

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146862.24 - phase: 0
(580 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
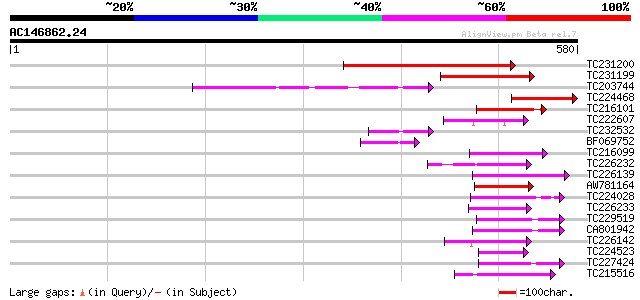
Score E
Sequences producing significant alignments: (bits) Value
TC231200 similar to UP|Q84LE6 (Q84LE6) RelA-SpoT like protein RS... 307 1e-83
TC231199 similar to UP|Q84LE6 (Q84LE6) RelA-SpoT like protein RS... 152 3e-37
TC203744 similar to UP|Q76JT3 (Q76JT3) RelA-SpoT like protein Ps... 124 2e-28
TC224468 similar to UP|Q84LE6 (Q84LE6) RelA-SpoT like protein RS... 117 1e-26
TC216101 similar to UP|Q9SCA1 (Q9SCA1) Calcium-binding protein, ... 57 2e-08
TC222607 weakly similar to UP|POC7_PHLPR (O82040) Polcalcin Phl ... 53 3e-07
TC232532 weakly similar to UP|Q8EBQ4 (Q8EBQ4) GTP pyrophosphokin... 52 6e-07
BF069752 homologue to GP|7141304|gb|A RSH1 {Arabidopsis thaliana... 52 7e-07
TC216099 similar to UP|Q9SCA1 (Q9SCA1) Calcium-binding protein, ... 52 1e-06
TC226232 weakly similar to UP|Q93YA8 (Q93YA8) Calcium binding pr... 51 2e-06
TC226139 similar to UP|Q93YA8 (Q93YA8) Calcium binding protein, ... 50 3e-06
AW781164 similar to GP|18146862|dbj| pollen-specific calcium-bin... 50 3e-06
TC224028 similar to UP|Q9AXG2 (Q9AXG2) Regulator of gene silenci... 49 8e-06
TC226233 similar to UP|TCH2_ARATH (P25070) Calmodulin-related pr... 48 1e-05
TC229519 weakly similar to UP|Q93YA8 (Q93YA8) Calcium binding pr... 48 1e-05
CA801942 48 1e-05
TC226142 similar to UP|Q93YA8 (Q93YA8) Calcium binding protein, ... 47 2e-05
TC224523 weakly similar to PIR|C96763|C96763 protein calmodulin ... 46 5e-05
TC227424 UP|Q39890 (Q39890) Calmodulin, complete 45 9e-05
TC215516 weakly similar to UP|Q8VWY6 (Q8VWY6) Pollen-specific ca... 43 3e-04
>TC231200 similar to UP|Q84LE6 (Q84LE6) RelA-SpoT like protein RSH4
(Fragment), partial (30%)
Length = 535
Score = 307 bits (786), Expect = 1e-83
Identities = 156/177 (88%), Positives = 165/177 (93%), Gaps = 1/177 (0%)
Frame = +3
Query: 342 VVLNPKSRENALEAGERACYRAHQIIQSMWKEIPSRTKDYISRPKGNGYRSLHMAVDVSE 401
V+LNPK+ ENALEAGERACYR HQIIQSMWKEIP RTKDYI+RPK NGY+SLHMAVDVSE
Sbjct: 3 VILNPKAGENALEAGERACYRTHQIIQSMWKEIPYRTKDYIARPKANGYKSLHMAVDVSE 182
Query: 402 IGRTRPLMEIQIRTTEMDRLAVGGMASHSLYKAGLTNPEEAKRLKTIMLAAAELAALRLK 461
G+TRPLMEIQIRTTEMDRLAVGG A+HSLYKAGLT+PEEAKRLKTIMLAAAELAALRLK
Sbjct: 183 NGKTRPLMEIQIRTTEMDRLAVGGTAAHSLYKAGLTDPEEAKRLKTIMLAAAELAALRLK 362
Query: 462 DFP-SANHKGIEFDQRDRVFRLLDKNGDGKISIEELTEVIEELGAPGEDAHDMMLLL 517
DFP S NHKGIE QRDRVFRLLDKNGDGKISIEELTEV+EELGAPGEDA +MM LL
Sbjct: 363 DFPTSTNHKGIEIHQRDRVFRLLDKNGDGKISIEELTEVMEELGAPGEDAREMMQLL 533
>TC231199 similar to UP|Q84LE6 (Q84LE6) RelA-SpoT like protein RSH4
(Fragment), partial (13%)
Length = 448
Score = 152 bits (385), Expect = 3e-37
Identities = 81/98 (82%), Positives = 88/98 (89%), Gaps = 1/98 (1%)
Frame = -3
Query: 441 EAKRLKTIMLAAAELAALRLKDF-PSANHKGIEFDQRDRVFRLLDKNGDGKISIEELTEV 499
+AK LKTI+LAAAELAALRLKDF S NH GIE DQRDRVFRLLDKNGDGKISIEELT+V
Sbjct: 398 QAKCLKTIILAAAELAALRLKDFLTSTNH*GIEIDQRDRVFRLLDKNGDGKISIEELTQV 219
Query: 500 IEELGAPGEDAHDMMLLLDSNSDGSLSSDEFQMFQKQV 537
+EELGAPGEDA +MM LLDSNS+G LSSDEF MFQ+QV
Sbjct: 218 MEELGAPGEDAREMMQLLDSNSNGPLSSDEFHMFQEQV 105
>TC203744 similar to UP|Q76JT3 (Q76JT3) RelA-SpoT like protein PsRSH1,
partial (54%)
Length = 1422
Score = 124 bits (310), Expect = 2e-28
Identities = 82/246 (33%), Positives = 134/246 (54%)
Frame = +1
Query: 188 AAALRKFCLTYYDIRALILDLALKLDMMRHLGHLPRYQQQIISLQVMKIYAPLAHAVGTN 247
A L L D RA+++ LA +L M L LP +QQ + + ++I+APLA+ +G +
Sbjct: 10 ADRLHTMFLGMADARAVLVKLADRLHNMMTLDALPGAKQQRFAKETLEIFAPLANRLGIS 189
Query: 248 YISLELEDLSFQYLFPYSYLYVDTWLRSQETGGISLIDVYKDELLESLKSDPILAELVDD 307
+LE+L F++L P + + + L E+ ++I + L ++LK + I ++
Sbjct: 190 TWKEQLENLCFKHLNPSQHEELSSKL--VESYDDAMITSAIERLEQALKDEGISYNVIS- 360
Query: 308 ISVKGRYKSRYSTMKKLLKDGRRPEDVNDVLGLRVVLNPKSRENALEAGERACYRAHQII 367
GR+KS YS K+LK +D++D+ GLR++++ E CY+A ++
Sbjct: 361 ----GRHKSLYSIYCKMLKKKLTIDDIHDIYGLRLIVDK----------EEDCYKALTVV 498
Query: 368 QSMWKEIPSRTKDYISRPKGNGYRSLHMAVDVSEIGRTRPLMEIQIRTTEMDRLAVGGMA 427
+W E+P + KDYI RPK NGY+SLH V +G + +E+QIRT +M A G A
Sbjct: 499 HRLWSEVPGKLKDYICRPKFNGYQSLHTVV----MGEGKVPLEVQIRTKDMHLQADFGFA 666
Query: 428 SHSLYK 433
+H YK
Sbjct: 667 AHWRYK 684
>TC224468 similar to UP|Q84LE6 (Q84LE6) RelA-SpoT like protein RSH4
(Fragment), partial (11%)
Length = 359
Score = 117 bits (293), Expect = 1e-26
Identities = 57/67 (85%), Positives = 63/67 (93%)
Frame = +1
Query: 514 MLLLDSNSDGSLSSDEFQMFQKQVEMVRNLEDRDDEYKKILDEKLHMADDSGLIQVYNKE 573
M LLDSNSDGSLSSDEF MFQ+QVE+VRNLE RDD+YKK+LDEKLHMAD+SGLIQVY+KE
Sbjct: 7 MQLLDSNSDGSLSSDEFHMFQEQVELVRNLEVRDDQYKKLLDEKLHMADESGLIQVYSKE 186
Query: 574 FGNRLVS 580
FGNRL S
Sbjct: 187 FGNRLAS 207
>TC216101 similar to UP|Q9SCA1 (Q9SCA1) Calcium-binding protein, partial
(65%)
Length = 796
Score = 57.0 bits (136), Expect = 2e-08
Identities = 29/74 (39%), Positives = 47/74 (63%), Gaps = 2/74 (2%)
Frame = +1
Query: 478 RVFRLLDKNGDGKISIEELTEVIEELG--APGEDAHDMMLLLDSNSDGSLSSDEFQMFQK 535
RVF++ D+NGDG+IS++EL++ +E LG P +D M+ +D N DG + DEF +
Sbjct: 241 RVFQMFDRNGDGRISLKELSDSLENLGILIPDKDLAQMIERIDVNGDGCVDMDEFGDLYE 420
Query: 536 QVEMVRNLEDRDDE 549
+ +E+RD+E
Sbjct: 421 SI-----MEERDEE 447
Score = 37.4 bits (85), Expect = 0.019
Identities = 22/68 (32%), Positives = 36/68 (52%), Gaps = 4/68 (5%)
Frame = +1
Query: 472 EFDQRDRVFRLLDKNGDGKISIEELTEVIEELGAPG----EDAHDMMLLLDSNSDGSLSS 527
E D R+ F + D+N DG I++EEL V+ LG ++ M+ +D + DG ++
Sbjct: 445 EEDMRE-AFNVFDQNRDGFITVEELRRVLASLGLKQGRTLDECKKMITKVDVDGDGMVNY 621
Query: 528 DEFQMFQK 535
EF+ K
Sbjct: 622 KEFRQMMK 645
>TC222607 weakly similar to UP|POC7_PHLPR (O82040) Polcalcin Phl p 7
(Calcium-binding pollen allergen Phl p 7) (P7), partial
(32%)
Length = 645
Score = 53.1 bits (126), Expect = 3e-07
Identities = 35/98 (35%), Positives = 52/98 (52%), Gaps = 11/98 (11%)
Frame = +2
Query: 444 RLKTIMLAAAELAALRLKDFPSANHKGIEF-------DQRDRVFRLLDKNGDGKISIEEL 496
R+K +M A + +K F S++ Q +VF+L+D NGDGKISI EL
Sbjct: 347 RIKEMMGACCKRRQSNIKLFSSSSDLSTSSFLHMKLSTQFHQVFKLIDTNGDGKISINEL 526
Query: 497 TEVIEELG----APGEDAHDMMLLLDSNSDGSLSSDEF 530
+E++ LG ++A M+ +LD N DG + DEF
Sbjct: 527 SELLSSLGYNKCTAVKEAEGMVKVLDFNRDGFVDLDEF 640
>TC232532 weakly similar to UP|Q8EBQ4 (Q8EBQ4) GTP pyrophosphokinase, partial
(8%)
Length = 459
Score = 52.4 bits (124), Expect = 6e-07
Identities = 28/66 (42%), Positives = 36/66 (54%)
Frame = +1
Query: 368 QSMWKEIPSRTKDYISRPKGNGYRSLHMAVDVSEIGRTRPLMEIQIRTTEMDRLAVGGMA 427
+ +W I DYI PK +GY+SLH AV G +E+QIRT M A G+A
Sbjct: 19 ERLWTPIDGEFDDYIINPKPSGYQSLHTAVQ----GPDNSPLEVQIRTQRMHECAEQGLA 186
Query: 428 SHSLYK 433
+H LYK
Sbjct: 187 AHWLYK 204
>BF069752 homologue to GP|7141304|gb|A RSH1 {Arabidopsis thaliana}, partial
(7%)
Length = 198
Score = 52.0 bits (123), Expect = 7e-07
Identities = 26/60 (43%), Positives = 35/60 (58%)
Frame = +1
Query: 360 CYRAHQIIQSMWKEIPSRTKDYISRPKGNGYRSLHMAVDVSEIGRTRPLMEIQIRTTEMD 419
CY +I +W IP KDYI+ PK NGY+SL V + + + +E+QIRT EMD
Sbjct: 16 CYHVLGLIHGIWTPIPRSVKDYIATPKPNGYQSLQTTV-IPFLYESMFRLEVQIRTEEMD 192
>TC216099 similar to UP|Q9SCA1 (Q9SCA1) Calcium-binding protein, partial
(63%)
Length = 946
Score = 51.6 bits (122), Expect = 1e-06
Identities = 28/83 (33%), Positives = 49/83 (58%), Gaps = 3/83 (3%)
Frame = +3
Query: 471 IEFDQRDRVFRLLDKNGDGKISIEELTEVIEELG--APGEDAHDMMLLLDSNSDGSLSSD 528
+E + RVF++ D+NGDG+I+ +EL + +E LG +D M+ +D N DG + D
Sbjct: 54 MEAQELKRVFQMFDRNGDGRITKKELNDSLENLGIFISDKDLSQMIQRIDVNGDGCVDMD 233
Query: 529 EF-QMFQKQVEMVRNLEDRDDEY 550
EF +++Q ++ N ED + +
Sbjct: 234 EFGELYQTIMDERDNEEDMREAF 302
>TC226232 weakly similar to UP|Q93YA8 (Q93YA8) Calcium binding protein,
partial (67%)
Length = 876
Score = 50.8 bits (120), Expect = 2e-06
Identities = 38/108 (35%), Positives = 51/108 (47%), Gaps = 2/108 (1%)
Frame = +2
Query: 428 SHSLYKAGLTNPEEAKRLKTIMLAAAELAALRLKDFPSANHKGIEFDQRDRVFRLLDKNG 487
S +YKA T+P+ EL L F S ++ + R ++F DKNG
Sbjct: 176 SRFIYKARDTSPQPV-----------ELTELNSSFFSSLGFIAMDEEVR-KIFSKFDKNG 319
Query: 488 DGKISIEELTEVIEELGA--PGEDAHDMMLLLDSNSDGSLSSDEFQMF 533
DGKIS EL E++ LG+ E+ MM LD N DG + EF F
Sbjct: 320 DGKISCAELKEMMVALGSKTTSEEVKRMMAELDRNGDGYIDLKEFGEF 463
>TC226139 similar to UP|Q93YA8 (Q93YA8) Calcium binding protein, partial
(79%)
Length = 519
Score = 50.1 bits (118), Expect = 3e-06
Identities = 32/101 (31%), Positives = 50/101 (48%), Gaps = 2/101 (1%)
Frame = +3
Query: 474 DQRDRVFRLLDKNGDGKISIEELTEVIEELGA--PGEDAHDMMLLLDSNSDGSLSSDEFQ 531
++ RVF D NGDGKIS+ EL V+ LG+ P E+ +M LD++ DG ++ EF
Sbjct: 156 EELKRVFSRFDANGDGKISVSELDNVLRSLGSGVPPEELQRVMEDLDTDHDGFINLSEFA 335
Query: 532 MFQKQVEMVRNLEDRDDEYKKILDEKLHMADDSGLIQVYNK 572
F + + D + +K + + L QV N+
Sbjct: 336 AFCRSDTADGGDTELHDAFNLYDQDKNGLISATELCQVLNR 458
>AW781164 similar to GP|18146862|dbj| pollen-specific calcium-binding protein
{Nicotiana tabacum}, partial (98%)
Length = 250
Score = 50.1 bits (118), Expect = 3e-06
Identities = 26/61 (42%), Positives = 38/61 (61%), Gaps = 1/61 (1%)
Frame = +2
Query: 476 RDRVFRLLDKNGDGKISIEELTEVIEELGA-PGEDAHDMMLLLDSNSDGSLSSDEFQMFQ 534
R+R+F+ D NGDG+IS EL E ++ LG+ E+ MM +D++ DG +S EF F
Sbjct: 29 RERIFKRFDANGDGQISSAELGEALKALGSVTAEEVKRMMEEIDTDGDGYISYQEFTEFA 208
Query: 535 K 535
K
Sbjct: 209 K 211
>TC224028 similar to UP|Q9AXG2 (Q9AXG2) Regulator of gene silencing, partial
(34%)
Length = 843
Score = 48.5 bits (114), Expect = 8e-06
Identities = 32/98 (32%), Positives = 52/98 (52%), Gaps = 2/98 (2%)
Frame = +1
Query: 472 EFDQRDRVFRLLDKNGDGKISIEELTEVIEELGA--PGEDAHDMMLLLDSNSDGSLSSDE 529
+ Q RVF D+NGD KIS EL + +E +G +DA + LLD + DG + ++
Sbjct: 289 KLSQYKRVFNQFDENGDSKISPSELRQCVEAIGGELSEKDAEVAVTLLDRDGDGLVGFED 468
Query: 530 FQMFQKQVEMVRNLEDRDDEYKKILDEKLHMADDSGLI 567
F F ++ + E+++D+ K+ K + D SG I
Sbjct: 469 FVRFLEEGKE----EEKEDDLKEAF--KRYEMDGSGCI 564
>TC226233 similar to UP|TCH2_ARATH (P25070) Calmodulin-related protein 2,
touch-induced, partial (55%)
Length = 800
Score = 48.1 bits (113), Expect = 1e-05
Identities = 27/68 (39%), Positives = 38/68 (55%), Gaps = 4/68 (5%)
Frame = +3
Query: 470 GIEFDQRD--RVFRLLDKNGDGKISIEELTEVIEELGA--PGEDAHDMMLLLDSNSDGSL 525
G+ D+ + ++F DKNGDGKIS EL E++ LG+ ++ MM LD N DG +
Sbjct: 192 GVAMDEEEVRKIFSKFDKNGDGKISCAELKEMMAALGSKTTSDEVKRMMAELDRNGDGYI 371
Query: 526 SSDEFQMF 533
EF F
Sbjct: 372 DLKEFGEF 395
Score = 37.4 bits (85), Expect = 0.019
Identities = 23/92 (25%), Positives = 43/92 (46%)
Frame = +3
Query: 454 ELAALRLKDFPSANHKGIEFDQRDRVFRLLDKNGDGKISIEELTEVIEELGAPGEDAHDM 513
+++ LK+ +A D+ R+ LD+NGDG I ++E E G G + +
Sbjct: 258 KISCAELKEMMAALGSKTTSDEVKRMMAELDRNGDGYIDLKEFGEFHCGGGGDGRELREA 437
Query: 514 MLLLDSNSDGSLSSDEFQMFQKQVEMVRNLED 545
L D + +G +S+ E +++ +L D
Sbjct: 438 FELYDLDKNGLISAKELHSVMRRLGEKCSLSD 533
>TC229519 weakly similar to UP|Q93YA8 (Q93YA8) Calcium binding protein,
partial (67%)
Length = 1060
Score = 47.8 bits (112), Expect = 1e-05
Identities = 34/93 (36%), Positives = 48/93 (51%), Gaps = 3/93 (3%)
Frame = +3
Query: 478 RVFRLLDKNGDGKISIEELTEVIEELGA--PGEDAHDMMLLLDSNSDGSLSSDEFQMFQK 535
++F DKNGDGKIS+ EL +++ LG+ E+ MM LD N DG + EF F
Sbjct: 111 QIFNKFDKNGDGKISVTELKDMLAALGSKTTDEELKRMMEELDQNGDGFIDLKEFADFH- 287
Query: 536 QVEMVRNLEDRDDEYKKILDE-KLHMADDSGLI 567
N D+ K++ D L+ D +GLI
Sbjct: 288 -----CNGGAGKDDSKELRDAFDLYDVDKNGLI 371
Score = 33.9 bits (76), Expect = 0.21
Identities = 25/96 (26%), Positives = 47/96 (48%), Gaps = 3/96 (3%)
Frame = +3
Query: 454 ELAALRLKDFPSANHKGIEFDQRDRVFRLLDKNGDGKISIEELTEVIEELGAPGEDA--- 510
+++ LKD +A ++ R+ LD+NGDG I ++E + GA +D+
Sbjct: 147 KISVTELKDMLAALGSKTTDEELKRMMEELDQNGDGFIDLKEFADFHCNGGAGKDDSKEL 326
Query: 511 HDMMLLLDSNSDGSLSSDEFQMFQKQVEMVRNLEDR 546
D L D + +G +S+ E +++RNL ++
Sbjct: 327 RDAFDLYDVDKNGLISAKELH------DVLRNLGEK 416
Score = 32.0 bits (71), Expect = 0.78
Identities = 18/54 (33%), Positives = 32/54 (58%), Gaps = 2/54 (3%)
Frame = +3
Query: 480 FRLLDKNGDGKISIEELTEVIEELGAPGE--DAHDMMLLLDSNSDGSLSSDEFQ 531
F L D + +G IS +EL +V+ LG D M+ +D++ DG+++ +EF+
Sbjct: 336 FDLYDVDKNGLISAKELHDVLRNLGEKCSLSDCRRMISNVDADGDGNVNFEEFK 497
>CA801942
Length = 446
Score = 47.8 bits (112), Expect = 1e-05
Identities = 34/97 (35%), Positives = 50/97 (51%), Gaps = 3/97 (3%)
Frame = +2
Query: 474 DQRDRVFRLLDKNGDGKISIEELTEVIEELGA--PGEDAHDMMLLLDSNSDGSLSSDEFQ 531
D+ ++F DKNGDGKIS+ EL +++ LG+ E+ M+ LD N DG + EF
Sbjct: 53 DEVQQIFNKFDKNGDGKISMAELKDMLSALGSKTTDEELKRMIEELDQNGDGFIDLKEFA 232
Query: 532 MFQKQVEMVRNLEDRDDEYKKILDE-KLHMADDSGLI 567
F N D+ K++ D L+ D +GLI
Sbjct: 233 DFH------CNGGAGKDDSKELRDAFDLYDVDKNGLI 325
Score = 33.9 bits (76), Expect = 0.21
Identities = 26/96 (27%), Positives = 46/96 (47%), Gaps = 3/96 (3%)
Frame = +2
Query: 454 ELAALRLKDFPSANHKGIEFDQRDRVFRLLDKNGDGKISIEELTEVIEELGAPGEDA--- 510
+++ LKD SA ++ R+ LD+NGDG I ++E + GA +D+
Sbjct: 101 KISMAELKDMLSALGSKTTDEELKRMIEELDQNGDGFIDLKEFADFHCNGGAGKDDSKEL 280
Query: 511 HDMMLLLDSNSDGSLSSDEFQMFQKQVEMVRNLEDR 546
D L D + +G +S+ E ++RNL ++
Sbjct: 281 RDAFDLYDVDKNGLISAKELH------HVLRNLGEK 370
>TC226142 similar to UP|Q93YA8 (Q93YA8) Calcium binding protein, partial
(89%)
Length = 572
Score = 47.0 bits (110), Expect = 2e-05
Identities = 34/97 (35%), Positives = 49/97 (50%), Gaps = 8/97 (8%)
Frame = +3
Query: 445 LKTIMLAAAELAALRLKDFPSANHKGI------EFDQRDRVFRLLDKNGDGKISIEELTE 498
LKTI++A + A P+ N + ++ RVF D N DGKIS+ EL
Sbjct: 51 LKTIVMATNPIEAGNGDAAPNPNATTKPSVYLQDTEELKRVFSRFDANCDGKISVTELDN 230
Query: 499 VIEELGA--PGEDAHDMMLLLDSNSDGSLSSDEFQMF 533
V+ LG+ P ED +M LD++ DG ++ EF F
Sbjct: 231 VLRSLGSGVPPEDIQRVMDDLDTDHDGFINLSEFAAF 341
Score = 38.5 bits (88), Expect = 0.008
Identities = 20/54 (37%), Positives = 33/54 (61%), Gaps = 2/54 (3%)
Frame = +3
Query: 480 FRLLDKNGDGKISIEELTEVIEELG--APGEDAHDMMLLLDSNSDGSLSSDEFQ 531
F L D + +G IS EL +V+ LG E+ H+M+ +DS+ DG+++ EF+
Sbjct: 390 FNLYDHDKNGHISATELCQVLNRLGMKCSVEECHNMIKSVDSDGDGNVNFPEFK 551
>TC224523 weakly similar to PIR|C96763|C96763 protein calmodulin F25P22.4
[imported] - Arabidopsis thaliana {Arabidopsis
thaliana;} , partial (33%)
Length = 416
Score = 45.8 bits (107), Expect = 5e-05
Identities = 23/53 (43%), Positives = 32/53 (59%), Gaps = 2/53 (3%)
Frame = +1
Query: 480 FRLLDKNGDGKISIEELTEVIEELG--APGEDAHDMMLLLDSNSDGSLSSDEF 530
FR D+NGDG+IS EE+ E + LG ED M+ +D++ DG + DEF
Sbjct: 49 FRTFDRNGDGRISAEEVKETLGRLGERCSIEDCRRMVRAVDTDGDGMVDMDEF 207
>TC227424 UP|Q39890 (Q39890) Calmodulin, complete
Length = 1424
Score = 45.1 bits (105), Expect = 9e-05
Identities = 30/90 (33%), Positives = 47/90 (51%), Gaps = 2/90 (2%)
Frame = +3
Query: 480 FRLLDKNGDGKISIEELTEVIEEL--GAPGEDAHDMMLLLDSNSDGSLSSDEFQMFQKQV 537
F L DK+GDG I++EEL VI L E+ DM+ +D++ +G++ DEF
Sbjct: 705 FGLFDKDGDGCITVEELATVIRSLDQNPTEEELQDMISEVDADGNGTIEFDEFLSL---- 872
Query: 538 EMVRNLEDRDDEYKKILDEKLHMADDSGLI 567
M + ++D D E + K+ D +G I
Sbjct: 873 -MAKKVKDTDAEEELKEAFKVFDKDQNGYI 959
>TC215516 weakly similar to UP|Q8VWY6 (Q8VWY6) Pollen-specific
calcium-binding protein, partial (51%)
Length = 649
Score = 43.1 bits (100), Expect = 3e-04
Identities = 29/105 (27%), Positives = 53/105 (49%), Gaps = 2/105 (1%)
Frame = +1
Query: 456 AALRLKDFPSANHKGIEFDQRDRVFRLLDKNGDGKISIEELTEVIEELGAP--GEDAHDM 513
+++ L+ F EF+ RV + D++GDGKIS EL + +G +DA +
Sbjct: 67 SSIFLRVFSKVMRMNTEFE---RVLKYFDEDGDGKISPCELRNRLGMIGGELLAKDAEKL 237
Query: 514 MLLLDSNSDGSLSSDEFQMFQKQVEMVRNLEDRDDEYKKILDEKL 558
+ LDS+ DG LS ++F + L+D ++ ++ D ++
Sbjct: 238 IEELDSDGDGFLSLEDFVKLMEAAGEDEKLKDLEEAFEMYNDTEM 372
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.319 0.135 0.379
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,635,546
Number of Sequences: 63676
Number of extensions: 281075
Number of successful extensions: 1775
Number of sequences better than 10.0: 98
Number of HSP's better than 10.0 without gapping: 1734
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1759
length of query: 580
length of database: 12,639,632
effective HSP length: 102
effective length of query: 478
effective length of database: 6,144,680
effective search space: 2937157040
effective search space used: 2937157040
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 62 (28.5 bits)
Medicago: description of AC146862.24


