

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146862.7 - phase: 0
(198 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
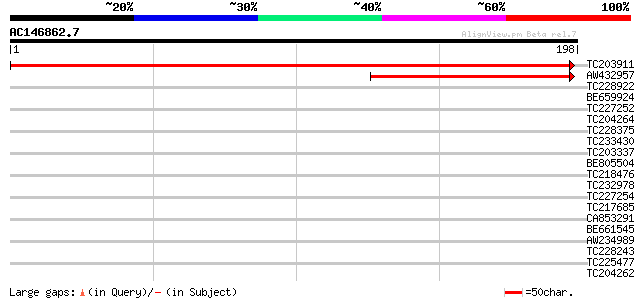
Sequences producing significant alignments: (bits) Value
TC203911 similar to UP|Q95RQ4 (Q95RQ4) LD16074p (CG6393-PA), par... 352 5e-98
AW432957 116 8e-27
TC228922 similar to UP|KADD_ARATH (Q9FIJ7) Probable adenylate ki... 32 0.15
BE659924 30 0.56
TC227252 similar to UP|Q9ZPI1 (Q9ZPI1) Lysyl-tRNA synthetase , ... 30 0.73
TC204264 similar to UP|Q8LL70 (Q8LL70) 15.8 kDa oleosin, partial... 30 0.95
TC228375 UP|Q6FYW8 (Q6FYW8) VirB3 protein homolog, partial (10%) 29 1.2
TC233430 similar to UP|Q9LU19 (Q9LU19) Gb|AAB63649.1, partial (4%) 29 1.2
TC203337 similar to UP|Q9S728 (Q9S728) En/Spm-like transposon pr... 29 1.2
BE805504 29 1.6
TC218476 homologue to UP|Q9M5K5 (Q9M5K5) Lipoamide dehydrogenase... 29 1.6
TC232978 weakly similar to UP|Q41402 (Q41402) Nodulin precursor,... 28 2.1
TC227254 homologue to UP|Q9FEL1 (Q9FEL1) Lysyl-tRNA synthetase (... 28 2.1
TC217685 similar to GB|AAM16165.1|20334818|AY094009 AT4g26750/F1... 28 2.1
CA853291 28 2.1
BE661545 28 2.1
AW234989 28 2.8
TC228243 similar to UP|Q8H6S8 (Q8H6S8) Translation initiation fa... 28 2.8
TC225477 homologue to UP|Q6R4U3 (Q6R4U3) PPase, partial (57%) 28 2.8
TC204262 weakly similar to UP|Q8LL70 (Q8LL70) 15.8 kDa oleosin, ... 28 2.8
>TC203911 similar to UP|Q95RQ4 (Q95RQ4) LD16074p (CG6393-PA), partial (12%)
Length = 2055
Score = 352 bits (904), Expect = 5e-98
Identities = 162/197 (82%), Positives = 175/197 (88%)
Frame = +1
Query: 1 MQITCEASLKCLQGKGPPFTFQCNGSSMEVFPELNNEPGNHPSGNVPEPNHRLGSEFLEP 60
MQI CEASLKCLQ KG P FQ NG+SME + EL NEPG HP+G+V EPN LGSEFLEP
Sbjct: 157 MQINCEASLKCLQSKGFPCNFQSNGNSMEGYTELKNEPGTHPAGDVAEPNCHLGSEFLEP 336
Query: 61 SNEFHNKPTYHHDYSTWTACHFHPHKMQQCQMNAFENHFYPHPVENQFQYAPINMVSQGY 120
SNEFH KPTYH +YSTWT CHF+ HK+QQCQMNAFE+H+YP+PVEN QY PINMV+QGY
Sbjct: 337 SNEFHTKPTYHQNYSTWTPCHFNSHKVQQCQMNAFESHYYPYPVENPLQYVPINMVAQGY 516
Query: 121 PREQYQEFQNFVVIDFEATCDKDKNPHPQEIIEFPSVIVSSVTGQLEACFQTYVRPTCNQ 180
PREQYQEFQ FVVIDFEATCDKDKNPHPQEIIEFPSVIVSS+TGQLEACFQTYVRPTCNQ
Sbjct: 517 PREQYQEFQYFVVIDFEATCDKDKNPHPQEIIEFPSVIVSSITGQLEACFQTYVRPTCNQ 696
Query: 181 HLSDFCKDLTGIQQIQV 197
L+DFCKDLTGIQQIQV
Sbjct: 697 LLTDFCKDLTGIQQIQV 747
>AW432957
Length = 400
Score = 116 bits (290), Expect = 8e-27
Identities = 54/71 (76%), Positives = 61/71 (85%)
Frame = +2
Query: 127 EFQNFVVIDFEATCDKDKNPHPQEIIEFPSVIVSSVTGQLEACFQTYVRPTCNQHLSDFC 186
EF++ V+IDFEATCD DKNPHPQ+I+EFPSVI S+ TG+LEAC QTYV TCNQ LSDFC
Sbjct: 2 EFESSVLIDFEATCDMDKNPHPQQIMEFPSVIESNFTGRLEACIQTYVMSTCNQLLSDFC 181
Query: 187 KDLTGIQQIQV 197
DLTGIQQI V
Sbjct: 182 MDLTGIQQIHV 214
>TC228922 similar to UP|KADD_ARATH (Q9FIJ7) Probable adenylate kinase 2,
chloroplast precursor (ATP-AMP transphosphorylase) ,
partial (75%)
Length = 1133
Score = 32.3 bits (72), Expect = 0.15
Identities = 17/58 (29%), Positives = 26/58 (44%)
Frame = -3
Query: 26 SSMEVFPELNNEPGNHPSGNVPEPNHRLGSEFLEPSNEFHNKPTYHHDYSTWTACHFH 83
SS ++ P+ N+ N S + P P+ S +F PT Y+ W A H+H
Sbjct: 501 SSWDIHPKANHSLSNLVSRDAP*PS----------SQQFRQGPTVPSPYTVWLAFHYH 358
>BE659924
Length = 588
Score = 30.4 bits (67), Expect = 0.56
Identities = 19/68 (27%), Positives = 28/68 (40%)
Frame = -2
Query: 46 VPEPNHRLGSEFLEPSNEFHNKPTYHHDYSTWTACHFHPHKMQQCQMNAFENHFYPHPVE 105
+P P R + F S H P +HH ++W C+ P Y P+
Sbjct: 470 LPAPTTRNSTGFRSRSAIPHPPPPHHHQRNSW-YCNDPP-------------THYTEPLP 333
Query: 106 NQFQYAPI 113
+QFQY P+
Sbjct: 332 SQFQYTPL 309
>TC227252 similar to UP|Q9ZPI1 (Q9ZPI1) Lysyl-tRNA synthetase , partial (59%)
Length = 1524
Score = 30.0 bits (66), Expect = 0.73
Identities = 12/27 (44%), Positives = 16/27 (58%), Gaps = 1/27 (3%)
Frame = -2
Query: 47 PEPNHR-LGSEFLEPSNEFHNKPTYHH 72
P PNH+ + + LEP FH +P HH
Sbjct: 1088 PNPNHQ*VANHTLEPYKMFHRRPLLHH 1008
>TC204264 similar to UP|Q8LL70 (Q8LL70) 15.8 kDa oleosin, partial (82%)
Length = 853
Score = 29.6 bits (65), Expect = 0.95
Identities = 16/46 (34%), Positives = 19/46 (40%), Gaps = 2/46 (4%)
Frame = +2
Query: 60 PSNEFHNKPTYHHDYST--WTACHFHPHKMQQCQMNAFENHFYPHP 103
P+ E HN Y H T W H H H+ C N + PHP
Sbjct: 191 PNREVHNCCNYWHHTLTPVWVDPHRHCHRFDHC--NPSSCYLQPHP 322
>TC228375 UP|Q6FYW8 (Q6FYW8) VirB3 protein homolog, partial (10%)
Length = 680
Score = 29.3 bits (64), Expect = 1.2
Identities = 25/86 (29%), Positives = 36/86 (41%), Gaps = 1/86 (1%)
Frame = -3
Query: 62 NEFHNKPTYHHDYSTWTACHFHPHKMQQCQMNAFENHFYPHPVENQFQYAPINMVSQGYP 121
N FH+ +HH A +FH H +Q N + H F Y+P+ Q Y
Sbjct: 489 NPFHSHIQFHHWKQIPLATNFHQHD*EQ-------NWIHLH-----FDYSPL*YYLQIYS 346
Query: 122 R-EQYQEFQNFVVIDFEATCDKDKNP 146
R EQ QE ++ + C +NP
Sbjct: 345 RKEQLQEHHSYKPL**AEYCLPFQNP 268
>TC233430 similar to UP|Q9LU19 (Q9LU19) Gb|AAB63649.1, partial (4%)
Length = 689
Score = 29.3 bits (64), Expect = 1.2
Identities = 17/59 (28%), Positives = 23/59 (38%), Gaps = 6/59 (10%)
Frame = -1
Query: 50 NHRLGSEFLEPSNEFHNKPTYHHD------YSTWTACHFHPHKMQQCQMNAFENHFYPH 102
NH F P ++ H+ P YHH Y CH + QQ ++HF H
Sbjct: 680 NHHCQVHFYSPDHDSHDDPIYHHQQSCLKRYQHALCCH*NEMTHQQ-----RDHHFLVH 519
>TC203337 similar to UP|Q9S728 (Q9S728) En/Spm-like transposon protein
(Protodermal factor 1), partial (46%)
Length = 1410
Score = 29.3 bits (64), Expect = 1.2
Identities = 18/77 (23%), Positives = 31/77 (39%)
Frame = +3
Query: 33 ELNNEPGNHPSGNVPEPNHRLGSEFLEPSNEFHNKPTYHHDYSTWTACHFHPHKMQQCQM 92
E+ +P +HP +H + P ++ H++P HH H H H + Q ++
Sbjct: 510 EVTIQPHHHPQA-ATVAHHHMTPLLQHPQSQQHHQPQQHHQP------HQHHHTLPQEEV 668
Query: 93 NAFENHFYPHPVENQFQ 109
+H V Q Q
Sbjct: 669 GTITHHQLMEVVAPQHQ 719
>BE805504
Length = 284
Score = 28.9 bits (63), Expect = 1.6
Identities = 10/34 (29%), Positives = 21/34 (61%)
Frame = -3
Query: 64 FHNKPTYHHDYSTWTACHFHPHKMQQCQMNAFEN 97
+ ++P H+ +W +CH H K+ CQ+++ +N
Sbjct: 276 YQDQPNTHNH--SWVSCHTHSDKLVFCQVHSQQN 181
>TC218476 homologue to UP|Q9M5K5 (Q9M5K5) Lipoamide dehydrogenase
(AT3g16950/K14A17_7), partial (44%)
Length = 782
Score = 28.9 bits (63), Expect = 1.6
Identities = 15/43 (34%), Positives = 20/43 (45%)
Frame = -1
Query: 71 HHDYSTWTACHFHPHKMQQCQMNAFENHFYPHPVENQFQYAPI 113
HH Y+T FH H++ C +NH Q QY+PI
Sbjct: 635 HHQYNTNEEPGFHLHQVLSCAHRQEQNH-----AALQHQYSPI 522
>TC232978 weakly similar to UP|Q41402 (Q41402) Nodulin precursor, partial
(28%)
Length = 590
Score = 28.5 bits (62), Expect = 2.1
Identities = 13/33 (39%), Positives = 17/33 (51%)
Frame = +1
Query: 40 NHPSGNVPEPNHRLGSEFLEPSNEFHNKPTYHH 72
+HP N+ NHR S L S+ +N P Y H
Sbjct: 184 SHPFTNLLSRNHRFTSHLLW*SHTIYNPPPYGH 282
>TC227254 homologue to UP|Q9FEL1 (Q9FEL1) Lysyl-tRNA synthetase (Fragment),
partial (41%)
Length = 448
Score = 28.5 bits (62), Expect = 2.1
Identities = 12/27 (44%), Positives = 16/27 (58%), Gaps = 1/27 (3%)
Frame = -3
Query: 47 PEPNHR-LGSEFLEPSNEFHNKPTYHH 72
P PNH+ + + LEP FH +P HH
Sbjct: 128 PNPNHQ*VVNHILEPYKMFHLRPLLHH 48
>TC217685 similar to GB|AAM16165.1|20334818|AY094009 AT4g26750/F10M23_90
{Arabidopsis thaliana;} , partial (57%)
Length = 1586
Score = 28.5 bits (62), Expect = 2.1
Identities = 26/110 (23%), Positives = 41/110 (36%), Gaps = 16/110 (14%)
Frame = +3
Query: 38 PGNHPSGNVPEPNHRLGSEFLEPSNEFHNKPTYHHDYSTWTACHFHPHKMQQCQMNAFEN 97
PG +PS + P PS ++H+ P +S ++ +HP Q
Sbjct: 801 PGVYPSQDYHSP---------PPSRDYHSPPPSQDYHSPPSSQDYHPPPPSQDYHPPPSQ 953
Query: 98 HFYPHPVE---------NQFQYAPINMVSQG--YPREQ-----YQEFQNF 131
++P P N QY+P N G YP + Y FQ++
Sbjct: 954 DYHPPPARSEGSYSELYNHQQYSPENSQHLGPNYPSHETSSYSYPHFQSY 1103
>CA853291
Length = 576
Score = 28.5 bits (62), Expect = 2.1
Identities = 13/41 (31%), Positives = 20/41 (48%), Gaps = 3/41 (7%)
Frame = +2
Query: 47 PEPNHRLGSEFLEPSNEFHNKPTYHH---DYSTWTACHFHP 84
P PNHR + +++ HN+ +HH D +T T P
Sbjct: 23 PPPNHRKQPNLIRNASDRHNRERHHHFLSDLATRTITPLQP 145
>BE661545
Length = 676
Score = 28.5 bits (62), Expect = 2.1
Identities = 14/58 (24%), Positives = 24/58 (41%)
Frame = +3
Query: 49 PNHRLGSEFLEPSNEFHNKPTYHHDYSTWTACHFHPHKMQQCQMNAFENHFYPHPVEN 106
PNH + ++ H+ +HHD+ H H + +NH Y H ++N
Sbjct: 180 PNHYY-DHYHNHNHHHHHHHNHHHDFDD------HDHNCGDDNQDHNDNHGYDHQIQN 332
>AW234989
Length = 407
Score = 28.1 bits (61), Expect = 2.8
Identities = 15/44 (34%), Positives = 26/44 (59%), Gaps = 4/44 (9%)
Frame = -3
Query: 131 FVVIDFEATCDKDKNPHPQEII----EFPSVIVSSVTGQLEACF 170
FV++ F+AT K N PQ+I F ++ + ++TG+L+ F
Sbjct: 327 FVMVSFKATKAKTANNTPQKIFTFIGTFLNLSIKTLTGELKLLF 196
>TC228243 similar to UP|Q8H6S8 (Q8H6S8) Translation initiation factor,
partial (34%)
Length = 1442
Score = 28.1 bits (61), Expect = 2.8
Identities = 10/24 (41%), Positives = 16/24 (66%)
Frame = -1
Query: 49 PNHRLGSEFLEPSNEFHNKPTYHH 72
P H++ S +LE +FHN+P + H
Sbjct: 911 PTHQVSSLYLEKILQFHNQPPFLH 840
>TC225477 homologue to UP|Q6R4U3 (Q6R4U3) PPase, partial (57%)
Length = 1488
Score = 28.1 bits (61), Expect = 2.8
Identities = 11/44 (25%), Positives = 22/44 (50%)
Frame = -2
Query: 145 NPHPQEIIEFPSVIVSSVTGQLEACFQTYVRPTCNQHLSDFCKD 188
N H +E + ++S+ E C+Q+ CN + ++C+D
Sbjct: 260 NTHTEEQLPALYFLLSTKVENCEGCWQSNPTNNCNSNSHEYCRD 129
>TC204262 weakly similar to UP|Q8LL70 (Q8LL70) 15.8 kDa oleosin, partial
(73%)
Length = 444
Score = 28.1 bits (61), Expect = 2.8
Identities = 16/46 (34%), Positives = 18/46 (38%), Gaps = 2/46 (4%)
Frame = +3
Query: 60 PSNEFHNKPTYHHDYST--WTACHFHPHKMQQCQMNAFENHFYPHP 103
P+ E HN Y H T W H H H+ C N H HP
Sbjct: 132 PNREVHNCCNYWHHTLTPVWVDPHRH*HRFDHC--NHCSCHLQAHP 263
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.320 0.135 0.441
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,691,117
Number of Sequences: 63676
Number of extensions: 239056
Number of successful extensions: 1361
Number of sequences better than 10.0: 62
Number of HSP's better than 10.0 without gapping: 1331
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1354
length of query: 198
length of database: 12,639,632
effective HSP length: 92
effective length of query: 106
effective length of database: 6,781,440
effective search space: 718832640
effective search space used: 718832640
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 56 (26.2 bits)
Medicago: description of AC146862.7


