

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146817.2 + phase: 0
(145 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
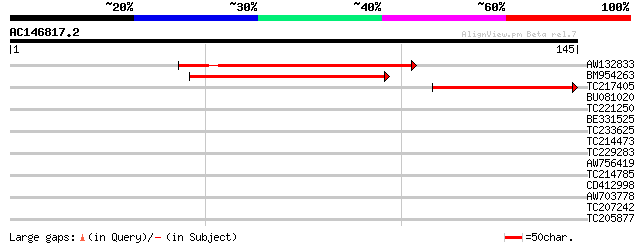
Score E
Sequences producing significant alignments: (bits) Value
AW132833 69 8e-13
BM954263 68 1e-12
TC217405 similar to UP|Q9FLL9 (Q9FLL9) Gb|AAB80666.1, partial (20%) 54 3e-08
BU081020 31 0.18
TC221250 weakly similar to GB|AAP37739.1|30725434|BT008380 At5g0... 31 0.24
BE331525 30 0.31
TC233625 similar to UP|Q85DB9 (Q85DB9) NADH dehydrogenase subuni... 30 0.31
TC214473 similar to UP|Q39449 (Q39449) Specific tissue protein 1... 27 3.4
TC229283 homologue to UP|P93078 (P93078) Ap19 protein, complete 27 3.4
AW756419 27 4.5
TC214785 similar to UP|Q6V8T3 (Q6V8T3) Chlorophyll a/b-binding p... 27 4.5
CD412998 26 5.9
AW703778 26 5.9
TC207242 UP|O22107 (O22107) Squalene synthase , complete 26 7.7
TC205877 similar to UP|Q9FJJ6 (Q9FJJ6) Arabidopsis thaliana geno... 26 7.7
>AW132833
Length = 385
Score = 68.9 bits (167), Expect = 8e-13
Identities = 32/61 (52%), Positives = 44/61 (71%)
Frame = +2
Query: 44 NSLKVKEKDNMKHHCDEKREVYEYMIIQPKEKGKSNSGKGEHITSRPLQAAHDEYEEEAT 103
N L++ E+ MK CDEKR++Y+YM+ + +E G S GKGE + + LQ AHDEY+EEAT
Sbjct: 116 NELRIVEE--MKRQCDEKRDLYDYMVTRYREGGXSKGGKGETFSLQQLQTAHDEYDEEAT 289
Query: 104 L 104
L
Sbjct: 290 L 292
>BM954263
Length = 454
Score = 68.2 bits (165), Expect = 1e-12
Identities = 32/51 (62%), Positives = 38/51 (73%)
Frame = +3
Query: 47 KVKEKDNMKHHCDEKREVYEYMIIQPKEKGKSNSGKGEHITSRPLQAAHDE 97
+++ ++MK CDEKR VYEYMI Q KEKGKS SGKGE T + LQAAH E
Sbjct: 300 ELRTVEDMKRQCDEKRNVYEYMIAQQKEKGKSKSGKGESFTLQQLQAAHAE 452
>TC217405 similar to UP|Q9FLL9 (Q9FLL9) Gb|AAB80666.1, partial (20%)
Length = 1652
Score = 53.5 bits (127), Expect = 3e-08
Identities = 22/37 (59%), Positives = 29/37 (77%)
Frame = +3
Query: 109 CPHIAKLGKGRRIKPLKHSNFNERNCRDKFRQDPCRF 145
C HI +L +G RI+P +S+FN RNCRDKFR++P RF
Sbjct: 306 CFHITELSRGGRIRPFIYSSFNPRNCRDKFREEPRRF 416
>BU081020
Length = 421
Score = 31.2 bits (69), Expect = 0.18
Identities = 24/90 (26%), Positives = 41/90 (44%), Gaps = 2/90 (2%)
Frame = -2
Query: 18 KNIKKNGKFG--PIRVPRGVNNQIFEDSNSLKVKEKDNMKHHCDEKREVYEYMIIQPKEK 75
KN +K K G +R N+ + N+L VK+ D++ H YE ++
Sbjct: 309 KNHRKGTKHGFDGMRYDSASNSSDHYEENNLVVKQTDSLLHG-------YEKCNLK---- 163
Query: 76 GKSNSGKGEHITSRPLQAAHDEYEEEATLA 105
N +GE++ S+PL H + A+L+
Sbjct: 162 ---NVHRGEYVKSKPLNLVHKSIQSTASLS 82
>TC221250 weakly similar to GB|AAP37739.1|30725434|BT008380 At5g05190
{Arabidopsis thaliana;} , partial (12%)
Length = 1059
Score = 30.8 bits (68), Expect = 0.24
Identities = 20/87 (22%), Positives = 36/87 (40%)
Frame = +3
Query: 37 NQIFEDSNSLKVKEKDNMKHHCDEKREVYEYMIIQPKEKGKSNSGKGEHITSRPLQAAHD 96
N++ + SN V D++ + D V GKS S +G+ +++ PL HD
Sbjct: 315 NEVIDGSNPHSVSHADHISYSDDYGHSV-----------GKSYSSEGDPVSAAPLHPLHD 461
Query: 97 EYEEEATLAKQGCPHIAKLGKGRRIKP 123
++ T++ I + K P
Sbjct: 462 SAYDKQTVSSGTLEPITEKDKNASRSP 542
>BE331525
Length = 397
Score = 30.4 bits (67), Expect = 0.31
Identities = 10/16 (62%), Positives = 14/16 (87%)
Frame = +3
Query: 54 MKHHCDEKREVYEYMI 69
MK CDEKR++Y+YM+
Sbjct: 336 MKRQCDEKRDLYDYMV 383
>TC233625 similar to UP|Q85DB9 (Q85DB9) NADH dehydrogenase subunit 4, partial
(5%)
Length = 890
Score = 30.4 bits (67), Expect = 0.31
Identities = 25/92 (27%), Positives = 41/92 (44%), Gaps = 6/92 (6%)
Frame = +3
Query: 42 DSNSLKVKEK------DNMKHHCDEKREVYEYMIIQPKEKGKSNSGKGEHITSRPLQAAH 95
DS+ L V+E D+ H E+ E Y+ + + + +SG + + +A
Sbjct: 153 DSSHLSVEELNEADTVDDDSSHLLEQNEGYKCQGKEESCEARESSGSEKEVEENNEEALQ 332
Query: 96 DEYEEEATLAKQGCPHIAKLGKGRRIKPLKHS 127
E E L+KQ IAK KGR+ ++S
Sbjct: 333 SEAELVKNLSKQRRCAIAKAHKGRKTVAARNS 428
>TC214473 similar to UP|Q39449 (Q39449) Specific tissue protein 1, partial
(19%)
Length = 946
Score = 26.9 bits (58), Expect = 3.4
Identities = 21/89 (23%), Positives = 40/89 (44%), Gaps = 13/89 (14%)
Frame = +3
Query: 31 VPRGVNNQI-FEDSNSLKVKEK--DNMKHHCDEKR----------EVYEYMIIQPKEKGK 77
+P G+ + F+ N+ K +E+ K+HC+E E + + K
Sbjct: 150 MPEGLQGLVSFQSENNPKTQEQLGKGSKNHCEESLVTNTQVNSDFEPIPSVTKYDDLEFK 329
Query: 78 SNSGKGEHITSRPLQAAHDEYEEEATLAK 106
S+ K + SRP HD++E ++++ K
Sbjct: 330 SSIIKNDDFESRPSVTKHDDFELKSSVTK 416
>TC229283 homologue to UP|P93078 (P93078) Ap19 protein, complete
Length = 988
Score = 26.9 bits (58), Expect = 3.4
Identities = 12/24 (50%), Positives = 15/24 (62%)
Frame = -3
Query: 114 KLGKGRRIKPLKHSNFNERNCRDK 137
K+GK +RI P NFN R C+ K
Sbjct: 815 KMGKRKRILPRNLFNFNLRTCKVK 744
>AW756419
Length = 172
Score = 26.6 bits (57), Expect = 4.5
Identities = 12/39 (30%), Positives = 24/39 (60%)
Frame = -1
Query: 51 KDNMKHHCDEKREVYEYMIIQPKEKGKSNSGKGEHITSR 89
+ ++ H+ +K+ + + I+ KEK K+ S + +HIT R
Sbjct: 130 EQSLSHYF*QKKFIRDQDILLQKEKNKNTSLQQKHITLR 14
>TC214785 similar to UP|Q6V8T3 (Q6V8T3) Chlorophyll a/b-binding protein type
I (Fragment), partial (33%)
Length = 733
Score = 26.6 bits (57), Expect = 4.5
Identities = 13/40 (32%), Positives = 23/40 (57%)
Frame = -3
Query: 49 KEKDNMKHHCDEKREVYEYMIIQPKEKGKSNSGKGEHITS 88
+ ++ +KH+C K+E+ ++ KEKGK KG+ S
Sbjct: 422 RTENMLKHYCT-KQEI*DWFTG*KKEKGKRKEEKGDRFES 306
>CD412998
Length = 465
Score = 26.2 bits (56), Expect = 5.9
Identities = 10/29 (34%), Positives = 15/29 (51%)
Frame = +1
Query: 114 KLGKGRRIKPLKHSNFNERNCRDKFRQDP 142
+L +GRR P H + ++ C DK P
Sbjct: 292 ELARGRRAGPPPHKHIKQKLCSDKCHNGP 378
>AW703778
Length = 311
Score = 26.2 bits (56), Expect = 5.9
Identities = 11/42 (26%), Positives = 21/42 (49%)
Frame = +3
Query: 68 MIIQPKEKGKSNSGKGEHITSRPLQAAHDEYEEEATLAKQGC 109
++I+P+++ + G+ H R L AA + E+ Q C
Sbjct: 9 LVIEPQQQAEGRGGQVAHGLGRRLTAAPQQAPEQLVPCAQNC 134
>TC207242 UP|O22107 (O22107) Squalene synthase , complete
Length = 1671
Score = 25.8 bits (55), Expect = 7.7
Identities = 9/14 (64%), Positives = 13/14 (92%)
Frame = +2
Query: 14 SKVDKNIKKNGKFG 27
SK D++++KNGKFG
Sbjct: 164 SKFDRSVEKNGKFG 205
>TC205877 similar to UP|Q9FJJ6 (Q9FJJ6) Arabidopsis thaliana genomic DNA,
chromosome 5, TAC clone:K19B1 (AT5g62460/K19B1_7),
partial (60%)
Length = 1321
Score = 25.8 bits (55), Expect = 7.7
Identities = 12/34 (35%), Positives = 19/34 (55%)
Frame = +2
Query: 43 SNSLKVKEKDNMKHHCDEKREVYEYMIIQPKEKG 76
S SLK + ++H CDEK ++ + QP + G
Sbjct: 458 SGSLKYAHRKCVQHWCDEKGDITCEICHQPYQPG 559
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.315 0.133 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,474,104
Number of Sequences: 63676
Number of extensions: 76749
Number of successful extensions: 312
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 312
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 312
length of query: 145
length of database: 12,639,632
effective HSP length: 88
effective length of query: 57
effective length of database: 7,036,144
effective search space: 401060208
effective search space used: 401060208
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 54 (25.4 bits)
Medicago: description of AC146817.2


