

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146862.18 - phase: 0 /pseudo
(155 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
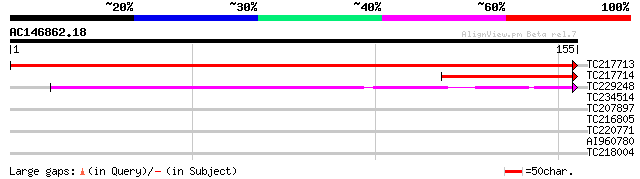
Score E
Sequences producing significant alignments: (bits) Value
TC217713 similar to UP|Q9FI75 (Q9FI75) Similarity to 5'-nucleoti... 287 1e-78
TC217714 homologue to UP|Q9FI75 (Q9FI75) Similarity to 5'-nucleo... 80 3e-16
TC229248 66 6e-12
TC234514 30 0.61
TC207897 weakly similar to UP|Q9LRZ3 (Q9LRZ3) Arabidopsis thalia... 27 3.9
TC216805 similar to GB|AAR11051.1|38098800|AY436891 ANA immunogl... 26 6.7
TC220771 similar to UP|Q9ZS50 (Q9ZS50) Purple acid phosphatase ... 26 8.8
AI960780 26 8.8
TC218004 similar to UP|Q9LGB7 (Q9LGB7) ESTs C73688(E20155), part... 26 8.8
>TC217713 similar to UP|Q9FI75 (Q9FI75) Similarity to 5'-nucleotidase,
partial (36%)
Length = 1174
Score = 287 bits (735), Expect = 1e-78
Identities = 144/155 (92%), Positives = 149/155 (95%)
Frame = +1
Query: 1 MVENSLGIHGDEILYVGDHIYTDVSQSKVHLRWRTALICRELEDEYSALIRCRGDRESLV 60
MVENSL IHGDEILYVGDHIYTDVSQSKVHLRWRTALICRELE+EY ALI R RESLV
Sbjct: 160 MVENSLDIHGDEILYVGDHIYTDVSQSKVHLRWRTALICRELEEEYDALIGSRSYRESLV 339
Query: 61 ELINQKEVVGDLFNQLRLALQRRSKDRPAQTLAATNMDDEDLTESMQKLLIVMQRLDEKI 120
ELINQKE+VGDLFNQLRLALQRRSKDRPAQTLAATN++DEDLT+SMQKLLIVMQRLDEKI
Sbjct: 340 ELINQKEIVGDLFNQLRLALQRRSKDRPAQTLAATNLNDEDLTDSMQKLLIVMQRLDEKI 519
Query: 121 APMLEADGELFNSRWGFLSRAGLWDKSHLMRQIEK 155
APMLEADGELFNSRWGFLSRAGLWDKSHLMRQIEK
Sbjct: 520 APMLEADGELFNSRWGFLSRAGLWDKSHLMRQIEK 624
>TC217714 homologue to UP|Q9FI75 (Q9FI75) Similarity to 5'-nucleotidase,
partial (5%)
Length = 643
Score = 80.5 bits (197), Expect = 3e-16
Identities = 37/37 (100%), Positives = 37/37 (100%)
Frame = +2
Query: 119 KIAPMLEADGELFNSRWGFLSRAGLWDKSHLMRQIEK 155
KIAPMLEADGELFNSRWGFLSRAGLWDKSHLMRQIEK
Sbjct: 2 KIAPMLEADGELFNSRWGFLSRAGLWDKSHLMRQIEK 112
>TC229248
Length = 1630
Score = 66.2 bits (160), Expect = 6e-12
Identities = 42/144 (29%), Positives = 75/144 (51%)
Frame = +1
Query: 12 EILYVGDHIYTDVSQSKVHLRWRTALICRELEDEYSALIRCRGDRESLVELINQKEVVGD 71
++LYVGDHIY D+ +SK L WRT L+ ELE E L R R+ L L ++++ + D
Sbjct: 754 QVLYVGDHIYGDILRSKKVLGWRTMLVIPELEKEVKLLWESRDTRKELQFLRSERDRIED 933
Query: 72 LFNQLRLALQRRSKDRPAQTLAATNMDDEDLTESMQKLLIVMQRLDEKIAPMLEADGELF 131
+ L+ +L+ ++ D ++ ++ + E L +K+ + Q +K+ + F
Sbjct: 934 EIHHLKWSLKFKNPDADSKQKLSSEL--EKLELEREKVRLSHQEAQKKL-------HQRF 1086
Query: 132 NSRWGFLSRAGLWDKSHLMRQIEK 155
+ WG L + G + S Q+E+
Sbjct: 1087HEPWGQLMKTG-YQNSRFAHQVER 1155
>TC234514
Length = 627
Score = 29.6 bits (65), Expect = 0.61
Identities = 14/26 (53%), Positives = 19/26 (72%), Gaps = 1/26 (3%)
Frame = +3
Query: 5 SLGIHGDEILYVGDHIYTD-VSQSKV 29
+L G ++LYVGDHIY D +S+ KV
Sbjct: 360 NLVAEGSQVLYVGDHIYGDTLSRKKV 437
>TC207897 weakly similar to UP|Q9LRZ3 (Q9LRZ3) Arabidopsis thaliana genomic
DNA, chromosome 3, TAC clone:K20I9 (AT3g16810/K20I9_3),
partial (17%)
Length = 929
Score = 26.9 bits (58), Expect = 3.9
Identities = 21/69 (30%), Positives = 32/69 (45%), Gaps = 8/69 (11%)
Frame = +2
Query: 76 LRLALQRRSKDRPAQTLAATNMDDEDLTES--------MQKLLIVMQRLDEKIAPMLEAD 127
+ +A+ +KD+ T+ DD DL ES Q+LLI D + +E+
Sbjct: 68 IEVAVNEVNKDK-------TSADDSDLAESGKKDPFVRRQELLIKSGLADSLLDICIESV 226
Query: 128 GELFNSRWG 136
GEL S +G
Sbjct: 227 GELIQSNFG 253
>TC216805 similar to GB|AAR11051.1|38098800|AY436891 ANA immunoglobulin kappa
light chain {Mus musculus;} , partial (16%)
Length = 1545
Score = 26.2 bits (56), Expect = 6.7
Identities = 20/85 (23%), Positives = 38/85 (44%)
Frame = +1
Query: 40 RELEDEYSALIRCRGDRESLVELINQKEVVGDLFNQLRLALQRRSKDRPAQTLAATNMDD 99
+ LEDEY+ + R RE +++ ++Q+ + ++ L L R + P A D
Sbjct: 220 QRLEDEYAKISAEREARERVIDSLHQEATMWT--DRFALTLNRSQELPPLLAKAKAVADT 393
Query: 100 EDLTESMQKLLIVMQRLDEKIAPML 124
E + LL Q + + + M+
Sbjct: 394 YSAPEEIHGLLSYCQHMIDLMDHMI 468
>TC220771 similar to UP|Q9ZS50 (Q9ZS50) Purple acid phosphatase , partial
(46%)
Length = 783
Score = 25.8 bits (55), Expect = 8.8
Identities = 11/26 (42%), Positives = 14/26 (53%)
Frame = +1
Query: 10 GDEILYVGDHIYTDVSQSKVHLRWRT 35
G +LYVGD Y D ++RW T
Sbjct: 568 GQAVLYVGDLSYADNHPDHDNVRWDT 645
>AI960780
Length = 202
Score = 25.8 bits (55), Expect = 8.8
Identities = 12/30 (40%), Positives = 16/30 (53%)
Frame = +2
Query: 30 HLRWRTALICRELEDEYSALIRCRGDRESL 59
H RW + I RE E+E ++ DRE L
Sbjct: 98 HFRWASVFIEREREEEEMGVLSNMIDREQL 187
>TC218004 similar to UP|Q9LGB7 (Q9LGB7) ESTs C73688(E20155), partial (12%)
Length = 866
Score = 25.8 bits (55), Expect = 8.8
Identities = 18/52 (34%), Positives = 29/52 (55%), Gaps = 8/52 (15%)
Frame = +3
Query: 98 DDEDLTE---SMQKLLIVMQRLDEKIA-----PMLEADGELFNSRWGFLSRA 141
D E++TE +MQ L V++ +E+I P L ++ E+FN G+L A
Sbjct: 171 DAENVTERDPTMQGLDSVIKSFEEEILSPLPNPNLASEAEVFNPNLGYLLEA 326
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.320 0.136 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,849,668
Number of Sequences: 63676
Number of extensions: 61539
Number of successful extensions: 287
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 287
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 287
length of query: 155
length of database: 12,639,632
effective HSP length: 89
effective length of query: 66
effective length of database: 6,972,468
effective search space: 460182888
effective search space used: 460182888
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 55 (25.8 bits)
Medicago: description of AC146862.18


