

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146819.13 + phase: 0
(97 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
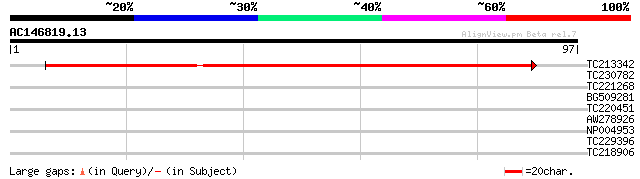
Score E
Sequences producing significant alignments: (bits) Value
TC213342 weakly similar to UP|O82445 (O82445) Response regulator... 69 5e-13
TC230782 similar to UP|Q9C5U1 (Q9C5U1) Histidine kinase, partial... 37 0.002
TC221268 similar to UP|Q9SXL4 (Q9SXL4) Histidine kinase 1, parti... 36 0.003
BG509281 similar to GP|14023997|dbj Hybrid sensory histidine kin... 35 0.005
TC220451 similar to UP|Q9SXL4 (Q9SXL4) Histidine kinase 1, parti... 35 0.008
AW278926 similar to GP|28852408|gb response regulator {Pseudomon... 33 0.023
NP004953 possible GTP binding protein 29 0.43
TC229396 similar to UP|Q9LVG4 (Q9LVG4) Emb|CAB86035.1 (Pseudo-re... 27 1.3
TC218906 25 8.2
>TC213342 weakly similar to UP|O82445 (O82445) Response regulator protein,
partial (41%)
Length = 599
Score = 68.6 bits (166), Expect = 5e-13
Identities = 37/84 (44%), Positives = 53/84 (63%)
Frame = +2
Query: 7 LIDIGVETHAVKNGKESLDLISTVERKFDLILMARIMPVMDGITATKKLRDLEYPGQIVG 66
L +G++ V+NG+E++D+ +R FDLILM MP+M+GI ATK+LR + IVG
Sbjct: 86 LESVGMKNQGVENGQEAVDIHCHGQR-FDLILMDMDMPIMNGIEATKELRSMGIGSMIVG 262
Query: 67 VSRHLTNDEYAEFCNAGLDDLHHE 90
VS T E +F AGL+D H +
Sbjct: 263 VSSRCTEAEIRKFMEAGLNDYHEK 334
>TC230782 similar to UP|Q9C5U1 (Q9C5U1) Histidine kinase, partial (14%)
Length = 869
Score = 37.0 bits (84), Expect = 0.002
Identities = 21/71 (29%), Positives = 37/71 (51%)
Frame = +1
Query: 11 GVETHAVKNGKESLDLISTVERKFDLILMARIMPVMDGITATKKLRDLEYPGQIVGVSRH 70
G + V +GK+++ + +FD M MP MDG ATK++R++E V+R
Sbjct: 535 GADVVCVSSGKDAISSLKP-PHQFDACFMDIQMPEMDGFEATKRIREME-----DSVNRE 696
Query: 71 LTNDEYAEFCN 81
++ D++ N
Sbjct: 697 VSMDDFENITN 729
>TC221268 similar to UP|Q9SXL4 (Q9SXL4) Histidine kinase 1, partial (10%)
Length = 978
Score = 36.2 bits (82), Expect = 0.003
Identities = 21/40 (52%), Positives = 24/40 (59%)
Frame = +1
Query: 32 RKFDLILMARIMPVMDGITATKKLRDLEYPGQIVGVSRHL 71
R +DLILM MP MDG ATK +R E VG SRH+
Sbjct: 514 RPYDLILMDCQMPKMDGYEATKAIRKSE-----VGTSRHI 618
>BG509281 similar to GP|14023997|dbj Hybrid sensory histidine kinase
{Mesorhizobium loti}, partial (6%)
Length = 420
Score = 35.4 bits (80), Expect = 0.005
Identities = 18/56 (32%), Positives = 32/56 (57%), Gaps = 6/56 (10%)
Frame = +1
Query: 10 IGVETHAVKNGKESLDLI------STVERKFDLILMARIMPVMDGITATKKLRDLE 59
+G +NG++++ + ++ R D ILM MPVMDG AT+++R++E
Sbjct: 4 LGASVMECENGEQAVQTVEEGLTRNSSNRPCDFILMDCQMPVMDGYEATRRIREIE 171
>TC220451 similar to UP|Q9SXL4 (Q9SXL4) Histidine kinase 1, partial (9%)
Length = 501
Score = 34.7 bits (78), Expect = 0.008
Identities = 23/58 (39%), Positives = 31/58 (52%), Gaps = 6/58 (10%)
Frame = +2
Query: 34 FDLILMARIMPVMDGITATKKLR------DLEYPGQIVGVSRHLTNDEYAEFCNAGLD 85
+DLILM MP MDG ATK +R DL P IV ++ H + + A+ G+D
Sbjct: 197 YDLILMDCQMPKMDGYEATKAIRKSEVGTDLHIP--IVALTAHAMSCDEAKCLEVGMD 364
>AW278926 similar to GP|28852408|gb response regulator {Pseudomonas syringae
pv. tomato str. DC3000}, partial (21%)
Length = 397
Score = 33.1 bits (74), Expect = 0.023
Identities = 25/81 (30%), Positives = 45/81 (54%), Gaps = 2/81 (2%)
Frame = +3
Query: 19 NGKESLDLISTVERKFDLILMARIMPVMDGITATKKLRDL--EYPGQIVGVSRHLTNDEY 76
NG +++++ T ER L+LM +MPVMDG A ++++ L E I+ ++ ++
Sbjct: 24 NGAQAVEVF-TRERP-QLVLMDALMPVMDGFEAARRIKQLAGEALVPIIFLTSLRESEAL 197
Query: 77 AEFCNAGLDDLHHEDGIPLAI 97
A+ +AG DD + PL +
Sbjct: 198 AQCLDAGGDDFLPKPYNPLIL 260
>NP004953 possible GTP binding protein
Length = 903
Score = 28.9 bits (63), Expect = 0.43
Identities = 15/39 (38%), Positives = 23/39 (58%)
Frame = -1
Query: 30 VERKFDLILMARIMPVMDGITATKKLRDLEYPGQIVGVS 68
V + DLI+ MPV+DG T++ R +EY I G++
Sbjct: 156 VRHRIDLIVN---MPVVDGYELTERSRKMEYRKPIFGIT 49
>TC229396 similar to UP|Q9LVG4 (Q9LVG4) Emb|CAB86035.1 (Pseudo-response
regulator 3), partial (28%)
Length = 1013
Score = 27.3 bits (59), Expect = 1.3
Identities = 12/34 (35%), Positives = 19/34 (55%)
Frame = +1
Query: 16 AVKNGKESLDLISTVERKFDLILMARIMPVMDGI 49
AV NG ++ ++ E DL+L MP++ GI
Sbjct: 67 AVSNGLQAWKVLEDPENGIDLVLTEVAMPILSGI 168
>TC218906
Length = 1258
Score = 24.6 bits (52), Expect = 8.2
Identities = 26/94 (27%), Positives = 43/94 (45%), Gaps = 13/94 (13%)
Frame = -1
Query: 3 LFEWLIDIGVETHAVKNGKES-LDLISTVERKFDLILMARIMPVMDG------ITATKKL 55
L + L+ I + A K ES I + ++F+L+L +I P+ + + L
Sbjct: 814 LLQLLVSILIIRKAFKLDSESTFKTIFLLSQRFNLLLQHQIFPLKSRCCPGTCVCSD*FL 635
Query: 56 RDLEY-----PGQI-VGVSRHLTNDEYAEFCNAG 83
+LE PG I +RHL+N + FC+ G
Sbjct: 634 *NLEQSSSTGPGIINTSSTRHLSNKQVNPFCSIG 533
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.322 0.142 0.417
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,647,727
Number of Sequences: 63676
Number of extensions: 30243
Number of successful extensions: 111
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 111
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 111
length of query: 97
length of database: 12,639,632
effective HSP length: 73
effective length of query: 24
effective length of database: 7,991,284
effective search space: 191790816
effective search space used: 191790816
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 51 (24.3 bits)
Medicago: description of AC146819.13


