

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146818.6 + phase: 0 /pseudo
(642 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
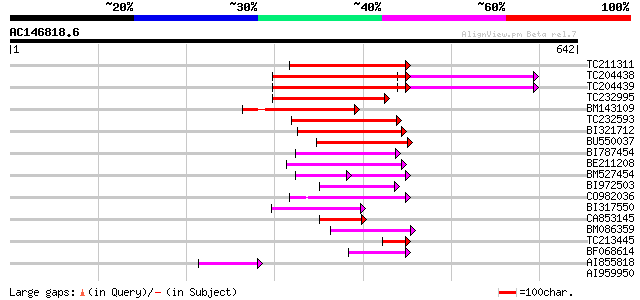
Sequences producing significant alignments: (bits) Value
TC211311 weakly similar to UP|O24587 (O24587) Pol protein, parti... 158 8e-39
TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, co... 154 9e-38
TC204439 UP|Q84VI4 (Q84VI4) Gag-pol polyprotein, complete 153 2e-37
TC232995 146 2e-35
BM143109 118 9e-27
TC232593 weakly similar to UP|Q9XG91 (Q9XG91) Tpv2-1c protein (F... 115 6e-26
BI321712 115 8e-26
BU550037 93 4e-19
BI787454 weakly similar to GP|21434|emb|CA ORF4 {Solanum tuberos... 78 1e-14
BE211208 68 1e-11
BM527454 weakly similar to GP|27901709|gb| gag-pol polyprotein {... 47 2e-10
BI972503 weakly similar to GP|18378607|gb polyprotein {Oryza sat... 63 4e-10
CO982036 62 1e-09
BI317550 weakly similar to GP|9759590|dbj| polyprotein-like {Ara... 56 4e-08
CA853145 weakly similar to GP|27901709|gb| gag-pol polyprotein {... 54 2e-07
BM086359 50 4e-06
TC213445 49 5e-06
BF068614 similar to GP|27817858|db OJ1081_B12.7 {Oryza sativa (j... 45 8e-05
AI855818 weakly similar to GP|21741393|e OSJNBb0051N19.6 {Oryza ... 42 6e-04
AI959950 41 0.001
>TC211311 weakly similar to UP|O24587 (O24587) Pol protein, partial (15%)
Length = 1213
Score = 158 bits (399), Expect = 8e-39
Identities = 77/137 (56%), Positives = 100/137 (72%)
Frame = +3
Query: 318 KKYILVVQIYVDDIIFGSTNASLCKEFSK*MQDEFEKSMMGELKFFLGIQINQRKDGVYV 377
K+ L++ IYVDDIIFG+T+ +CKEF + M+D FE SM GELKF LG+QI Q+ G+++
Sbjct: 477 KETFLIIHIYVDDIIFGATSKRMCKEFFELMKDGFETSMKGELKFLLGLQIIQKVYGIFI 656
Query: 378 HQTKYTKELLKKFKLEDCKVMNTPMHPTCT*SKEDIGTKVDQKLYIGMIGSLLYLTASRP 437
HQ KYTK LK+F++++ K M TPMH + K++ G K Y GMI SL YLT+SRP
Sbjct: 657 HQEKYTKSHLKRFRMDEAKPMATPMHRSTIIDKDEKGNHTS*KEYSGMIDSLSYLTSSRP 836
Query: 438 DILFSVCLCARFQSNPK 454
DI+F VCLCARFQS PK
Sbjct: 837 DIVFVVCLCARFQSYPK 887
>TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, complete
Length = 4734
Score = 154 bits (390), Expect = 9e-38
Identities = 73/157 (46%), Positives = 108/157 (68%)
Frame = +1
Query: 298 LIKNDFKRGQVDTTLFRGTLKKYILVVQIYVDDIIFGSTNASLCKEFSK*MQDEFEKSMM 357
L + +++G +D TLF + +++ QIYVDDI+FG + + + F + MQ EFE S++
Sbjct: 3697 LTQQGYRKGGIDKTLFVKQDAENLMIAQIYVDDIVFGGMSNEMLRHFVQQMQSEFEMSLV 3876
Query: 358 GELKFFLGIQINQRKDGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCT*SKEDIGTKV 417
GEL +FLG+Q+ Q +D +++ Q+KY K ++KKF +E+ TP SK++ GT V
Sbjct: 3877 GELTYFLGLQVKQMEDSIFLSQSKYAKNIVKKFGMENASHKRTPAPTHLKLSKDEAGTSV 4056
Query: 418 DQKLYIGMIGSLLYLTASRPDILFSVCLCARFQSNPK 454
DQ LY MIGSLLYLTASRPDI ++V +CAR+Q+NPK
Sbjct: 4057 DQSLYRSMIGSLLYLTASRPDITYAVGVCARYQANPK 4167
Score = 64.7 bits (156), Expect = 1e-10
Identities = 51/159 (32%), Positives = 81/159 (50%)
Frame = +3
Query: 440 LFSVCLCARFQSNPKRISFNCC*ENLHVFEGNN*SWTALYEIPRLSSGWIL*C*LCW*QD 499
L S CLC + S S EN + + + * W + + R ++GW+L*C*L W
Sbjct: 4125 LCSRCLC-KISSQS*DKSLESSKENSEICKWHQ*LWDYVLSLFRFNAGWVL*C*LGWKCR 4301
Query: 500 *MKINQWKLSVPGRESDIMG*QKTGNHCYVYSKSIIHFSCKLLYTTTLDETSVGRLSDKC 559
* K + W + + G +S M Q+ +Y S ++ S K L+TT+LDE + +
Sbjct: 4302 *QKKHFWWMFLFGNQSYFMVQQEAELCVPIYC*SRVYCSRKQLFTTSLDEADAEGVQCRT 4481
Query: 560 *QYSHLL**YYCYLFVKESNSTFKSQAYINQTPLYQRLC 598
+ +L* + CY + +S ST ++QA+ + T LY R C
Sbjct: 4482 RCHDIVL*QHECY*YF*KSCSTQQNQAH*H*TSLY*RSC 4598
>TC204439 UP|Q84VI4 (Q84VI4) Gag-pol polyprotein, complete
Length = 4731
Score = 153 bits (387), Expect = 2e-37
Identities = 72/157 (45%), Positives = 108/157 (67%)
Frame = +1
Query: 298 LIKNDFKRGQVDTTLFRGTLKKYILVVQIYVDDIIFGSTNASLCKEFSK*MQDEFEKSMM 357
L + +++G +D TLF + +++ QIYVDDI+FG + + + F + MQ EFE S++
Sbjct: 3694 LTQQGYRKGGIDKTLFVKQDAENLMIAQIYVDDIVFGGMSNEMLRHFVQQMQSEFEMSLV 3873
Query: 358 GELKFFLGIQINQRKDGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCT*SKEDIGTKV 417
GEL +FLG+Q+ Q +D +++ Q++Y K ++KKF +E+ TP SK++ GT V
Sbjct: 3874 GELTYFLGLQVKQMEDSIFLSQSRYAKNIVKKFGMENASHKRTPAPTHLKLSKDEAGTSV 4053
Query: 418 DQKLYIGMIGSLLYLTASRPDILFSVCLCARFQSNPK 454
DQ LY MIGSLLYLTASRPDI ++V +CAR+Q+NPK
Sbjct: 4054 DQSLYRSMIGSLLYLTASRPDITYAVGVCARYQANPK 4164
Score = 68.2 bits (165), Expect = 1e-11
Identities = 50/159 (31%), Positives = 85/159 (53%)
Frame = +3
Query: 440 LFSVCLCARFQSNPKRISFNCC*ENLHVFEGNN*SWTALYEIPRLSSGWIL*C*LCW*QD 499
L S CLC + S + S + EN + + + * W + + + ++GW+L*C*L W
Sbjct: 4122 LCSRCLC-KISSQSQDKSLDSSKENSEICKWH**LWDYVLSLFKSNAGWVL*C*LGWKCR 4298
Query: 500 *MKINQWKLSVPGRESDIMG*QKTGNHCYVYSKSIIHFSCKLLYTTTLDETSVGRLSDKC 559
* K + W + + G++ M Q+ +YS+S ++ S K L+T +LDE + +
Sbjct: 4299 *QKKHFWWMLLFGKQPYFMVQQEAELCVPIYSRSRVYCSRKQLFTASLDEADAEGVQCRT 4478
Query: 560 *QYSHLL**YYCYLFVKESNSTFKSQAYINQTPLYQRLC 598
+ +L* + CY + +S ST ++QA+ + T LYQR C
Sbjct: 4479 RCHDIVL*QHECY*YF*KSCSTQQNQAH*H*TSLYQRSC 4595
>TC232995
Length = 1009
Score = 146 bits (369), Expect = 2e-35
Identities = 73/133 (54%), Positives = 96/133 (71%)
Frame = +2
Query: 298 LIKNDFKRGQVDTTLFRGTLKKYILVVQIYVDDIIFGSTNASLCKEFSK*MQDEFEKSMM 357
L++ +F RG+VDTTLF IL+VQIYVDDIIFGSTN SLCKEFS MQ EFE SMM
Sbjct: 122 LLEKEFSRGKVDTTLFIKRKHNDILLVQIYVDDIIFGSTNDSLCKEFSLDMQSEFEMSMM 301
Query: 358 GELKFFLGIQINQRKDGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCT*SKEDIGTKV 417
GELK+FLG+QI Q + G++++Q+KY KEL+K+F ++ K M+TPM C K++ G +
Sbjct: 302 GELKYFLGLQIKQTQ*GIFINQSKYCKELIKRFGMDSAKHMSTPMSTNCYLDKDESGQSI 481
Query: 418 DQKLYIGMIGSLL 430
D K Y IG ++
Sbjct: 482 DIKQYRDAIGEVV 520
>BM143109
Length = 415
Score = 118 bits (295), Expect = 9e-27
Identities = 66/133 (49%), Positives = 88/133 (65%)
Frame = +1
Query: 264 NKVTVNKKVLSTLKNLLQLQDWKQSGYFYPMQ*ILIKNDFKRGQVDTTLFRGTLKKYILV 323
N V KKVL LK L+ ++ + L+ F +G+VDT LF IL+
Sbjct: 37 NHVFKLKKVLYGLKQALR-------AWYELLSKFLLDKGFSKGKVDTNLFI*KKLNDILL 195
Query: 324 VQIYVDDIIFGSTNASLCKEFSK*MQDEFEKSMMGELKFFLGIQINQRKDGVYVHQTKYT 383
VQIYVDDIIFGSTN SLCK+FS+ MQ+EFE SMM EL FFLG+QI Q K+G+++ Q+KY
Sbjct: 196 VQIYVDDIIFGSTNDSLCKKFSQDMQNEFEMSMMRELNFFLGLQIKQTKNGIFISQSKYC 375
Query: 384 KELLKKFKLEDCK 396
K+L+ +F +E+ K
Sbjct: 376 KDLIHRFGMENDK 414
>TC232593 weakly similar to UP|Q9XG91 (Q9XG91) Tpv2-1c protein (Fragment),
partial (16%)
Length = 562
Score = 115 bits (288), Expect = 6e-26
Identities = 55/124 (44%), Positives = 85/124 (68%)
Frame = +1
Query: 320 YILVVQIYVDDIIFGSTNASLCKEFSK*MQDEFEKSMMGELKFFLGIQINQRKDGVYVHQ 379
Y L+V +YVDD++ +A L +EF + M FE + +G + +FLGI+I Q ++ V + Q
Sbjct: 187 YFLIVSLYVDDLLVTRDDARLVEEFKQEMMQAFEMTNLGLMTYFLGIEIKQSQNKVLICQ 366
Query: 380 TKYTKELLKKFKLEDCKVMNTPMHPTCT*SKEDIGTKVDQKLYIGMIGSLLYLTASRPDI 439
KY KE+LKKF++E+CK ++TPM+ +K D K+D+ Y +IG L+YLTA+RPDI
Sbjct: 367 RKYAKEILKKFQMEECKSVSTPMNQKEKFNKVDGADKIDEGYYRSLIGCLMYLTATRPDI 546
Query: 440 LFSV 443
LF++
Sbjct: 547 LFAI 558
>BI321712
Length = 399
Score = 115 bits (287), Expect = 8e-26
Identities = 56/124 (45%), Positives = 81/124 (65%)
Frame = -3
Query: 326 IYVDDIIFGSTNASLCKEFSK*MQDEFEKSMMGELKFFLGIQINQRKDGVYVHQTKYTKE 385
+YVDD+IF N S+ +EF K M +EFE + MG + ++LGI++ Q G+++ Q Y KE
Sbjct: 379 LYVDDLIFTGNNPSMFEEFKKDMSNEFEMTDMGLMAYYLGIEVKQEDKGIFITQEGYAKE 200
Query: 386 LLKKFKLEDCKVMNTPMHPTCT*SKEDIGTKVDQKLYIGMIGSLLYLTASRPDILFSVCL 445
+LKKFK++D + TPM SK + G VD LY +IGSL YLT +RPDIL+ V +
Sbjct: 199 VLKKFKMDDANPVGTPMECGSKLSKHEKGENVDPTLYKSLIGSLRYLTCTRPDILYVVGV 20
Query: 446 CARF 449
+R+
Sbjct: 19 VSRY 8
>BU550037
Length = 728
Score = 92.8 bits (229), Expect = 4e-19
Identities = 43/109 (39%), Positives = 74/109 (67%)
Frame = -3
Query: 348 MQDEFEKSMMGELKFFLGIQINQRKDGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCT 407
M+ EFE + +G++K+FLG+ I Q +DG+++ Q KY E+L+KF +E CK + T +
Sbjct: 363 MESEFEMTDLGQMKYFLGM*IFQSEDGIFISQKKYAWEILRKFHMERCKPIATVLVVNEK 184
Query: 408 *SKEDIGTKVDQKLYIGMIGSLLYLTASRPDILFSVCLCARFQSNPKRI 456
SK++ + D +Y +IGSLLYL+A+RP+++F+ L +RF +P ++
Sbjct: 183 FSKDEEDNQGDASVYRSLIGSLLYLSATRPNLMFAATLLSRFTKSPSQV 37
>BI787454 weakly similar to GP|21434|emb|CA ORF4 {Solanum tuberosum}, partial
(21%)
Length = 421
Score = 78.2 bits (191), Expect = 1e-14
Identities = 37/119 (31%), Positives = 70/119 (58%)
Frame = +2
Query: 324 VQIYVDDIIFGSTNASLCKEFSK*MQDEFEKSMMGELKFFLGIQINQRKDGVYVHQTKYT 383
+ +YVDDI+ +A+ + + + + F+ + LK+FLGI++ Q DGV + Q KY
Sbjct: 65 LMVYVDDIMITKKDATKIVQLKEHLFNHFQTKDLRYLKYFLGIEVAQSGDGVVISQRKYA 244
Query: 384 KELLKKFKLEDCKVMNTPMHPTCT*SKEDIGTKVDQKLYIGMIGSLLYLTASRPDILFS 442
++L++ +++C+++++PM P D + Y ++G L+YLT +RPDI F+
Sbjct: 245 LDILEETGMQNCRLVDSPMDPNLKLMAYQSEVYPDPERYRRLVGKLIYLTITRPDISFA 421
>BE211208
Length = 413
Score = 68.2 bits (165), Expect = 1e-11
Identities = 40/137 (29%), Positives = 70/137 (50%), Gaps = 1/137 (0%)
Frame = +2
Query: 314 RGTLKKYILVVQIYVDDIIFGSTNASLCKEFSK*MQDEFEKSMMGELKFFLGIQINQRKD 373
RG + ++ + +YVDDII + L + + F +G+L +FLGI+++
Sbjct: 2 RGKKDRNLVYLLVYVDDIIITGRSNYLIQSLVHHLNSNFSLKQLGQLDYFLGIEVHHTPT 181
Query: 374 G-VYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCT*SKEDIGTKVDQKLYIGMIGSLLYL 432
G V + Q+KY +LL K + + K +++PM SK D +Y ++G+L Y
Sbjct: 182 GSVLLTQSKYICDLLHKTDMAEAKPISSPMVTNLRLSKNGDDLLSDPTMYRSVVGALQYP 361
Query: 433 TASRPDILFSVCLCARF 449
T +RP+I F+ +F
Sbjct: 362 TITRPEISFAANKVCQF 412
>BM527454 weakly similar to GP|27901709|gb| gag-pol polyprotein {Vitis
vinifera}, partial (19%)
Length = 437
Score = 47.0 bits (110), Expect(2) = 2e-10
Identities = 19/64 (29%), Positives = 36/64 (55%)
Frame = +2
Query: 324 VQIYVDDIIFGSTNASLCKEFSK*MQDEFEKSMMGELKFFLGIQINQRKDGVYVHQTKYT 383
+ +YVDDI+ + + + F+ +G+ ++FLGI++ Q KDG+ + Q KY
Sbjct: 41 LMVYVDDIVITGNDQGKIAQLKGHLFSHFQTKDLGKFEYFLGIEVAQSKDGIIISQRKYA 220
Query: 384 KELL 387
++L
Sbjct: 221 LDIL 232
Score = 36.6 bits (83), Expect(2) = 2e-10
Identities = 21/72 (29%), Positives = 39/72 (54%), Gaps = 1/72 (1%)
Frame = +1
Query: 383 TKEL-LKKFKLEDCKVMNTPMHPTCT*SKEDIGTKVDQKLYIGMIGSLLYLTASRPDILF 441
TKE+ ++ + DC+ +++ M P D + Y ++G L+YLT +RP+I F
Sbjct: 205 TKEVCIRHTGMSDCRPIDSLMDPNKKLLPNQGKPYSDSERYRILVGKLIYLTITRPNISF 384
Query: 442 SVCLCARFQSNP 453
V + ++F +P
Sbjct: 385 VVGVVSQFMQSP 420
>BI972503 weakly similar to GP|18378607|gb polyprotein {Oryza sativa
(japonica cultivar-group)}, partial (5%)
Length = 327
Score = 63.2 bits (152), Expect = 4e-10
Identities = 30/90 (33%), Positives = 53/90 (58%)
Frame = +2
Query: 352 FEKSMMGELKFFLGIQINQRKDGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCT*SKE 411
F+ +G LK+FLGI++ Q V + Q KY ++L++ +++C+ +++PM P +
Sbjct: 56 FQTKDLGYLKYFLGIEVAQSGVDVVISQRKYALDILEETGMQNCRPVDSPMDPNLKLMAD 235
Query: 412 DIGTKVDQKLYIGMIGSLLYLTASRPDILF 441
D + Y ++G L+YLT +RPDI F
Sbjct: 236 QSEIYHDPERYRRLVGKLIYLTITRPDISF 325
>CO982036
Length = 674
Score = 61.6 bits (148), Expect = 1e-09
Identities = 41/138 (29%), Positives = 72/138 (51%), Gaps = 2/138 (1%)
Frame = -2
Query: 318 KKYILVVQ--IYVDDIIFGSTNASLCKEFSK*MQDEFEKSMMGELKFFLGIQINQRKDGV 375
K +IL V +YVD II GS+ +L + + + F ++G+L +F+ I++ D +
Sbjct: 670 KTHILTVYLLVYVDIIITGSS-CTLIQNLTSKLNSSFPLKLLGKLDYFVEIEVKSMPDLL 494
Query: 376 YVHQTKYTKELLKKFKLEDCKVMNTPMHPTCT*SKEDIGTKVDQKLYIGMIGSLLYLTAS 435
+ +T + +K + + + +++PM TC SK D Y ++G+L Y T
Sbjct: 493 FSLRTSIFEIFCRKPR*Q-AQPISSPMTTTCKLSKSDSDLFSGPTFYRSVVGALQYTTVI 317
Query: 436 RPDILFSVCLCARFQSNP 453
RP+I F+V +F SNP
Sbjct: 316 RPEISFAVNKVCQFMSNP 263
>BI317550 weakly similar to GP|9759590|dbj| polyprotein-like {Arabidopsis
thaliana}, partial (18%)
Length = 421
Score = 56.2 bits (134), Expect = 4e-08
Identities = 31/106 (29%), Positives = 53/106 (49%)
Frame = -2
Query: 297 ILIKNDFKRGQVDTTLFRGTLKKYILVVQIYVDDIIFGSTNASLCKEFSK*MQDEFEKSM 356
+L+K + + D +LF T + +YVDDII + + F+
Sbjct: 363 LLLKEGYIQSISDYSLFTLTKGNTFTALLVYVDDIILAGDSIDEFDRIKNVLDLAFKIKN 184
Query: 357 MGELKFFLGIQINQRKDGVYVHQTKYTKELLKKFKLEDCKVMNTPM 402
+G+LK+FLG+++ + G+ + Q KY +LLK L CK +TP+
Sbjct: 183 LGKLKYFLGLEVAHSRLGITISQRKYCLDLLKDSGLLGCKPASTPL 46
>CA853145 weakly similar to GP|27901709|gb| gag-pol polyprotein {Vitis
vinifera}, partial (29%)
Length = 563
Score = 53.9 bits (128), Expect = 2e-07
Identities = 22/53 (41%), Positives = 37/53 (69%)
Frame = +3
Query: 352 FEKSMMGELKFFLGIQINQRKDGVYVHQTKYTKELLKKFKLEDCKVMNTPMHP 404
F+ +G+LK+FLGI++ Q K G+ + Q KY ++LK +++CK ++TPM P
Sbjct: 72 FQTKDLGQLKYFLGIEVAQSKTGIAICQRKYALDILKDTGMQNCKPVDTPMDP 230
>BM086359
Length = 427
Score = 49.7 bits (117), Expect = 4e-06
Identities = 30/96 (31%), Positives = 46/96 (47%)
Frame = +1
Query: 364 LGIQINQRKDGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCT*SKEDIGTKVDQKLYI 423
LGI + Q G+ + Q KY ++L + + DC NTPM P D
Sbjct: 1 LGIDVAQSSYGIVISQWKYALDILTETGMLDCLPSNTPMDPNVKLLSGQGEALEDPGR*C 180
Query: 424 GMIGSLLYLTASRPDILFSVCLCARFQSNPKRISFN 459
++G L YLT +R DI F+V + ++F +P +N
Sbjct: 181 CLVGRLNYLTVTRLDITFAVGVLSQFLKDPTDSQWN 288
>TC213445
Length = 705
Score = 49.3 bits (116), Expect = 5e-06
Identities = 20/32 (62%), Positives = 26/32 (80%)
Frame = +2
Query: 423 IGMIGSLLYLTASRPDILFSVCLCARFQSNPK 454
+ MI S LYL+ SRP I+FSVC+C R+Q+NPK
Sbjct: 191 LAMIESFLYLSTSRPHIMFSVCMCVRYQANPK 286
Score = 39.3 bits (90), Expect = 0.005
Identities = 29/79 (36%), Positives = 43/79 (53%)
Frame = +3
Query: 515 SDIMG*QKTGNHCYVYSKSIIHFSCKLLYTTTLDETSVGRLSDKC*QYSHLL**YYCYLF 574
S IM * K C + +S I+F KLL T LDET+ L + * Y++ +* Y C
Sbjct: 450 SSIMA**KAK*CCLINCRSRIYFC*KLLCTNLLDETTTF*LWFET*SYTYPM*QYKCN*S 629
Query: 575 VKESNSTFKSQAYINQTPL 593
+++S S ++AY N+ L
Sbjct: 630 IQKSYSVL*NKAY*NKASL 686
>BF068614 similar to GP|27817858|db OJ1081_B12.7 {Oryza sativa (japonica
cultivar-group)}, partial (3%)
Length = 413
Score = 45.4 bits (106), Expect = 8e-05
Identities = 23/70 (32%), Positives = 39/70 (54%)
Frame = +1
Query: 384 KELLKKFKLEDCKVMNTPMHPTCT*SKEDIGTKVDQKLYIGMIGSLLYLTASRPDILFSV 443
++LL K K+ + + +++PM +C SK D LY ++G+L Y T +RP+ +SV
Sbjct: 199 RDLLPKTKMAEAQPISSPMVSSCKLSKSGSDLFQDPTLYGSVVGALQYATLTRPEFSYSV 378
Query: 444 CLCARFQSNP 453
+F S P
Sbjct: 379 NKVCQFMSQP 408
Score = 36.2 bits (82), Expect = 0.046
Identities = 21/80 (26%), Positives = 39/80 (48%), Gaps = 1/80 (1%)
Frame = +2
Query: 309 DTTLFRGTLKKYILVVQIYVDDIIFGSTNASLCKEFSK*MQDEFEKSMMGELKFFLGIQI 368
D +LF ++Y + + +YVDDI L ++ + +F +G L +FLGI++
Sbjct: 2 DPSLFVYKQQQYTVFLLVYVDDITITGNCVLLIQQLISQLNSQFALKQLGLLDYFLGIEV 181
Query: 369 NQRKD-GVYVHQTKYTKELL 387
D G+ + K+ + L
Sbjct: 182 KYLPDKGISCPRLKWQRHNL 241
>AI855818 weakly similar to GP|21741393|e OSJNBb0051N19.6 {Oryza sativa
(japonica cultivar-group)}, partial (10%)
Length = 463
Score = 42.4 bits (98), Expect = 6e-04
Identities = 30/73 (41%), Positives = 41/73 (56%)
Frame = -1
Query: 214 CKKS*ISFKEMMCEIWYPNLSRRTLLEQNGYSETS*MNRVK*SETKLYLLNKVTVNKKVL 273
CKK+*I+ KE+MC NL EQNG+ E +*MN E +L K + K+
Sbjct: 454 CKKN*INLKEIMCGN**KNLKIILS*EQNGFLEIN*MNMA*LLEIRLD**QKGIIKKRE* 275
Query: 274 STLKNLLQLQDWK 286
+ K++LQLQD K
Sbjct: 274 TMKKHMLQLQD*K 236
>AI959950
Length = 466
Score = 41.2 bits (95), Expect = 0.001
Identities = 34/86 (39%), Positives = 45/86 (51%)
Frame = -2
Query: 214 CKKS*ISFKEMMCEIWYPNLSRRTLLEQNGYSETS*MNRVK*SETKLYLLNKVTVNKKVL 273
CKK+ ISFK +M R LE NGY T+* V+ +TK L KVT N+KV
Sbjct: 387 CKKNLISFKRIMSRSSLNYQKERR*LE*NGYFVTN*TRMVRL*DTKQD*LLKVTHNRKV* 208
Query: 274 STLKNLLQLQDWKQSGYFYPMQ*ILI 299
+T K L L K ++ +Q I+I
Sbjct: 207 TTQKPLHLLHV*K*YASYFHLQPIVI 130
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.352 0.155 0.517
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 27,388,148
Number of Sequences: 63676
Number of extensions: 377616
Number of successful extensions: 3197
Number of sequences better than 10.0: 55
Number of HSP's better than 10.0 without gapping: 2385
Number of HSP's successfully gapped in prelim test: 81
Number of HSP's that attempted gapping in prelim test: 786
Number of HSP's gapped (non-prelim): 2519
length of query: 642
length of database: 12,639,632
effective HSP length: 103
effective length of query: 539
effective length of database: 6,081,004
effective search space: 3277661156
effective search space used: 3277661156
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (22.0 bits)
S2: 62 (28.5 bits)
Medicago: description of AC146818.6


