

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146794.6 + phase: 0
(1018 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
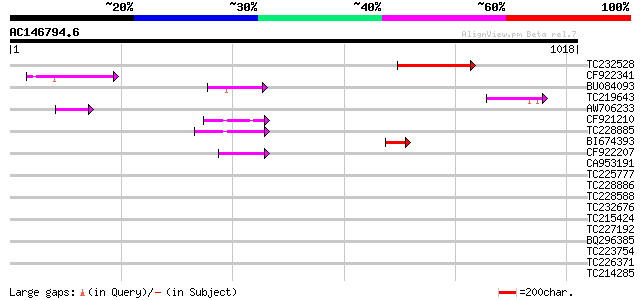
Score E
Sequences producing significant alignments: (bits) Value
TC232528 weakly similar to UP|Q6WAY5 (Q6WAY5) Gag/pol polyprotei... 152 7e-37
CF922341 98 2e-20
BU084093 66 9e-11
TC219643 weakly similar to UP|Q6WAY7 (Q6WAY7) Gag/pol polyprotei... 65 1e-10
AW706233 55 2e-07
CF921210 54 3e-07
TC228885 54 3e-07
BI674393 52 1e-06
CF922207 46 7e-05
CA953191 42 0.001
TC225777 weakly similar to UP|Q868B4 (Q868B4) C. elegans GRL-25 ... 41 0.002
TC228886 40 0.004
TC228588 similar to UP|Q6NLV4 (Q6NLV4) At5g13480, partial (29%) 40 0.007
TC232676 similar to UP|Q6K709 (Q6K709) DNA-binding protein-like,... 39 0.009
TC215424 similar to UP|Q9MA04 (Q9MA04) F20B17.16, partial (26%) 39 0.012
TC227192 similar to UP|Q6L8U3 (Q6L8U3) Auxin response factor 1, ... 37 0.034
BQ296385 37 0.044
TC223754 similar to UP|Q86EQ4 (Q86EQ4) Clone ZZD1536 mRNA sequen... 37 0.044
TC226371 similar to UP|Q9M0H8 (Q9M0H8) Predicted proline-rich pr... 36 0.075
TC214285 similar to UP|Q8RU73 (Q8RU73) Ribose-5-phosphate isomer... 36 0.098
>TC232528 weakly similar to UP|Q6WAY5 (Q6WAY5) Gag/pol polyprotein
(Fragment), partial (3%)
Length = 449
Score = 152 bits (384), Expect = 7e-37
Identities = 72/139 (51%), Positives = 104/139 (74%)
Frame = +1
Query: 697 SKISILSLLLSSAAHRDTLLKVLEQAYVDHEVTVDHFGSIVGNITACSNLWFSEDELPEA 756
+++S+L LL+SS HR L+KVL +A+V +++V+ FG +V NITA + L F+E+E+P
Sbjct: 31 ARVSLLELLMSSEPHRALLVKVLNEAHVAQDISVEGFGGLVNNITANNYLAFAEEEIPAE 210
Query: 757 GKHHNLALHISVNCKSDMLSNVLVDTGSSLNVMPKSTLDQLSYRGTPLRRSTFLVKAFDG 816
G+ HN ALH+SV C +++ VL+D G SLNVMPKSTLD+L + + L+ S+ +V+AFDG
Sbjct: 211 GRGHNKALHVSVKCMDHIVAKVLIDNGYSLNVMPKSTLDKLPFNASHLKPSSMVVRAFDG 390
Query: 817 TRKSVLGEIDLPITIGPET 835
TR+ V GEIDLP+ IGP T
Sbjct: 391 TRREVRGEIDLPVQIGPHT 447
>CF922341
Length = 675
Score = 98.2 bits (243), Expect = 2e-20
Identities = 59/175 (33%), Positives = 90/175 (50%), Gaps = 10/175 (5%)
Frame = +1
Query: 30 IPATNVVSINTTLPQTTAAVTEPLVHAIPQSVNIN--THHGNIPVIKTMEER----MEEL 83
IP N+ T L T A V IP + +H + ++ + + M E+
Sbjct: 130 IPHHNLADFETCL---TYATEGQAVGGIPLRNTLEGPQYHPQLHLLHSTTSKNPHVMAEM 300
Query: 84 AK--ELRHEIKANQGNADSF--KTQDLCLVPKVDVPKKFKIPDFDRYNGLTCPQNHIIKY 139
K L ++A +G D ++L LVP + P KFK+ DFD+Y G TCP+NH+ Y
Sbjct: 301 GKLDHLEEGLRAIEGGEDYAFANLEELFLVPNIITPPKFKVLDFDKYKGTTCPKNHLKMY 480
Query: 140 VRKMGNYKDNDSLMIHCFQDSLMEDATEWYTSLSKNDIHTFDELAAAFKSHYGFN 194
+KMG Y ++ L+IH FQ+SL A WYT+L + +H++ +L AF Y +N
Sbjct: 481 CQKMGAYAKDEELLIHSFQESLTGVAVTWYTNLEPSRVHSWKDLMVAFVRQYQYN 645
>BU084093
Length = 421
Score = 65.9 bits (159), Expect = 9e-11
Identities = 40/115 (34%), Positives = 53/115 (45%), Gaps = 7/115 (6%)
Frame = -1
Query: 356 PGQPQVPVNAIAQQM---KQQLPVQQQQQHQQARPT----FPPIPMLYAELLPTLLHRGH 408
P QP+ P A Q+ PV + P F P+PM Y +LLP+L+
Sbjct: 346 PVQPKAPAQREAPQVPTPNTTRPVGNSNTTRNFPPRPFQEFTPLPMTYEDLLPSLIANHL 167
Query: 409 CTTRQGKPPPDPLPPRFRSDLKCDFHQGALGHDVEGCYALKYIVKKLIDQGKLTF 463
G+ P P + + C +H G GH VE C ALKY V+ L+D G LTF
Sbjct: 166 AVVTPGRVLQPPFPKWYDPNATCKYHGGVPGHSVEKCLALKYKVQHLMDAGWLTF 2
>TC219643 weakly similar to UP|Q6WAY7 (Q6WAY7) Gag/pol polyprotein
(Fragment), partial (8%)
Length = 1320
Score = 65.5 bits (158), Expect = 1e-10
Identities = 48/125 (38%), Positives = 55/125 (43%), Gaps = 15/125 (12%)
Frame = +1
Query: 856 PWIHDAGAVTSTLHQKLKFAKSGKLVTIHGEEAYLVSQLSSFSCIEAGSAE-GTAFQGLT 914
PWIH G V STLHQKLKF G LV + GEE LVS SS +EA TAFQ
Sbjct: 1 PWIHSVGVVPSTLHQKLKFVVEGHLVIVSGEEDILVSCPSSMPYVEAAEESLETAFQSFE 180
Query: 915 VEGTEPKRDGTAMASLKD-----AQRAVQEGQAAGWG---------RLIQLRENKHKEGL 960
V L D A+ + G G G LI + N+ K GL
Sbjct: 181 VVSISSVDSLFGQPCLSDAAVMMARVMLGNGYEPGMGLGKDNGGITSLINTQGNRGKYGL 360
Query: 961 GFSPT 965
G+ PT
Sbjct: 361 GYKPT 375
>AW706233
Length = 376
Score = 54.7 bits (130), Expect = 2e-07
Identities = 27/71 (38%), Positives = 40/71 (56%), Gaps = 2/71 (2%)
Frame = -3
Query: 82 ELAKELRHEIKANQGNADSF--KTQDLCLVPKVDVPKKFKIPDFDRYNGLTCPQNHIIKY 139
E L+ KA +G D ++L LV + P KFK+ +FD+Y G TCP+NH+ Y
Sbjct: 347 EKLDHLKERFKAIEGGQDYAFANLEELFLVXNIISPPKFKVLNFDKYKGTTCPKNHLKMY 168
Query: 140 VRKMGNYKDND 150
+KMG Y ++
Sbjct: 167 CQKMGAYAKDE 135
>CF921210
Length = 790
Score = 53.9 bits (128), Expect = 3e-07
Identities = 41/118 (34%), Positives = 55/118 (45%)
Frame = +2
Query: 349 YHQYPLPPGQPQVPVNAIAQQMKQQLPVQQQQQHQQARPTFPPIPMLYAELLPTLLHRGH 408
YH+ LP Q + P A + Q++ ++ + F PIP+ YA+LL LL
Sbjct: 236 YHKVCLPHFQ*RTPPLAQTKNTNQEVNFAARKPVE-----FTPIPVSYADLLSYLLDNSM 400
Query: 409 CTTRQGKPPPDPLPPRFRSDLKCDFHQGALGHDVEGCYALKYIVKKLIDQGKLTFENN 466
K PL + S+ C GAL H +E C ALK V+ LID G L FE N
Sbjct: 401 VAITLAKVHQPPLF*GYDSNATCG---GALRHSIEHCRALKRKVQGLIDAGWLKFEEN 565
>TC228885
Length = 901
Score = 53.9 bits (128), Expect = 3e-07
Identities = 43/134 (32%), Positives = 58/134 (43%)
Frame = -3
Query: 333 SYPYAPYSQHPFFPPFYHQYPLPPGQPQVPVNAIAQQMKQQLPVQQQQQHQQARPTFPPI 392
S P+ P H P H+ LP Q + P A + Q++ ++ + F PI
Sbjct: 767 SLPHHPVI-HKGHP*INHKVCLPHFQ*RTPPLAQTKNTNQEMNFAARKPVE-----FTPI 606
Query: 393 PMLYAELLPTLLHRGHCTTRQGKPPPDPLPPRFRSDLKCDFHQGALGHDVEGCYALKYIV 452
P+ YA+LLP LL K P + S+ C H A G +E ALK V
Sbjct: 605 PVSYADLLPYLLDNSMVAITLAKVHQPPFLREYDSNAMCACHGEAPGRSIEHYRALKRKV 426
Query: 453 KKLIDQGKLTFENN 466
+ LID G L FE N
Sbjct: 425 QGLIDAGWLKFEEN 384
>BI674393
Length = 152
Score = 52.0 bits (123), Expect = 1e-06
Identities = 22/44 (50%), Positives = 37/44 (84%)
Frame = +1
Query: 676 EILRLIKRSDYKIVDQLLQTPSKISILSLLLSSAAHRDTLLKVL 719
E LR+I++S++K+++QL +TP+++S+L LL+SS HR L+KVL
Sbjct: 16 EFLRIIQQSEFKVIEQLNKTPARVSLLELLMSSEPHRALLVKVL 147
>CF922207
Length = 616
Score = 46.2 bits (108), Expect = 7e-05
Identities = 31/95 (32%), Positives = 40/95 (41%), Gaps = 3/95 (3%)
Frame = -1
Query: 375 PVQQQQQ---HQQARPTFPPIPMLYAELLPTLLHRGHCTTRQGKPPPDPLPPRFRSDLKC 431
P + QQ+ Q+A P L+P L K P P + S+ C
Sbjct: 613 PKKHQQKGILQQKACRVHPKFRCHMLNLIPYPLDNSMVAITPTKVPQPPFFREYDSNATC 434
Query: 432 DFHQGALGHDVEGCYALKYIVKKLIDQGKLTFENN 466
+H GA GH +E C K+ V LID G L FE N
Sbjct: 433 AYHGGAPGHSIEHCMTPKHKV*SLIDTG*LKFEEN 329
>CA953191
Length = 422
Score = 42.0 bits (97), Expect = 0.001
Identities = 28/58 (48%), Positives = 32/58 (54%), Gaps = 1/58 (1%)
Frame = -3
Query: 845 INASYSCLLGRPW-IHDAGAVTSTLHQKLKFAKSGKLVTIHGEEAYLVSQLSSFSCIE 901
I +Y+ L GRPW IH V STLH K K GKLV I +E LV + SS IE
Sbjct: 420 ITPTYNGLQGRPWRIHCVKLVPSTLH*K*KIVIDGKLVIIFVKEDLLVGEPSSTPYIE 247
>TC225777 weakly similar to UP|Q868B4 (Q868B4) C. elegans GRL-25 protein
(Corresponding sequence ZK643.8), partial (7%)
Length = 762
Score = 41.2 bits (95), Expect = 0.002
Identities = 28/93 (30%), Positives = 37/93 (39%), Gaps = 7/93 (7%)
Frame = +3
Query: 350 HQYPLPPGQPQVPVNAIAQQMKQQLPVQQQQQHQQARPTFPPIPMLYAELLPTL----LH 405
HQ LPP QP V Q + Q P QQ Q A PP P ++ + P H
Sbjct: 132 HQVELPPPQPPVQQQQQPPQQQLQQPPQQPPQQHLAPQQLPPQPPIFQQTYPPFHLPQFH 311
Query: 406 RGHCT---TRQGKPPPDPLPPRFRSDLKCDFHQ 435
+ +Q + PP P P F+ + HQ
Sbjct: 312 HHQPSLQFQQQQQQPPQPPPQLFQHQAQPPSHQ 410
>TC228886
Length = 748
Score = 40.4 bits (93), Expect = 0.004
Identities = 19/42 (45%), Positives = 24/42 (56%)
Frame = +3
Query: 425 FRSDLKCDFHQGALGHDVEGCYALKYIVKKLIDQGKLTFENN 466
+ S+ C +H GA GH +E C K+ V LID G L FE N
Sbjct: 153 YDSNATCAYHGGASGHSIEHCMTPKHKV*SLIDTGWLKFEEN 278
>TC228588 similar to UP|Q6NLV4 (Q6NLV4) At5g13480, partial (29%)
Length = 1178
Score = 39.7 bits (91), Expect = 0.007
Identities = 42/141 (29%), Positives = 53/141 (36%), Gaps = 4/141 (2%)
Frame = +1
Query: 286 AESSVNASKRYGNGHHKKKETEVGMVSAGAGQSMATVAPINAAQMP---PSYPYAPYSQH 342
AE S A + GN + T G + G ++ T+ + A MP PS Q
Sbjct: 415 AEQSPVAGRTGGNFTIAEGPTTPGPFAPGLTRNEGTIPGVGVA-MPLSIPSLDMPQGEQK 591
Query: 343 PFFPPFYHQYPLPPGQPQVPVNAIAQQMKQQLPVQQ-QQQHQQARPTFPPIPMLYAELLP 401
P PLPPG +NA QQ QQ P Q Q QHQ +PM +
Sbjct: 592 QPHPVSMGAPPLPPGPHPSLINANQQQPYQQNPQQMPQHQHQALPQQMGSLPM--PPNMQ 765
Query: 402 TLLHRGHCTTRQGKPPPDPLP 422
L H H PP +P
Sbjct: 766 QLQHPSHSPFPNMPRPPPQMP 828
>TC232676 similar to UP|Q6K709 (Q6K709) DNA-binding protein-like, partial
(57%)
Length = 926
Score = 39.3 bits (90), Expect = 0.009
Identities = 30/84 (35%), Positives = 38/84 (44%), Gaps = 3/84 (3%)
Frame = +1
Query: 349 YHQYPLPPGQPQVPVNAIAQQMKQQLPVQQQQQHQQARPTFPPIPMLYAELLPTLLHRGH 408
Y + PL Q + + + QQ +QQ QQQQQ QQ++ + M L P LLH G
Sbjct: 451 YERLPLEDDQGEEEMQ-VQQQQQQQQQQQQQQQQQQSQGLGEQVSMPMYNLPPNLLHNGQ 627
Query: 409 CTTRQ---GKPPPDPLPPRFRSDL 429
G PP PP F S L
Sbjct: 628 NMPHDVFWGAPPRP--PPSF*SPL 693
>TC215424 similar to UP|Q9MA04 (Q9MA04) F20B17.16, partial (26%)
Length = 1030
Score = 38.9 bits (89), Expect = 0.012
Identities = 31/99 (31%), Positives = 36/99 (36%), Gaps = 4/99 (4%)
Frame = +3
Query: 329 QMPPSYPYAPYSQH--PFFPPFYHQ--YPLPPGQPQVPVNAIAQQMKQQLPVQQQQQHQQ 384
Q+PP+ Y QH P+ PP H YP PP P P A Q
Sbjct: 48 QIPPNSNYHQNQQHHVPYAPPNPHHPHYPYPPPPPPPPPEA----------------SYQ 179
Query: 385 ARPTFPPIPMLYAELLPTLLHRGHCTTRQGKPPPDPLPP 423
P PP PM Y + + Q PPP PL P
Sbjct: 180 PPPPPPPAPMYYPS--------NNQYSNQPPPPPPPLSP 272
>TC227192 similar to UP|Q6L8U3 (Q6L8U3) Auxin response factor 1, partial
(14%)
Length = 875
Score = 37.4 bits (85), Expect = 0.034
Identities = 27/71 (38%), Positives = 36/71 (50%), Gaps = 2/71 (2%)
Frame = +3
Query: 355 PPGQPQVPVNAIAQQMKQQL--PVQQQQQHQQARPTFPPIPMLYAELLPTLLHRGHCTTR 412
PP + N QQ+ QL +QQQQQHQQ R P+L ++LL + T +
Sbjct: 105 PPATQMLQQNPSEQQLHLQLLQKLQQQQQHQQ-RLLSTSTPLLLSQLL-----QQQNTHQ 266
Query: 413 QGKPPPDPLPP 423
Q + P PLPP
Sbjct: 267 QNQQLPQPLPP 299
>BQ296385
Length = 399
Score = 37.0 bits (84), Expect = 0.044
Identities = 33/124 (26%), Positives = 42/124 (33%), Gaps = 10/124 (8%)
Frame = +2
Query: 314 GAGQSMATVAPINAAQM-------PPSYPYAPYSQHPFFPPFYHQYPLPPGQPQVPVNAI 366
G G + P N ++ PP P P QHP PP H + P P ++
Sbjct: 5 GGGTKRRSFRPHNPGRLRLHRQIRPPRSPQIP--QHPILPPQIHSHSGPKPTKTNPGPSV 178
Query: 367 AQQMKQQLPVQQQQQHQQARPTFPPIPMLYAELLPTLLHRGHCTTRQGKPPPD---PLPP 423
Q Q P+ RP PP P+ R PPP PLPP
Sbjct: 179 GLQ-AQPSPLPPHPHRGHRRPLLPPFPL-----------------RPHGPPPQLRGPLPP 304
Query: 424 RFRS 427
R+
Sbjct: 305 PRRA 316
>TC223754 similar to UP|Q86EQ4 (Q86EQ4) Clone ZZD1536 mRNA sequence, partial
(22%)
Length = 742
Score = 37.0 bits (84), Expect = 0.044
Identities = 32/100 (32%), Positives = 40/100 (40%), Gaps = 7/100 (7%)
Frame = -2
Query: 335 PYAPYSQHPFFPPFYHQYPLPPGQPQVPVNAIAQQMKQQLPVQQQQQHQQARPTFPPIPM 394
P P + FPP PLPP P +P + M Q L H++ P PP PM
Sbjct: 444 PGFPQLKQVLFPP---PPPLPPPPPNLPPS-----MPQSLAK*PTSPHRKQPPPLPPPPM 289
Query: 395 L---YAELLPTLLHRGHCTTRQGKPPPD----PLPPRFRS 427
L A+ L + H PPP P PPR +S
Sbjct: 288 LPFPVAQSLAMCPNPPHLKQDPPPPPPPMPTLPPPPRTQS 169
>TC226371 similar to UP|Q9M0H8 (Q9M0H8) Predicted proline-rich protein,
partial (10%)
Length = 1099
Score = 36.2 bits (82), Expect = 0.075
Identities = 26/77 (33%), Positives = 31/77 (39%), Gaps = 6/77 (7%)
Frame = +1
Query: 321 TVAPINAAQMPPSYPYAPYSQHPFFPPFYHQYPLP----PGQPQVPVNAIAQQMKQQLPV 376
T P +A P Y P Q P P P P P + Q +QQ P
Sbjct: 28 TPMPNSALPQHPQNQYLPSDQQYRTPQLVAPQPTPSQVTPSPPVQQFSHYQQPQQQQQPP 207
Query: 377 QQQQQH--QQARPTFPP 391
QQQQQ QQ +P+ PP
Sbjct: 208 QQQQQQWSQQVQPSQPP 258
>TC214285 similar to UP|Q8RU73 (Q8RU73) Ribose-5-phosphate isomerase
precursor , partial (81%)
Length = 951
Score = 35.8 bits (81), Expect = 0.098
Identities = 34/112 (30%), Positives = 47/112 (41%), Gaps = 10/112 (8%)
Frame = +2
Query: 322 VAPINAAQMPPSYPYAPYSQHPFFPPFYHQYPLPPGQPQ-VPVNAIAQQMKQQLPVQQQQ 380
++PI+ + P +APYS HP P H PLPP P+ P Q+ +++ +Q
Sbjct: 17 LSPISLFRTPQRL-HAPYSTHP*LP*TAHSTPLPPRHPRHHPHPGRPQETRRRQGRGVRQ 193
Query: 381 QHQQARPTFPPIPMLY-------AELLPTLLHR--GHCTTRQGKPPPDPLPP 423
+ RP L + L PTL HR H T G P PP
Sbjct: 194 ERHGPRPRHRLHCCLRRRQAWRPSRLRPTLRHRRCPHLQTHGGAGPLPRHPP 349
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.317 0.134 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 44,292,503
Number of Sequences: 63676
Number of extensions: 669314
Number of successful extensions: 5630
Number of sequences better than 10.0: 161
Number of HSP's better than 10.0 without gapping: 4941
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 5387
length of query: 1018
length of database: 12,639,632
effective HSP length: 107
effective length of query: 911
effective length of database: 5,826,300
effective search space: 5307759300
effective search space used: 5307759300
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 64 (29.3 bits)
Medicago: description of AC146794.6


