

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
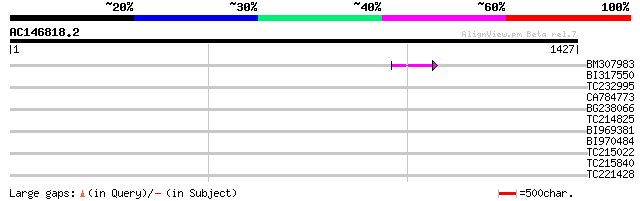
Query= AC146818.2 - phase: 0 /pseudo
(1427 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BM307983 44 5e-04
BI317550 weakly similar to GP|9759590|dbj| polyprotein-like {Ara... 38 0.036
TC232995 37 0.048
CA784773 weakly similar to GP|27901698|gb gag-pol polyprotein {V... 34 0.40
BG238066 similar to SP|P49199|RS8_ 40S ribosomal protein S8. [Ri... 33 0.90
TC214825 similar to UP|RS8_ORYSA (P49199) 40S ribosomal protein ... 33 1.2
BI969381 31 4.5
BI970484 30 5.8
TC215022 homologue to UP|PSAG_ARATH (Q9S7N7) Photosystem I react... 30 5.8
TC215840 homologue to UP|Q41122 (Q41122) Proline-rich protein pr... 30 5.8
TC221428 similar to UP|BRH2_DROME (Q24256) Homeobox B-H2 protein... 30 9.9
>BM307983
Length = 406
Score = 43.9 bits (102), Expect = 5e-04
Identities = 41/119 (34%), Positives = 56/119 (46%), Gaps = 4/119 (3%)
Frame = +3
Query: 961 VVNGYIRSKDMQMVP*NVSKHVWLLKVLIKLKGLIILKHSPLLPS---YPPLGYCLLLHL 1017
VV GYI+ M + WL + K G I+ +H LL Y + L
Sbjct: 6 VVGGYIQLNIRLMTHWTGIRRGWLQRDTSKPMGSIMRRH--LLSGKNEYSQDHHLLSSKH 179
Query: 1018 FMVGICINLMSITPSCMEIYMKQYI*K-SLKVLLLPNLDKFASSRNPFMVLSKQVGNGL 1075
+VG CINLM PS ME + K+Y + L ++ + KFA R P+MV S +G GL
Sbjct: 180 NLVGRCINLMLKMPSYMEAWKKKYTWRFHLDMVPVMEEIKFAD*RRPYMVSSSLLGLGL 356
>BI317550 weakly similar to GP|9759590|dbj| polyprotein-like {Arabidopsis
thaliana}, partial (18%)
Length = 421
Score = 37.7 bits (86), Expect = 0.036
Identities = 22/51 (43%), Positives = 30/51 (58%)
Frame = -3
Query: 1068 SKQVGNGLRSLLSFFMLRDSLKLMQITHYLLKSLLLPTLWFWFMWMTLF*Q 1118
SKQVG+G++S L F + + QIT YLL + +L W+M MT F Q
Sbjct: 401 SKQVGSGMKS*LICFSKKVTYNQFQITPYLLSPKEILSLHCWYM*MTSFLQ 249
>TC232995
Length = 1009
Score = 37.4 bits (85), Expect = 0.048
Identities = 21/71 (29%), Positives = 38/71 (52%)
Frame = +3
Query: 1046 LKVLLLPNLDKFASSRNPFMVLSKQVGNGLRSLLSFFMLRDSLKLMQITHYLLKSLLLPT 1105
L + L N F + + FMV +K +G+G+ + FF+ ++S ++ I HY + ++
Sbjct: 15 LVLKFLINQTMFINYKRLFMV*NKPLGHGMND*VIFFLKKNSPEVKWIPHYS*RESIMIF 194
Query: 1106 LWFWFMWMTLF 1116
WF +M M F
Sbjct: 195 CWFKYMLMI*F 227
>CA784773 weakly similar to GP|27901698|gb gag-pol polyprotein {Vitis
vinifera}, partial (34%)
Length = 409
Score = 34.3 bits (77), Expect = 0.40
Identities = 41/127 (32%), Positives = 60/127 (46%), Gaps = 3/127 (2%)
Frame = +1
Query: 919 PLVIFQLVKILGG*LLCRQR*RLLMQIILGNLLIFLQMLYP*VVNGYIRSKDMQMVP*NV 978
PL + L+ IL G L + RLL ++LG+L + L + *V +G+ K MV
Sbjct: 25 PLFVRHLI-ILAGDKLWLMKCRLLKIMVLGSLFLSLLVRRL*VADGFTLLKLGPMVRLIG 201
Query: 979 SKHVWLLKVLIKLKGL--IILKHSPLLPSYPPLGYC-LLLHLFMVGICINLMSITPSCME 1035
+ W L+ + L +IL LL + PL C LL LF G I+L+ PS
Sbjct: 202 LRLAWSLRATHRYMALNTVILS---LLYFFSPLFVCSLLWRLFDTGPFISLILRMPSSTV 372
Query: 1036 IYMKQYI 1042
I + +I
Sbjct: 373 ILRRIFI 393
>BG238066 similar to SP|P49199|RS8_ 40S ribosomal protein S8. [Rice] {Oryza
sativa}, partial (66%)
Length = 511
Score = 33.1 bits (74), Expect = 0.90
Identities = 21/59 (35%), Positives = 33/59 (55%), Gaps = 1/59 (1%)
Frame = -1
Query: 805 LSIPLVLLQFPLHIFTLIVKLLTLCLLMTLL-*ILLKIMMRFLMKIHLLILKKGMMKWS 862
LSI + LLQFPLH+ TL + + TLL +L + F ++ ++LK + +WS
Sbjct: 499 LSIYVALLQFPLHMVTLRGVFCSCSSITTLLCSFVLGSCLLFPTNLNTIVLKIPLFEWS 323
>TC214825 similar to UP|RS8_ORYSA (P49199) 40S ribosomal protein S8, partial
(95%)
Length = 860
Score = 32.7 bits (73), Expect = 1.2
Identities = 22/59 (37%), Positives = 32/59 (53%), Gaps = 1/59 (1%)
Frame = -2
Query: 805 LSIPLVLLQFPLHIFTLIVKLLTLCLLMTLL-*ILLKIMMRFLMKIHLLILKKGMMKWS 862
LSI L LLQF LH+ TL + + TLL LL + F ++ ++LK + +WS
Sbjct: 496 LSIFLALLQFSLHMVTLFGFFCSCSSITTLLCSFLLGSCLLFPTNLNTIVLKIPLFEWS 320
>BI969381
Length = 602
Score = 30.8 bits (68), Expect = 4.5
Identities = 21/73 (28%), Positives = 36/73 (48%), Gaps = 2/73 (2%)
Frame = +2
Query: 1049 LLLPNLDKFASSRNPFMVLSKQVGNGLRSLLSFFMLRDSLKLMQIT--HYLLKSLLLPTL 1106
L L N SS F++LS N L +F +R S + + H + S+ LPT+
Sbjct: 308 LEL*NATSMTSSSALFLLLSLIFPNLANILSNFHSIRRSTREPSLCEPHSIKSSISLPTM 487
Query: 1107 WFWFMWMTLF*QE 1119
W +W+++ *++
Sbjct: 488 REWHLWVSIL*EK 526
>BI970484
Length = 797
Score = 30.4 bits (67), Expect = 5.8
Identities = 18/54 (33%), Positives = 26/54 (47%), Gaps = 8/54 (14%)
Frame = -1
Query: 1063 PFMVLSKQVGNGLRSLLSFFMLRDSLKLMQITHYLLKSLL--------LPTLWF 1108
PF + S + G L SL FF++ + L+ + H L L+ LP LWF
Sbjct: 203 PFALASLRAGKNLLSLNLFFLILNCHPLLGLMHLLKAGLIHFLCIYTQLPCLWF 42
>TC215022 homologue to UP|PSAG_ARATH (Q9S7N7) Photosystem I reaction center
subunit V, chloroplast precursor (PSI-G), partial (63%)
Length = 1012
Score = 30.4 bits (67), Expect = 5.8
Identities = 13/35 (37%), Positives = 21/35 (59%)
Frame = +2
Query: 1077 SLLSFFMLRDSLKLMQITHYLLKSLLLPTLWFWFM 1111
++++ M R SLK+ Q+ H L+ SL +WF M
Sbjct: 686 NVINVMMYRFSLKIQQVQHMLI*SLFSAAIWFAIM 790
>TC215840 homologue to UP|Q41122 (Q41122) Proline-rich protein precursor,
partial (74%)
Length = 1171
Score = 30.4 bits (67), Expect = 5.8
Identities = 26/118 (22%), Positives = 58/118 (49%)
Frame = +3
Query: 804 FLSIPLVLLQFPLHIFTLIVKLLTLCLLMTLL*ILLKIMMRFLMKIHLLILKKGMMKWSL 863
FL + + PL + +L++ +T+ LL+ +L+ ++ RF+++++LL +++L
Sbjct: 282 FLHLLTTITLLPLPLLSLLLTTITITLLL----LLIHLLFRFILQLNLL------FRFTL 431
Query: 864 *PPENPPEVLNHQFTYKIMSVIQQVVITL*QILLVIQLYLLNTVLMLTL*ILRLSPLV 921
LN F + + + L ++ L++ L L L+L L + R +P +
Sbjct: 432 --------QLNLLFRFTLQ------LNLLFRLTLLLNLRLFRFTLLLNLRLFRFTPFL 563
>TC221428 similar to UP|BRH2_DROME (Q24256) Homeobox B-H2 protein (Homeobox
BarH2 protein), partial (4%)
Length = 1188
Score = 29.6 bits (65), Expect = 9.9
Identities = 18/55 (32%), Positives = 29/55 (52%), Gaps = 11/55 (20%)
Frame = +2
Query: 1155 PQKAFLSVNANTVWSFLMMLA*LVANQFLLLLIL-----------LYVYLKILGL 1198
P A L ++ WS L++L L ++FLLLLI+ LY+++ +L L
Sbjct: 524 PTMAILLALSSIYWSLLLLLLPLPLSRFLLLLIIITTPK*YLLLPLYLFIALLSL 688
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.370 0.167 0.644
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 74,542,552
Number of Sequences: 63676
Number of extensions: 1224623
Number of successful extensions: 18491
Number of sequences better than 10.0: 23
Number of HSP's better than 10.0 without gapping: 7607
Number of HSP's successfully gapped in prelim test: 831
Number of HSP's that attempted gapping in prelim test: 10281
Number of HSP's gapped (non-prelim): 9716
length of query: 1427
length of database: 12,639,632
effective HSP length: 109
effective length of query: 1318
effective length of database: 5,698,948
effective search space: 7511213464
effective search space used: 7511213464
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.8 bits)
S2: 65 (29.6 bits)
Medicago: description of AC146818.2


