

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146864.6 - phase: 0 /pseudo
(180 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
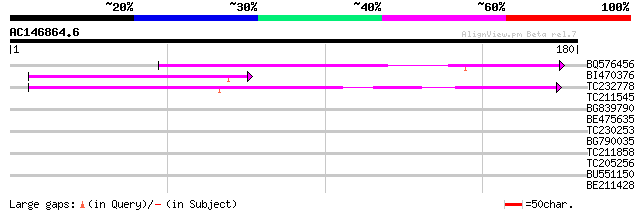
Score E
Sequences producing significant alignments: (bits) Value
BQ576456 74 5e-14
BI470376 weakly similar to GP|14090338|dbj P0638D12.5 {Oryza sat... 62 2e-10
TC232778 similar to UP|Q9LNG5 (Q9LNG5) F21D18.16, partial (5%) 61 2e-10
TC211545 weakly similar to UP|Q7XHQ1 (Q7XHQ1) Calcineurin-like p... 37 0.007
BG839790 similar to PIR|D96521|D96 protein F21D18.16 [imported] ... 33 0.009
BE475635 30 0.81
TC230253 28 1.8
BG790035 similar to GP|6319167|gb|A branched-chain amino acid am... 27 4.0
TC211858 27 4.0
TC205256 similar to UP|BCA2_ARATH (Q9M439) Branched-chain-amino-... 27 4.0
BU551150 27 5.2
BE211428 27 6.8
>BQ576456
Length = 424
Score = 73.6 bits (179), Expect = 5e-14
Identities = 46/133 (34%), Positives = 68/133 (50%), Gaps = 4/133 (3%)
Frame = +1
Query: 48 GEMTITLDDVSCLLHIPVGGNLLFHESLSIHQGTQYLVNYLGLEFEESAAQTKRLRSAHI 107
GE+TITLDD++CLLH+P+ G L E L + + L+ L + EE+ A+ R A++
Sbjct: 31 GELTITLDDMACLLHLPITGALHRFEPLGVDEAVLLLMELLEVSREEARAKIVRAHRAYV 210
Query: 108 TYDTLLSIYTSYLTEAKSYANQPEEEDSMEWYRTRC----IRAFLLYLVGCTLFSDKAGN 163
L +Y S + RC +RA+L +LV CTLF++K+
Sbjct: 211 RLSWLREVYQSRC-------------------QARCWIVAVRAYLFHLVDCTLFANKSAT 333
Query: 164 SCCVVYLKYFDDL 176
VV+LK F DL
Sbjct: 334 HVHVVHLKGF*DL 372
>BI470376 weakly similar to GP|14090338|dbj P0638D12.5 {Oryza sativa
(japonica cultivar-group)}, partial (5%)
Length = 429
Score = 61.6 bits (148), Expect = 2e-10
Identities = 36/82 (43%), Positives = 47/82 (56%), Gaps = 11/82 (13%)
Frame = -1
Query: 7 VIRASGLYLLLETNYGQVDHGLLIAFSERWHSETSSFHLPVGEMTITLDDVSCLLHIPVG 66
+++ SGLY +++ +V+ L+ AF ERW ET +FHL GE TITL DVS LL IPV
Sbjct: 396 LLQISGLYPIMKLAQLKVNGALVNAFIERWRPETHTFHLKYGEATITLQDVSVLLGIPVD 217
Query: 67 GN-----------LLFHESLSI 77
G LFHE L +
Sbjct: 216 GRPLIGNTNIDWFELFHELLGV 151
>TC232778 similar to UP|Q9LNG5 (Q9LNG5) F21D18.16, partial (5%)
Length = 746
Score = 61.2 bits (147), Expect = 2e-10
Identities = 48/170 (28%), Positives = 78/170 (45%), Gaps = 1/170 (0%)
Frame = -1
Query: 7 VIRASGLYLLLETNYGQVDHGLLIAFSERWHSETSSFHLPVGEMTITLDDVSCLLHIPV- 65
++R G Y +++ Y +++ L+ AF ERW ET +FHL GE TITL DV LL +
Sbjct: 545 LLRQCGFYWIMKMGYLKINAALISAFIERWRPETHTFHLRCGEATITLQDVLILLGLRTD 366
Query: 66 GGNLLFHESLSIHQGTQYLVNYLGLEFEESAAQTKRLRSAHITYDTLLSIYTSYLTEAKS 125
G L+ +L + L+ E E + K AH Y S++ +
Sbjct: 365 GAPLIGSTNLDWADLCEELLGVRPQEGEIEGSVVKLSWLAH---------YFSHINIDEG 213
Query: 126 YANQPEEEDSMEWYRTRCIRAFLLYLVGCTLFSDKAGNSCCVVYLKYFDD 175
Q + R IRA++L +G +F +K+ + + YL++ D
Sbjct: 212 NVEQLQ----------RFIRAWILRFIGGVIFVNKSSSRVSLRYLQFLRD 93
>TC211545 weakly similar to UP|Q7XHQ1 (Q7XHQ1) Calcineurin-like
phosphoesterase-like, partial (11%)
Length = 1127
Score = 36.6 bits (83), Expect = 0.007
Identities = 20/54 (37%), Positives = 30/54 (55%)
Frame = +3
Query: 17 LETNYGQVDHGLLIAFSERWHSETSSFHLPVGEMTITLDDVSCLLHIPVGGNLL 70
++ Y +++ L+ ER +T +FH+ E TITL DVS LL + V G L
Sbjct: 3 IKMGYLKINASLITVLIER*RPKTHTFHMRCRECTITLQDVSVLLGMRVDGTPL 164
>BG839790 similar to PIR|D96521|D96 protein F21D18.16 [imported] -
Arabidopsis thaliana, partial (2%)
Length = 504
Score = 33.5 bits (75), Expect(2) = 0.009
Identities = 12/39 (30%), Positives = 24/39 (60%)
Frame = -1
Query: 7 VIRASGLYLLLETNYGQVDHGLLIAFSERWHSETSSFHL 45
++R G Y +++ +Y +++ L+ ERW ET +FH+
Sbjct: 261 LLRQCGFY*IMKMSYLKINASLMTVLIERWRPETHTFHM 145
Score = 21.6 bits (44), Expect(2) = 0.009
Identities = 11/17 (64%), Positives = 12/17 (69%)
Frame = -3
Query: 49 EMTITLDDVSCLLHIPV 65
E TITL DVS LL + V
Sbjct: 130 ECTITLQDVSXLLGLRV 80
>BE475635
Length = 412
Score = 29.6 bits (65), Expect = 0.81
Identities = 12/27 (44%), Positives = 19/27 (69%)
Frame = +3
Query: 151 LVGCTLFSDKAGNSCCVVYLKYFDDLT 177
L+GCTLF++K+ V++L F DL+
Sbjct: 3 LLGCTLFANKSATHVYVIFLDAFRDLS 83
>TC230253
Length = 1217
Score = 28.5 bits (62), Expect = 1.8
Identities = 35/122 (28%), Positives = 53/122 (42%), Gaps = 19/122 (15%)
Frame = +1
Query: 70 LFHESLSIHQGTQYLVNYLGLEFEESAA-----QTKRLRSAHITYDTL--------LSIY 116
+ H S HQGT ++ G+ FE A K +++A + L L I
Sbjct: 499 ILHTSSQSHQGTTS--SFAGI-FENLATLCIDFDLKTVKNAITLFSALKACPKLQNLEIN 669
Query: 117 TSYLTEAKSYANQPEEEDSMEWYRTR----CIRAFL--LYLVGCTLFSDKAGNSCCVVYL 170
+ A Y +Q +E+D+ME+YR + CI L L ++G AG V +L
Sbjct: 670 SEINDNAVDYYDQHDEDDAMEYYRRKEPCECIDQQLKTLSIMGF------AGKKTEVEFL 831
Query: 171 KY 172
KY
Sbjct: 832 KY 837
>BG790035 similar to GP|6319167|gb|A branched-chain amino acid
aminotransferase {Solanum tuberosum}, partial (17%)
Length = 430
Score = 27.3 bits (59), Expect = 4.0
Identities = 14/36 (38%), Positives = 19/36 (51%), Gaps = 9/36 (25%)
Frame = -3
Query: 128 NQPEEEDSMEWYRTRCIRAF---------LLYLVGC 154
NQPEE S+ W RT+ +F +L +VGC
Sbjct: 197 NQPEEAISLNWERTQFFESFCSLSFC*NSVLVVVGC 90
>TC211858
Length = 524
Score = 27.3 bits (59), Expect = 4.0
Identities = 10/15 (66%), Positives = 15/15 (99%)
Frame = -2
Query: 147 FLLYLVGCTLFSDKA 161
+LL+LVGCTLF++K+
Sbjct: 523 YLLHLVGCTLFANKS 479
>TC205256 similar to UP|BCA2_ARATH (Q9M439) Branched-chain-amino-acid
aminotransferase 2, chloroplast precursor (Atbcat-2) ,
partial (86%)
Length = 1440
Score = 27.3 bits (59), Expect = 4.0
Identities = 14/36 (38%), Positives = 19/36 (51%), Gaps = 9/36 (25%)
Frame = -3
Query: 128 NQPEEEDSMEWYRTRCIRAF---------LLYLVGC 154
NQPEE S+ W RT+ +F +L +VGC
Sbjct: 196 NQPEEAISLNWERTQFFESFCSLSFC*NSVLVVVGC 89
>BU551150
Length = 482
Score = 26.9 bits (58), Expect = 5.2
Identities = 16/66 (24%), Positives = 29/66 (43%)
Frame = +2
Query: 69 LLFHESLSIHQGTQYLVNYLGLEFEESAAQTKRLRSAHITYDTLLSIYTSYLTEAKSYAN 128
+ FH S S+H +V LG + AA+ +R+ + + ++ +Y Y
Sbjct: 113 ICFHRSTSLHHQNHCMVQMLGGQSSPIAAKGRRIWTPSSLHFPMMKLYL--------YXX 268
Query: 129 QPEEED 134
+ E ED
Sbjct: 269 EKEAED 286
>BE211428
Length = 353
Score = 26.6 bits (57), Expect = 6.8
Identities = 16/56 (28%), Positives = 28/56 (49%), Gaps = 5/56 (8%)
Frame = -2
Query: 89 GLEFEESAAQTKR---LRSAHITYDTLLSIYTSYL--TEAKSYANQPEEEDSMEWY 139
G+ F S + K L + H+ Y+ + S + + TE K +A +E ++ EWY
Sbjct: 289 GVNFS*SQCKEKNCDDLNTRHMNYELVSSFFFFFFF*TEDKKHALTFKESNTSEWY 122
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.322 0.137 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,138,358
Number of Sequences: 63676
Number of extensions: 125367
Number of successful extensions: 629
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 628
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 629
length of query: 180
length of database: 12,639,632
effective HSP length: 91
effective length of query: 89
effective length of database: 6,845,116
effective search space: 609215324
effective search space used: 609215324
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 56 (26.2 bits)
Medicago: description of AC146864.6


