

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
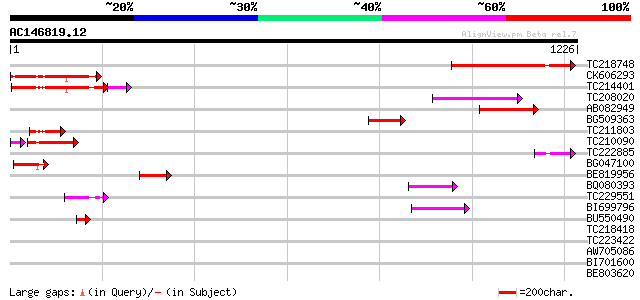
Query= AC146819.12 + phase: 0 /pseudo
(1226 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC218748 similar to UP|Q76CU1 (Q76CU1) PDR-type ABC transporter ... 262 8e-70
CK606293 218 1e-56
TC214401 weakly similar to UP|Q949G3 (Q949G3) ABC1 protein, part... 209 7e-56
TC208020 similar to UP|Q76CU2 (Q76CU2) PDR-type ABC transporter ... 130 5e-30
AB082949 125 1e-28
BG509363 similar to GP|14331118|e ABC1 protein {Nicotiana plumba... 93 8e-19
TC211803 similar to UP|Q76CU2 (Q76CU2) PDR-type ABC transporter ... 77 5e-14
TC210090 similar to UP|Q9LFH0 (Q9LFH0) PDR9 ABC transporter, par... 70 8e-14
TC222885 weakly similar to UP|Q8GU89 (Q8GU89) PDR-like ABC trans... 66 1e-10
BG047100 similar to GP|20522008|db pleiotropic drug resistance l... 63 7e-10
BE819956 similar to GP|20522008|db pleiotropic drug resistance l... 63 9e-10
BQ080393 49 1e-05
TC229551 weakly similar to UP|Q6X4V5 (Q6X4V5) ABC transporter, p... 49 1e-05
BI699796 similar to PIR|G85167|G85 ABC transporter like protein ... 49 2e-05
BU550490 48 2e-05
TC218418 homologue to UP|Q6X4V5 (Q6X4V5) ABC transporter, partia... 43 0.001
TC223422 39 0.011
AW705086 39 0.018
BI701600 38 0.024
BE803620 homologue to GP|13605839|gb| At2g01320/F10A8.20 {Arabid... 38 0.024
>TC218748 similar to UP|Q76CU1 (Q76CU1) PDR-type ABC transporter 2 (Fragment),
partial (37%)
Length = 1384
Score = 262 bits (669), Expect = 8e-70
Identities = 155/268 (57%), Positives = 183/268 (67%)
Frame = +2
Query: 956 GKSFNNVHG*ANFRARCKICRYCYESH*KYSGEWKNCCLCHSSIKY*YI*IF**AVIDEA 1015
GKSF N++G*ANF ARCK C YC+E+ *++S KN CL H S K+ +I*IF**A +EA
Sbjct: 17 GKSFYNIYG*ANFWARCKSCCYCHENS*EHSRHRKNSCLYHPSAKHRHI*IF**AFANEA 196
Query: 1016 RRASDICGANRTSFFPFNKLF*GD*RCQ*D*RWL*PSSLDVGNYFFRKRNAIGD*FF*SV 1075
RR +ICGA TSFFPFN L *G+ RCQ*D*RWL* ++D G+ F KRN IGD*F *SV
Sbjct: 197 RRPRNICGATWTSFFPFN*LL*GNPRCQ*D*RWL*SGNMDAGSLDFSKRNGIGD*FC*SV 376
Query: 1076 QKFRIIQEKQSSYCRIEHSSS*FCESSFSFKILKTPFRSIQGLLMETTLVLLAESAIQCI 1135
QKFR+IQEKQS+Y RIE+SSS F F +L S+ GLLMETTLVLLA+S+I C
Sbjct: 377 QKFRVIQEKQSTY*RIEYSSSWFKRPLFPITVLNLLPHSMHGLLMETTLVLLAQSSIHCY 556
Query: 1136 KISLHSRCLYLFRERVLWPWLQNVHFNQLQ*KETRSP*LHWLNVDYYSTYWH*ECWFCAS 1195
KISL + C VL PWLQN *+ TRS H L+V S YWH*EC A+
Sbjct: 557 KISLLNCCSCCAW*HVLGPWLQN-------*QTTRSFQCHGLHVCCCSPYWH*EC*CSAA 715
Query: 1196 SGDCRESSFL*GKCCQNVFSTGICFWSG 1223
SG C +S L*GK +NVFS +CF SG
Sbjct: 716 SGCC*ANSLL*GKSSRNVFSFTVCFCSG 799
>CK606293
Length = 782
Score = 218 bits (555), Expect = 1e-56
Identities = 118/202 (58%), Positives = 156/202 (76%), Gaps = 6/202 (2%)
Frame = -3
Query: 2 ESDEISLMKWDSIQRLPTVARLRRGLLTTPEGDSNEIDVHKIGLQERTYLLQRLLRNNTV 61
E+DE +L KW +IQ+LPTVARLR+ L+T+P+G+SNEIDV K+GLQE+ LL+RL++ T
Sbjct: 561 ENDEEAL-KWAAIQKLPTVARLRKALITSPDGESNEIDVKKLGLQEKKALLERLVK--TA 391
Query: 62 EVDNDHSFLLKLMRDRIDRAGVDIPTIEVRFEHLNVQAQVHVGKRALHTITNYMLDLVE- 120
+ DN+ FLLKL +DRIDR G+D+PTIEVRFE+L+++A+ G RAL T TN++++++E
Sbjct: 390 QEDNE-KFLLKL-KDRIDRVGIDLPTIEVRFENLSIEAEARAGTRALPTFTNFIVNILEG 217
Query: 121 -----YILKRRKQQLNILQDVSGILKHSRLTLLLGPPNSGKTILLLALAGKLDPNLKIAN 175
++L RKQ LNIL+DVSGI+K R+TLLLGPP+SGKT LLLALAGKLDP K
Sbjct: 216 LLNSLHVLPNRKQHLNILEDVSGIIKPGRMTLLLGPPSSGKTTLLLALAGKLDPKKK--- 46
Query: 176 EVQFYEQFAGKVSYNGHEMNEF 197
F+GKV+YNGH MNEF
Sbjct: 45 -------FSGKVTYNGHGMNEF 1
>TC214401 weakly similar to UP|Q949G3 (Q949G3) ABC1 protein, partial (18%)
Length = 1177
Score = 209 bits (531), Expect(2) = 7e-56
Identities = 111/217 (51%), Positives = 154/217 (70%), Gaps = 6/217 (2%)
Frame = +1
Query: 4 DEISLMKWDSIQRLPTVARLRRGLLTTPEGDSNEIDVHKIGLQERTYLLQRLLRNNTVEV 63
D+ +KW +I+RLPT R+RR +L +G E+D+ ++GL ER +++RL++ E
Sbjct: 373 DDEEALKWAAIERLPTYLRIRRSILNNEDGKGREVDIKQLGLTERKIIVERLVK--IAEE 546
Query: 64 DNDHSFLLKLMRDRIDRAGVDIPTIEVRFEHLNVQAQVHVGKRALHTITNYMLDLVE--- 120
DN+ FLLKL R+R+DR G+DIPTIEVRFEH+NV+AQV+VG RAL ++ N+ +++E
Sbjct: 547 DNER-FLLKL-RERMDRVGLDIPTIEVRFEHINVEAQVYVGGRALPSMLNFFANVIEGFL 720
Query: 121 ---YILKRRKQQLNILQDVSGILKHSRLTLLLGPPNSGKTILLLALAGKLDPNLKIANEV 177
+I+ K+ L ILQ+VSGI+K R+TLLLGPP SGKT LLLALAGKLD +L
Sbjct: 721 NYLHIIPSPKKPLRILQNVSGIIKPRRMTLLLGPPGSGKTTLLLALAGKLDKDL------ 882
Query: 178 QFYEQFAGKVSYNGHEMNEFVPQRTAAYVSQNDTHLG 214
+G+V+YNGH + EFVPQRT+AY+SQ D H+G
Sbjct: 883 ----NHSGRVTYNGHGLEEFVPQRTSAYISQYDNHIG 981
Score = 28.5 bits (62), Expect(2) = 7e-56
Identities = 24/52 (46%), Positives = 26/52 (49%)
Frame = +2
Query: 211 THLGN*LSEKPWPFQQEFKELDLDMIC*KRCVEERWRKTLFLIRISMSI*RL 262
T L *L EK W Q + K LD M C EER + L I I M I*RL
Sbjct: 971 TTLEK*L*EKLWLSQLDAKGLDRTMRCWLSY*EERSMQRLNRILILMPI*RL 1126
>TC208020 similar to UP|Q76CU2 (Q76CU2) PDR-type ABC transporter 1, partial
(14%)
Length = 765
Score = 130 bits (326), Expect = 5e-30
Identities = 85/196 (43%), Positives = 112/196 (56%), Gaps = 1/196 (0%)
Frame = +1
Query: 914 DVHRGSYGACRTDPT-EGYNCCAWCYWSLYTTAKKVDYCGRVSGKSFNNVHG*ANFRARC 972
DVH GS+G +P E A C WSL T K+ DYC +SG +N HG*A F RC
Sbjct: 157 DVH*GSHGTGGAEPIKELTGWIARCEWSLN*TTKETDYCS*ISG*PVHNFHG*AYFWVRC 336
Query: 973 KICRYCYESH*KYSGEWKNCCLCHSSIKY*YI*IF**AVIDEARRASDICGANRTSFFPF 1032
K C CYE+ +Y G WKN CL H S +*+I* *A+ +EA R +IC A + + F
Sbjct: 337 KSCCNCYENSQEYCGHWKNRCLHHPSA*H*HI*SIR*AIPNEAWRTRNICWAIGSPLYSF 516
Query: 1033 NKLF*GD*RCQ*D*RWL*PSSLDVGNYFFRKRNAIGD*FF*SVQKFRIIQEKQSSYCRIE 1092
+++F* R + + R +*P LDVG+Y F R G *F+* VQ+F I EKQ++Y R
Sbjct: 517 DQVF*EHWRGEQNQRRI*PGYLDVGSYNFSTRT*FGC*FY*LVQEF*SI*EKQAAYTRTG 696
Query: 1093 HSSS*FCESSFSFKIL 1108
+ S S F + IL
Sbjct: 697 STCSRLKGSLFPYSIL 744
>AB082949
Length = 403
Score = 125 bits (313), Expect = 1e-28
Identities = 74/128 (57%), Positives = 90/128 (69%)
Frame = +2
Query: 1016 RRASDICGANRTSFFPFNKLF*GD*RCQ*D*RWL*PSSLDVGNYFFRKRNAIGD*FF*SV 1075
RR DICGA TSF FN+L *G+*RC +* WL S+LDVG++ F KRN IGD*F *+V
Sbjct: 8 RRTRDICGATWTSFLSFNQLL*GN*RCPYN*GWLQSSNLDVGSHNFSKRNGIGD*FR*AV 187
Query: 1076 QKFRIIQEKQSSYCRIEHSSS*FCESSFSFKILKTPFRSIQGLLMETTLVLLAESAIQCI 1135
KFR+IQEKQ +Y RIE+SS F F FK+L + + GLLMETTLVLLA+ I
Sbjct: 188 *KFRLIQEKQGTY*RIEYSSPGFKGPLFFFKVL*VLYYPMHGLLMETTLVLLAQ**IHRP 367
Query: 1136 KISLHSRC 1143
KISL++ C
Sbjct: 368 KISLYNCC 391
>BG509363 similar to GP|14331118|e ABC1 protein {Nicotiana plumbaginifolia},
partial (8%)
Length = 455
Score = 92.8 bits (229), Expect = 8e-19
Identities = 44/80 (55%), Positives = 54/80 (67%)
Frame = +3
Query: 777 EERNDSFF*TTLYHL**SNIFS*YATGNEESTSCRGKVEPIEWCQWFIQTGCSHSSDGCD 836
E+RN S F* T YHL**S+I *+ATGNE + RG++ E C W IQ CSHS DGC
Sbjct: 213 EKRNGSSF*ATFYHL**SHILC*HATGNEGTGCTRGQIGAFEGC*WCIQAWCSHSFDGCK 392
Query: 837 WCWQNNSDGCTSWKKNQRIY 856
W W + DGC+ W +N+RIY
Sbjct: 393 WSW*DYFDGCSGW*ENRRIY 452
Score = 34.7 bits (78), Expect = 0.26
Identities = 29/96 (30%), Positives = 49/96 (50%), Gaps = 12/96 (12%)
Frame = +1
Query: 83 VDIPTIEVRFEHLNVQAQVHVGKRAL------HTITN----YMLDLVEYILKRRKQQ--L 130
V++P IE +V H K+ + H+IT Y +D+ + + ++ Q+ L
Sbjct: 148 VELPRIESSGRGDSVVESSHGKKKGMVLPFEPHSITFDEVIYSVDMPQEMKEQGVQEDRL 327
Query: 131 NILQDVSGILKHSRLTLLLGPPNSGKTILLLALAGK 166
+L+ VSG + LT L+G +GKT L+ LAG+
Sbjct: 328 VLLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGR 435
>TC211803 similar to UP|Q76CU2 (Q76CU2) PDR-type ABC transporter 1, partial
(6%)
Length = 255
Score = 77.0 bits (188), Expect = 5e-14
Identities = 42/77 (54%), Positives = 58/77 (74%)
Frame = +1
Query: 44 GLQERTYLLQRLLRNNTVEVDNDHSFLLKLMRDRIDRAGVDIPTIEVRFEHLNVQAQVHV 103
G+QER LL+RL++ E DN+ FLLKL ++RIDR G+DIPTIEVR+EHLN++A+ V
Sbjct: 4 GIQERQKLLERLVK--VAEEDNER-FLLKL-KERIDRVGLDIPTIEVRYEHLNIEAEAFV 171
Query: 104 GKRALHTITNYMLDLVE 120
G RAL + N + ++VE
Sbjct: 172 GSRALPSFINSVTNVVE 222
>TC210090 similar to UP|Q9LFH0 (Q9LFH0) PDR9 ABC transporter, partial (12%)
Length = 1189
Score = 69.7 bits (169), Expect(2) = 8e-14
Identities = 36/117 (30%), Positives = 77/117 (65%), Gaps = 6/117 (5%)
Frame = +3
Query: 38 IDVHKIGLQERTYLLQRLLRNNTVEVDNDHSFLLKLMRDRIDRAGVDIPTIEVRFEHLNV 97
+DV K+G QER +++L+++ ++ND+ LL+ R RID+ G+++PT+E+R+++L+V
Sbjct: 282 VDVRKLGAQERHTFIEKLIKH----IENDNLRLLQKFRKRIDKVGINLPTVELRYQNLSV 449
Query: 98 QAQVHV--GK---RALHTITNYMLDLVEY-ILKRRKQQLNILQDVSGILKHSRLTLL 148
+A+ + GK +T+ ++ D + +LK + +++I+++ +GI+K R +L
Sbjct: 450 EAECKIVQGKPIPTLWNTLKEWIFDTTKLSVLKSQNSKISIIKNDNGIIKPGRYAIL 620
Score = 26.6 bits (57), Expect(2) = 8e-14
Identities = 11/33 (33%), Positives = 18/33 (54%)
Frame = +2
Query: 1 MESDEISLMKWDSIQRLPTVARLRRGLLTTPEG 33
+++D ++W IQRLPT R+ L +G
Sbjct: 170 VDNDAGEALQWAEIQRLPTFERITSALFDVYDG 268
>TC222885 weakly similar to UP|Q8GU89 (Q8GU89) PDR-like ABC transporter (PDR4
ABC transporter), partial (9%)
Length = 484
Score = 65.9 bits (159), Expect = 1e-10
Identities = 40/88 (45%), Positives = 52/88 (58%)
Frame = +1
Query: 1136 KISLHSRCLYLFRERVLWPWLQNVHFNQLQ*KETRSP*LHWLNVDYYSTYWH*ECWFCAS 1195
KISL++ C + E +L PWLQN K TRS H ++V S+ W+*E F +
Sbjct: 85 KISLYNCCCLVVWEHILEPWLQNQ-------KATRSFQCHGIHVCCCSSSWY*EF*FSTT 243
Query: 1196 SGDCRESSFL*GKCCQNVFSTGICFWSG 1223
+G ++S L*GK C NVFS GICF SG
Sbjct: 244 AGGS*KNSVL*GKSCWNVFSFGICFCSG 327
>BG047100 similar to GP|20522008|db pleiotropic drug resistance like
protein {Nicotiana tabacum}, partial (4%)
Length = 229
Score = 63.2 bits (152), Expect = 7e-10
Identities = 34/81 (41%), Positives = 50/81 (60%), Gaps = 5/81 (6%)
Frame = +2
Query: 9 MKWDSIQRLPTVARLRRGLLTTPEGDSNEIDVHKIGLQERTYLLQRLLR-----NNTVEV 63
+KW ++++LPT RLR+GLLT G +NEIDV +G+QER L +RL++ N V V
Sbjct: 14 LKWAALEKLPTYNRLRKGLLTASHGVANEIDVSDLGIQERQKLFERLVKVAV*DNERVSV 193
Query: 64 DNDHSFLLKLMRDRIDRAGVD 84
+ +R+ IDR G+D
Sbjct: 194E---------LRESIDRVGLD 229
>BE819956 similar to GP|20522008|db pleiotropic drug resistance like protein
{Nicotiana tabacum}, partial (6%)
Length = 673
Score = 62.8 bits (151), Expect = 9e-10
Identities = 39/69 (56%), Positives = 44/69 (63%)
Frame = -2
Query: 282 DWTYVKILW*EMRS*KASLKDKGNVLQ*GRR*SDH*NLYSWMIYLLVWMTQQPFKL*NR* 341
DW V IL *EM+ * L DKGNVLQ GR D L SWM YLLVW+ +Q K * R
Sbjct: 444 DWRSVLILL*EMQC*GGYLVDKGNVLQHGRCWLD*LMLNSWMKYLLVWIARQLTKF*TRS 265
Query: 342 NNLSTFSKE 350
+N+ TFSKE
Sbjct: 264 SNVFTFSKE 238
>BQ080393
Length = 347
Score = 49.3 bits (116), Expect = 1e-05
Identities = 37/108 (34%), Positives = 55/108 (50%), Gaps = 1/108 (0%)
Frame = +1
Query: 862 NFWLFKEAGNLCKSLWIL*TKLYPFSLCYCI*IIALFGMAPIVCRNQC*NQEDVHRGSYG 921
N W+ +E NLCKSLW+L*T Y F+ Y I AL + P+ R+ + + S G
Sbjct: 22 NLWVPQEPRNLCKSLWLL*TN*YSFTPSYY*RIFALLCLPPVTKRS*QG*KNTICGPSDG 201
Query: 922 ACRTDPTEG-YNCCAWCYWSLYTTAKKVDYCGRVSGKSFNNVHG*ANF 968
R ++G Y+ + Y + T K+ D+ SFN+ HG* +F
Sbjct: 202 FGRVR*SQGCYSGASRSYRVVNRTKKEADHRC*ACC*SFNHFHG*THF 345
>TC229551 weakly similar to UP|Q6X4V5 (Q6X4V5) ABC transporter, partial (34%)
Length = 1436
Score = 48.9 bits (115), Expect = 1e-05
Identities = 32/95 (33%), Positives = 49/95 (50%)
Frame = +3
Query: 119 VEYILKRRKQQLNILQDVSGILKHSRLTLLLGPPNSGKTILLLALAGKLDPNLKIANEVQ 178
+E + K + IL ++G + RL ++GP SGK+ LL ALAG+L N+K
Sbjct: 36 LEATVTNGKNRKLILHGLTGYAQPGRLLAIIGPSGSGKSTLLDALAGRLTSNIK------ 197
Query: 179 FYEQFAGKVSYNGHEMNEFVPQRTAAYVSQNDTHL 213
GK+ NGH+ + T+ YV+Q+D L
Sbjct: 198 ----QTGKILINGHKQE--LAYGTSGYVTQDDAML 284
>BI699796 similar to PIR|G85167|G85 ABC transporter like protein [imported] -
Arabidopsis thaliana, partial (13%)
Length = 421
Score = 48.5 bits (114), Expect = 2e-05
Identities = 39/127 (30%), Positives = 61/127 (47%), Gaps = 1/127 (0%)
Frame = +3
Query: 869 AGNLCKSLWIL*TKLYPFSLCYCI*IIALFGMAPIVCRNQC*NQEDVHRGSYGACRTDPT 928
+GN+C S+ +L*TK + FS I +F +A N C N + + S+
Sbjct: 36 SGNIC*SIRLL*TK*HTFSKYNSRRICDVFCLASASFSN*CKN*S*ICK*SHSYY*A*WN 215
Query: 929 EGYNCCAWCY-WSLYTTAKKVDYCGRVSGKSFNNVHG*ANFRARCKICRYCYESH*KYSG 987
+ + Y W + T K D+ SFN+++G* ++R+RCK C ES +
Sbjct: 216 QRFFSRHAEY*WFID*TKKTADHSR*ACC*SFNHIYG*TDYRSRCKGSCSCNESSEECCW 395
Query: 988 EWKNCCL 994
WKNCC+
Sbjct: 396 NWKNCCM 416
>BU550490
Length = 616
Score = 48.1 bits (113), Expect = 2e-05
Identities = 23/32 (71%), Positives = 27/32 (83%)
Frame = +2
Query: 144 RLTLLLGPPNSGKTILLLALAGKLDPNLKIAN 175
RLTLLLGPP SGKT LL ALAGKLD +L++ +
Sbjct: 71 RLTLLLGPPRSGKTTLLQALAGKLDRDLRVGH 166
>TC218418 homologue to UP|Q6X4V5 (Q6X4V5) ABC transporter, partial (49%)
Length = 1220
Score = 42.7 bits (99), Expect = 0.001
Identities = 29/98 (29%), Positives = 48/98 (48%)
Frame = +1
Query: 117 DLVEYILKRRKQQLNILQDVSGILKHSRLTLLLGPPNSGKTILLLALAGKLDPNLKIANE 176
DL + + N+L+ ++G + T L+GP SGK+ LL AL+ +L N +
Sbjct: 253 DLTVMVTLSNGETQNVLEGLTGYAEPGTFTALMGPSGSGKSTLLDALSSRLAANAFL--- 423
Query: 177 VQFYEQFAGKVSYNGHEMNEFVPQRTAAYVSQNDTHLG 214
+G + NG + + TAAYV+Q+D +G
Sbjct: 424 -------SGTILLNGRKAK--LSFGTAAYVTQDDNLIG 510
>TC223422
Length = 426
Score = 39.3 bits (90), Expect = 0.011
Identities = 19/26 (73%), Positives = 22/26 (84%)
Frame = +2
Query: 147 LLLGPPNSGKTILLLALAGKLDPNLK 172
+ LGP +SGK ILLLALAG+LDP LK
Sbjct: 158 IALGPTSSGKNILLLALAGELDP*LK 235
>AW705086
Length = 427
Score = 38.5 bits (88), Expect = 0.018
Identities = 15/21 (71%), Positives = 20/21 (94%)
Frame = -3
Query: 80 RAGVDIPTIEVRFEHLNVQAQ 100
R G+DIPTIEVR++HLNV+A+
Sbjct: 242 RVGIDIPTIEVRYKHLNVEAE 180
>BI701600
Length = 426
Score = 38.1 bits (87), Expect = 0.024
Identities = 28/84 (33%), Positives = 45/84 (53%), Gaps = 1/84 (1%)
Frame = +3
Query: 128 QQLNILQDVSGILKHSRLTLLLGPPNSGKTILLLALAGKL-DPNLKIANEVQFYEQFAGK 186
++ IL+ V+GI + + +LGP SGK+ LL ALAG+L P L G
Sbjct: 60 KERTILKGVTGIAQPGEILAVLGPSGSGKSTLLHALAGRLHGPGL------------TGT 203
Query: 187 VSYNGHEMNEFVPQRTAAYVSQND 210
+ N ++ + V +RT +V+Q+D
Sbjct: 204 ILANSSKLTKPVLRRT-GFVTQDD 272
>BE803620 homologue to GP|13605839|gb| At2g01320/F10A8.20 {Arabidopsis
thaliana}, partial (8%)
Length = 413
Score = 38.1 bits (87), Expect = 0.024
Identities = 19/36 (52%), Positives = 25/36 (68%)
Frame = +3
Query: 132 ILQDVSGILKHSRLTLLLGPPNSGKTILLLALAGKL 167
+L++VSG K RL ++GP SGKT LL LAG+L
Sbjct: 294 LLKNVSGEAKPGRLLAIMGPSGSGKTTLLNVLAGQL 401
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.352 0.155 0.576
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 67,049,612
Number of Sequences: 63676
Number of extensions: 1174363
Number of successful extensions: 17706
Number of sequences better than 10.0: 69
Number of HSP's better than 10.0 without gapping: 9744
Number of HSP's successfully gapped in prelim test: 756
Number of HSP's that attempted gapping in prelim test: 7093
Number of HSP's gapped (non-prelim): 11927
length of query: 1226
length of database: 12,639,632
effective HSP length: 108
effective length of query: 1118
effective length of database: 5,762,624
effective search space: 6442613632
effective search space used: 6442613632
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.5 bits)
S2: 65 (29.6 bits)
Medicago: description of AC146819.12


