

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146853.5 - phase: 0 /pseudo
(235 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
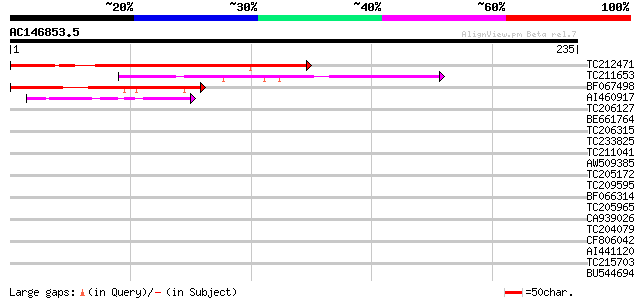
Score E
Sequences producing significant alignments: (bits) Value
TC212471 weakly similar to GB|AAA99226.1|896283|HUMMUT13 methylm... 112 1e-25
TC211653 similar to UP|Q38848 (Q38848) LRP1, partial (27%) 106 8e-24
BF067498 95 2e-20
AI460917 similar to GP|29150089|emb related to c-myb like protei... 42 2e-04
TC206127 weakly similar to PIR|A96697|A96697 protein F1N21.18 [i... 36 0.017
BE661764 33 0.11
TC206315 33 0.11
TC233825 similar to UP|Q9SVY1 (Q9SVY1) Zinc finger-like protein ... 33 0.15
TC211041 33 0.15
AW509385 weakly similar to PIR|A96697|A966 protein F1N21.18 [imp... 32 0.19
TC205172 similar to UP|DX_DROME (Q23985) Deltex protein, partial... 31 0.43
TC209595 homologue to GB|AAL62011.1|18252261|AY072620 AT5g08330/... 31 0.43
BF066314 weakly similar to GP|9858778|gb|A BAC19.10 {Lycopersico... 31 0.56
TC205965 31 0.56
CA939026 31 0.56
TC204079 similar to UP|Q9SWA8 (Q9SWA8) Glycine-rich RNA-binding ... 31 0.56
CF806042 31 0.56
AI441120 31 0.56
TC215703 homologue to PIR|G96755|G96755 developmental protein ho... 30 0.73
BU544694 weakly similar to GP|13676773|gb PF6 protein {Chlamydom... 30 1.2
>TC212471 weakly similar to GB|AAA99226.1|896283|HUMMUT13 methylmalonyl-CoA
mutase {Homo sapiens;} , partial (3%)
Length = 407
Score = 112 bits (281), Expect = 1e-25
Identities = 65/126 (51%), Positives = 77/126 (60%), Gaps = 1/126 (0%)
Frame = +2
Query: 1 MSMLGLRDLVLIAPSPSSLQHHHQQNQNQNQNQNQPISHDHNSNHSLPSSASLSVGFGIF 60
M+MLGLRDLV IAP+PS L HHHQQ+Q PIS +H+SN LPS ASLSVG GIF
Sbjct: 50 MNMLGLRDLVFIAPTPSQL-HHHQQHQ--------PISAEHHSNLPLPSQASLSVGLGIF 202
Query: 61 PLLTATPCMPQQQSQNNEVQENPSNNNNFWNLRMCPEV-VNLPKKGVITEEENHGKIAMM 119
PLLT Q + N + N N+WNL+MC VN +KGV+ E+ MM
Sbjct: 203 PLLTVPHTNDVQVQVQDCANNNNNTNTNYWNLKMCGATEVNSTRKGVMNMEDEGSNKQMM 382
Query: 120 ESEENG 125
ESEE G
Sbjct: 383 ESEEGG 400
>TC211653 similar to UP|Q38848 (Q38848) LRP1, partial (27%)
Length = 515
Score = 106 bits (265), Expect = 8e-24
Identities = 58/150 (38%), Positives = 79/150 (52%), Gaps = 15/150 (10%)
Frame = +3
Query: 46 SLPSSASLSVGFGIFPLLTATPCMPQQQSQNNEVQENPSNNN--NFWNLRMCPEVVNLPK 103
SL + +L VG G+ PLL ATPC+ +S NN + W + + K
Sbjct: 33 SLNPATALGVGVGVIPLLAATPCL---ESDNNILGSRTRGGGGIQLWQDQQQHHQSHYMK 203
Query: 104 K----GVITEE---------ENHGKIAMMESEENGVYGSEYRVCQDCGNRAKKDCVFRRC 150
K G++ +N G++ + G CQDCGN+AKKDC RRC
Sbjct: 204 KQLQQGLLDHNSNTSSGNLIQNSGEVTASGTSSGGT------TCQDCGNQAKKDCANRRC 365
Query: 151 RTCCKGRGYDCSTHLKSTWIPSTRRREREV 180
RTCCK RG+DC TH+KSTW+P+ RRRER++
Sbjct: 366 RTCCKSRGFDCPTHVKSTWVPAARRRERQL 455
>BF067498
Length = 308
Score = 95.1 bits (235), Expect = 2e-20
Identities = 56/89 (62%), Positives = 61/89 (67%), Gaps = 8/89 (8%)
Frame = +3
Query: 1 MSMLGLRDLVLIAPSPSSLQHHHQQNQNQNQNQNQPISHDHNSNHS-----LPSSA-SLS 54
MSMLGLRDLVLIAP+PSSL HH Q QP S +H + H+ LPSSA SLS
Sbjct: 69 MSMLGLRDLVLIAPTPSSLNHH----------QGQPFSENHTNTHTPNNIPLPSSAASLS 218
Query: 55 VGFGIFPLLTATPCMPQ--QQSQNNEVQE 81
VGFGIFPLLTATPC+PQ NNEVQE
Sbjct: 219 VGFGIFPLLTATPCVPQSHHHHHNNEVQE 305
>AI460917 similar to GP|29150089|emb related to c-myb like protein
{Neurospora crassa}, partial (4%)
Length = 421
Score = 42.4 bits (98), Expect = 2e-04
Identities = 26/70 (37%), Positives = 39/70 (55%)
Frame = +1
Query: 8 DLVLIAPSPSSLQHHHQQNQNQNQNQNQPISHDHNSNHSLPSSASLSVGFGIFPLLTATP 67
D+ ++AP+ SS HHH + N N + +NSN+ P++A G +FPLLTATP
Sbjct: 238 DIFVVAPA-SSFYHHHNDVVLSDPNNN---NGSNNSNN--PATAH---GVNVFPLLTATP 390
Query: 68 CMPQQQSQNN 77
C+ + N
Sbjct: 391 CLESEGIMGN 420
>TC206127 weakly similar to PIR|A96697|A96697 protein F1N21.18 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(34%)
Length = 960
Score = 35.8 bits (81), Expect = 0.017
Identities = 25/86 (29%), Positives = 36/86 (41%), Gaps = 9/86 (10%)
Frame = +1
Query: 120 ESEENGVYGSEYRVCQDCGNRAKKDCV----FRRCR-----TCCKGRGYDCSTHLKSTWI 170
E EE RVC+DCGN + CV R+ + CC RG C+ H + +
Sbjct: 109 EQEE*ARRTEASRVCEDCGNESLSVCVESLRLRKAKLGTFEICCWNRGRHCN-HRSGSRL 285
Query: 171 PSTRRREREVEMFAGGGDGEGCSGVK 196
P +R + F G CS ++
Sbjct: 286 PQIQRCSQPHPPFCRQQGG*SCSQIR 363
>BE661764
Length = 634
Score = 33.1 bits (74), Expect = 0.11
Identities = 26/90 (28%), Positives = 40/90 (43%), Gaps = 16/90 (17%)
Frame = +3
Query: 20 QHHHQQNQNQNQNQ-NQPISHDHNSNHSL-------PSSASLSVGFGIFPLLTATPCM-- 69
+HHH + + +NQ +P+ H N + P + + ++ PLLT P
Sbjct: 9 EHHHHIEKKRKENQWRKPLHHLPNPHQWTLKFKTPHPPTPTPTLQSLPLPLLTTWPLPLS 188
Query: 70 -PQQQSQNNE-----VQENPSNNNNFWNLR 93
P ++ NN V N +NNNN W LR
Sbjct: 189HPSPKTNNNNFSNS*VPLNNNNNNNLWFLR 278
>TC206315
Length = 583
Score = 33.1 bits (74), Expect = 0.11
Identities = 21/63 (33%), Positives = 32/63 (50%), Gaps = 9/63 (14%)
Frame = +1
Query: 18 SLQHHHQQNQNQNQNQNQPI---SHDHNSNHSLPSSASLSVGFGIFPLLTAT------PC 68
+L+HHH +Q+QN +QN P+ S +S+ S S+ + +PL T PC
Sbjct: 199 NLKHHHHHHQSQNLSQNHPLFFTSSHFSSSSSFSSTYPTNHTLLPWPLQKKTPNHQNQPC 378
Query: 69 MPQ 71
PQ
Sbjct: 379 PPQ 387
>TC233825 similar to UP|Q9SVY1 (Q9SVY1) Zinc finger-like protein (WIP2
protein), partial (7%)
Length = 527
Score = 32.7 bits (73), Expect = 0.15
Identities = 23/75 (30%), Positives = 37/75 (48%), Gaps = 7/75 (9%)
Frame = +2
Query: 14 PSPSSLQHHHQQ-----NQN-QNQNQNQPISHDHNSNHSLPSSASLSVGF-GIFPLLTAT 66
PSP+ HHH NQN N N + H H+ ++ SS+ S PLLT +
Sbjct: 170 PSPNPSPHHHPYAHNFLNQNGSNINNHNTFFHFHHFQTTICSSSPPSPPLREALPLLTLS 349
Query: 67 PCMPQQQSQNNEVQE 81
P ++Q ++++ Q+
Sbjct: 350 PRNEEEQEEDDQDQD 394
>TC211041
Length = 1090
Score = 32.7 bits (73), Expect = 0.15
Identities = 13/30 (43%), Positives = 21/30 (69%)
Frame = -2
Query: 37 ISHDHNSNHSLPSSASLSVGFGIFPLLTAT 66
++HDH+ +HS S S+++ IFPLL+ T
Sbjct: 693 VAHDHDHDHSSSSEMSITIASLIFPLLSPT 604
>AW509385 weakly similar to PIR|A96697|A966 protein F1N21.18 [imported] -
Arabidopsis thaliana, partial (22%)
Length = 449
Score = 32.3 bits (72), Expect = 0.19
Identities = 18/53 (33%), Positives = 26/53 (48%), Gaps = 9/53 (16%)
Frame = +2
Query: 132 RVCQDCGNRAKKDCV----FRRCR-----TCCKGRGYDCSTHLKSTWIPSTRR 175
RVC+DCGN + CV R+ + CC RG C+ H + +P +R
Sbjct: 59 RVCEDCGNESLSVCVESLRLRKAKLGTFEICCWNRGRHCN-HRSGSRLPQIQR 214
>TC205172 similar to UP|DX_DROME (Q23985) Deltex protein, partial (4%)
Length = 579
Score = 31.2 bits (69), Expect = 0.43
Identities = 26/74 (35%), Positives = 36/74 (48%), Gaps = 5/74 (6%)
Frame = +3
Query: 20 QHHHQQNQNQNQNQNQPISHDHNSNH-----SLPSSASLSVGFGIFPLLTATPCMPQQQS 74
QHHHQQ Q Q Q+Q ++H +++H + SS S + PLL P P
Sbjct: 39 QHHHQQQQQHQQQQHQ-LAHFDSTSHDDFLEQMLSSCSWTDLNHNKPLLW-DPNTPNDIK 212
Query: 75 QNNEVQENPSNNNN 88
+E PSNNN+
Sbjct: 213 PPDET--TPSNNND 248
>TC209595 homologue to GB|AAL62011.1|18252261|AY072620 AT5g08330/F8L15_60
{Arabidopsis thaliana;} , partial (56%)
Length = 869
Score = 31.2 bits (69), Expect = 0.43
Identities = 19/55 (34%), Positives = 25/55 (44%), Gaps = 4/55 (7%)
Frame = +1
Query: 8 DLVLIAPSPSSLQHHHQQNQNQNQNQNQPISHDHNSNHSLPSS----ASLSVGFG 58
+LV ++P S HHHQ +Q Q Q + + LP ASLS G G
Sbjct: 487 ELVSVSPQNSMFHHHHQHHQQQQQQAAMGEASAARLGNYLPGHLNLLASLSGGHG 651
>BF066314 weakly similar to GP|9858778|gb|A BAC19.10 {Lycopersicon
esculentum}, partial (4%)
Length = 308
Score = 30.8 bits (68), Expect = 0.56
Identities = 16/31 (51%), Positives = 18/31 (57%)
Frame = +1
Query: 15 SPSSLQHHHQQNQNQNQNQNQPISHDHNSNH 45
SPSS Q+H N NQNQN P + N NH
Sbjct: 220 SPSSRQNHQSSNLNQNQNLPPP---NLNPNH 303
>TC205965
Length = 1409
Score = 30.8 bits (68), Expect = 0.56
Identities = 24/74 (32%), Positives = 32/74 (42%)
Frame = -1
Query: 13 APSPSSLQHHHQQNQNQNQNQNQPISHDHNSNHSLPSSASLSVGFGIFPLLTATPCMPQQ 72
+PS LQHH + QN Q P+ L +S+S PL + PC+PQ
Sbjct: 959 SPSSQVLQHHPNCFEIQNIAQGLPLQ-------ILEASSS--------PLFSTKPCLPQ- 828
Query: 73 QSQNNEVQENPSNN 86
EV+ N NN
Sbjct: 827 -----EVRRNAENN 801
>CA939026
Length = 426
Score = 30.8 bits (68), Expect = 0.56
Identities = 25/95 (26%), Positives = 39/95 (40%), Gaps = 21/95 (22%)
Frame = +1
Query: 20 QHHHQQNQNQNQNQ-NQPISHDHNSNHSL-------PSSASLSVGFGIFPLLTATPCM-- 69
+HHH + + +NQ +P+ H N + P + + ++ PLLT P
Sbjct: 10 EHHHHIEKKRKENQWRKPLHHLPNPHQWTLKFKTPHPPTPTPTLQSLPLPLLTTWPLPLS 189
Query: 70 -PQQQSQNNEVQE----------NPSNNNNFWNLR 93
P ++ NN N +NNNN W LR
Sbjct: 190 HPSPKTNNNNFSNS*VPLNNNNNNNNNNNNLWFLR 294
>TC204079 similar to UP|Q9SWA8 (Q9SWA8) Glycine-rich RNA-binding protein,
complete
Length = 748
Score = 30.8 bits (68), Expect = 0.56
Identities = 13/36 (36%), Positives = 20/36 (55%), Gaps = 3/36 (8%)
Frame = -1
Query: 13 APSPSSL---QHHHQQNQNQNQNQNQPISHDHNSNH 45
+PS SSL +HHH+ N + + N + H H+ H
Sbjct: 586 SPSFSSLHHHRHHHESNHGHHHHHNHGLCHRHSHRH 479
>CF806042
Length = 604
Score = 30.8 bits (68), Expect = 0.56
Identities = 23/72 (31%), Positives = 36/72 (49%), Gaps = 3/72 (4%)
Frame = +1
Query: 9 LVLIAPSPS---SLQHHHQQNQNQNQNQNQPISHDHNSNHSLPSSASLSVGFGIFPLLTA 65
+ ++AP+ S +L HHH + + N P + N P++A +G G+ PLL
Sbjct: 355 MFVVAPASSFHPNLHHHH----HHQPSPNDPAVMVDSLN---PATA---LGVGVIPLLAP 504
Query: 66 TPCMPQQQSQNN 77
TPC +S NN
Sbjct: 505 TPC---HESDNN 531
>AI441120
Length = 378
Score = 30.8 bits (68), Expect = 0.56
Identities = 16/47 (34%), Positives = 22/47 (46%)
Frame = +3
Query: 25 QNQNQNQNQNQPISHDHNSNHSLPSSASLSVGFGIFPLLTATPCMPQ 71
Q Q Q Q QPI H HN+ + + +S S F P + C P+
Sbjct: 180 QRLQQQQQQQQPIGHKHNTMSKINNHSSSSCSFFTSPCI----CRPK 308
>TC215703 homologue to PIR|G96755|G96755 developmental protein homolog DG1118
[imported] - Arabidopsis thaliana {Arabidopsis
thaliana;} , complete
Length = 896
Score = 30.4 bits (67), Expect = 0.73
Identities = 12/26 (46%), Positives = 18/26 (69%)
Frame = -3
Query: 55 VGFGIFPLLTATPCMPQQQSQNNEVQ 80
+ F +FP+LTA C+ SQN+E+Q
Sbjct: 111 ISFSVFPILTALLCLGFFYSQNSEIQ 34
>BU544694 weakly similar to GP|13676773|gb PF6 protein {Chlamydomonas
reinhardtii}, partial (0%)
Length = 500
Score = 29.6 bits (65), Expect = 1.2
Identities = 13/32 (40%), Positives = 16/32 (49%)
Frame = +3
Query: 14 PSPSSLQHHHQQNQNQNQNQNQPISHDHNSNH 45
P+ H HQ N++Q P SH H SNH
Sbjct: 246 PTXXPYHHRHQPNRHQPPTTRNP-SHHHQSNH 338
Score = 27.3 bits (59), Expect = 6.2
Identities = 10/23 (43%), Positives = 14/23 (60%)
Frame = +1
Query: 13 APSPSSLQHHHQQNQNQNQNQNQ 35
AP P HH +Q ++Q+QNQ
Sbjct: 166 APEPKHQHHHQHHHQPKHQDQNQ 234
Score = 27.3 bits (59), Expect = 6.2
Identities = 16/35 (45%), Positives = 22/35 (62%), Gaps = 7/35 (20%)
Frame = +1
Query: 12 IAPSPSSL-QHHHQ---QNQNQNQNQN---QPISH 39
+AP+P QHHHQ Q ++Q+QNQ+ QP H
Sbjct: 157 LAPAPEPKHQHHHQHHHQPKHQDQNQHLVCQPXXH 261
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.313 0.128 0.389
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,571,953
Number of Sequences: 63676
Number of extensions: 232887
Number of successful extensions: 2219
Number of sequences better than 10.0: 127
Number of HSP's better than 10.0 without gapping: 2052
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2172
length of query: 235
length of database: 12,639,632
effective HSP length: 94
effective length of query: 141
effective length of database: 6,654,088
effective search space: 938226408
effective search space used: 938226408
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 57 (26.6 bits)
Medicago: description of AC146853.5


