sbjct: 629 GYMAPEYAMKGVVSIKTDVFSFGVLLLEILSSKKNNSRYHSDHPLNLIGYVS 784
>TC227811 similar to PIR|JQ1674|JQ1674 protein kinase TMK1 , receptor type
precursor - Arabidopsis thaliana {Arabidopsis thaliana;}
, partial (40%)
Length = 1436
Score = 201 bits (512), Expect = 7e-52
Identities = 108/231 (46%), Positives = 152/231 (65%), Gaps = 5/231 (2%)
Frame = +2
Query: 460 TDNFSTNNKLGEGGFGPVYKATLQDGHVIAVKRLSGNS--EQGSKEFKNEVILCVKLQHR 517
TDNFS N LG+GGFG VY+ L DG IAVKR+ + +G+ EFK+E+ + K++HR
Sbjct: 101 TDNFSEKNVLGQGGFGTVYRGELHDGTRIAVKRMECGAIAGKGAAEFKSEIAVLTKVRHR 280
Query: 518 NLVKVLGCCIEGDEKLLIYEYMPNKSLDSFLFDPTQSKL--LSWSMRLNILNAIARGIQY 575
+LV +LG C++G+EKLL+YEYMP +L LFD + L L W+ RL I +ARG++Y
Sbjct: 281 HLVSLLGYCLDGNEKLLVYEYMPQGTLSRHLFDWPEEGLEPLEWNRRLTIALDVARGVEY 460
Query: 576 LHQDSRLRIIHRDLKASNILLDNEMDPKISDFGMARMCGGDQIEGKTRRIVGTYGYMAPE 635
LH + IHRDLK SNILL ++M K++DFG+ R+ + +T RI GT+GY+APE
Sbjct: 461 LHGLAHQSFIHRDLKPSNILLGDDMRAKVADFGLVRLAPEGKASIET-RIAGTFGYLAPE 637
Query: 636 YVIHGLFSIKSDVFSFGVLLLETISGKKNRTLTYHEHDHNLI-WHVSTNMN 685
Y + G + K DVFSFGV+L+E I+G+K T E +L+ W ++N
Sbjct: 638 YAVTGRVTTKVDVFSFGVILMELITGRKALDETQPEDSMHLVTWFRRMSIN 790
>TC208425 serine/threonine kinase-related protein [Glycine max]
Length = 763
Score = 182 bits (463), Expect(2) = 2e-51
Identities = 90/134 (67%), Positives = 106/134 (78%)
Frame = +1
Query: 479 KATLQDGHVIAVKRLSGNSEQGSKEFKNEVILCVKLQHRNLVKVLGCCIEGDEKLLIYEY 538
K L G IAVKRLS S QG+ EF+NE L KLQHRNLV++LG C+EG EK+LIYEY
Sbjct: 358 KGVLPSGQEIAVKRLSVTSLQGAVEFRNEAALVAKLQHRNLVRLLGFCLEGQEKILIYEY 537
Query: 539 MPNKSLDSFLFDPTQSKLLSWSMRLNILNAIARGIQYLHQDSRLRIIHRDLKASNILLDN 598
+PNKSLD FLFDP + K L WS R I+ IARGIQYLH+DS+LRII+RDLKASN+LLD
Sbjct: 538 IPNKSLDYFLFDPAKQKELDWSRRYKIIVGIARGIQYLHEDSQLRIIYRDLKASNVLLDE 717
Query: 599 EMDPKISDFGMARM 612
M+PKISDFGMA++
Sbjct: 718 NMNPKISDFGMAKI 759
Score = 39.3 bits (90), Expect(2) = 2e-51
Identities = 17/29 (58%), Positives = 21/29 (71%)
Frame = +2
Query: 451 FDLATIIKATDNFSTNNKLGEGGFGPVYK 479
FDL T+ AT+ FS NK+G+GGFG YK
Sbjct: 272 FDLPTVEAATNRFSDENKIGQGGFGXGYK 358
>TC208424 weakly similar to UP|O65466 (O65466) Serine/threonine kinase-like
protein , partial (34%)
Length = 913
Score = 199 bits (507), Expect = 3e-51
Identities = 98/153 (64%), Positives = 116/153 (75%)
Frame = +2
Query: 531 EKLLIYEYMPNKSLDSFLFDPTQSKLLSWSMRLNILNAIARGIQYLHQDSRLRIIHRDLK 590
EK+LIYEY+ NKSLD FLFDP + K L WS R I+ IARGIQYLH+DS+LRIIHRDLK
Sbjct: 2 EKILIYEYITNKSLDHFLFDPVKQKELDWSRRYKIIVGIARGIQYLHEDSQLRIIHRDLK 181
Query: 591 ASNILLDNEMDPKISDFGMARMCGGDQIEGKTRRIVGTYGYMAPEYVIHGLFSIKSDVFS 650
ASN+LLD M+PKISDFGMA++ DQ + T RIVGTYGYM+PEY + G FS+KSDVFS
Sbjct: 182 ASNVLLDENMNPKISDFGMAKIFQADQTQVNTGRIVGTYGYMSPEYAMRGQFSVKSDVFS 361
Query: 651 FGVLLLETISGKKNRTLTYHEHDHNLIWHVSTN 683
FGVL+LE +SGKKN H +L+ H N
Sbjct: 362 FGVLVLEIVSGKKNTDFYQSNHADDLLSHAWKN 460
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.319 0.137 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 34,682,423
Number of Sequences: 63676
Number of extensions: 550086
Number of successful extensions: 4416
Number of sequences better than 10.0: 940
Number of HSP's better than 10.0 without gapping: 3754
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3824
length of query: 685
length of database: 12,639,632
effective HSP length: 104
effective length of query: 581
effective length of database: 6,017,328
effective search space: 3496067568
effective search space used: 3496067568
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 62 (28.5 bits)
Medicago: description of AC146866.11


