

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
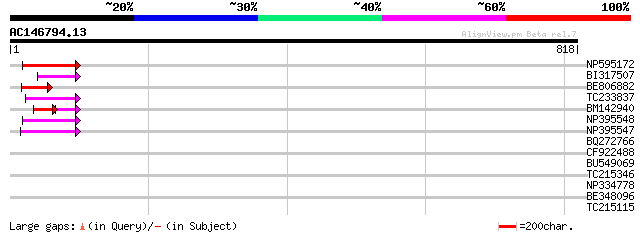
Query= AC146794.13 + phase: 0 /pseudo
(818 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
NP595172 polyprotein [Glycine max] 87 4e-17
BI317507 48 2e-05
BE806882 similar to GP|27764548|gb polyprotein {Glycine max}, pa... 47 3e-05
TC233837 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 45 1e-04
BM142940 39 2e-04
NP395548 reverse transcriptase [Glycine max] 44 3e-04
NP395547 reverse transcriptase [Glycine max] 43 5e-04
BQ272766 weakly similar to GP|28558781|gb| pol protein {Cucumis ... 41 0.002
CF922488 41 0.002
BU549069 35 0.17
TC215346 homologue to UP|Q8LP18 (Q8LP18) Cullin-like protein1, p... 34 0.23
NP334778 reverse transcriptase [Glycine max] 32 0.87
BE348096 29 7.3
TC215115 homologue to PIR|T02313|T02313 endoplasmic reticulum in... 29 9.6
>NP595172 polyprotein [Glycine max]
Length = 4659
Score = 86.7 bits (213), Expect = 4e-17
Identities = 41/83 (49%), Positives = 60/83 (71%)
Frame = +1
Query: 19 FRCYMNKIFHAYLDWFVVVFIDDILIYSRSEEEHAEHLKIMLQVLKEKKMYAKLSKCEFW 78
F+C MNKIF L FV+VF DDILIYS S ++H +HL+ +LQ LK+ +++A+LSKC F
Sbjct: 2188 FQCLMNKIFQFALRKFVLVFFDDILIYSASWKDHLKHLESVLQTLKQHQLFARLSKCSFG 2367
Query: 79 LSEVSFLGHIISDSGIVVTHQKL 101
+EV +LGH +S G+ + + K+
Sbjct: 2368 DTEVDYLGHKVSGLGVSMENTKV 2436
>BI317507
Length = 359
Score = 47.8 bits (112), Expect = 2e-05
Identities = 22/61 (36%), Positives = 36/61 (58%)
Frame = -1
Query: 41 DILIYSRSEEEHAEHLKIMLQVLKEKKMYAKLSKCEFWLSEVSFLGHIISDSGIVVTHQK 100
+ILIYS + H HL +L VLK++++ A KC F + + +LGH+IS + + K
Sbjct: 353 NILIYSPDWKSHIMHLTAVLDVLKKERLVANRKKCYFSQTTIEYLGHVISKDCVAMDSNK 174
Query: 101 L 101
+
Sbjct: 173 V 171
>BE806882 similar to GP|27764548|gb polyprotein {Glycine max}, partial (3%)
Length = 153
Score = 47.0 bits (110), Expect = 3e-05
Identities = 21/45 (46%), Positives = 31/45 (68%)
Frame = -1
Query: 18 AFRCYMNKIFHAYLDWFVVVFIDDILIYSRSEEEHAEHLKIMLQV 62
+F+C MN IF L +V+VF DDIL+YS + EH HL+++ +V
Sbjct: 135 SFQCLMNHIFQHALRKYVLVFFDDILVYSSTWHEHLCHLEVVFKV 1
>TC233837 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (6%)
Length = 402
Score = 45.1 bits (105), Expect = 1e-04
Identities = 23/79 (29%), Positives = 43/79 (54%)
Frame = +2
Query: 23 MNKIFHAYLDWFVVVFIDDILIYSRSEEEHAEHLKIMLQVLKEKKMYAKLSKCEFWLSEV 82
M +FH + + V++DD++ SR+E EH +L + L++ ++ +KC F +
Sbjct: 155 MVALFHDMMHKEIEVYVDDMIAKSRTETEHLVNLCKLFGRLQKYQLKLNPTKCTFGVKSG 334
Query: 83 SFLGHIISDSGIVVTHQKL 101
LG I+S GI + +K+
Sbjct: 335 KLLGFIVSQKGIEIDPEKV 391
>BM142940
Length = 422
Score = 38.5 bits (88), Expect(2) = 2e-04
Identities = 18/34 (52%), Positives = 24/34 (69%)
Frame = -1
Query: 35 VVVFIDDILIYSRSEEEHAEHLKIMLQVLKEKKM 68
+ F DILIYS +E +H +HLKI+LQ KE K+
Sbjct: 260 ICFFSYDILIYSINEGKHEKHLKIVLQTFKEAKL 159
Score = 25.4 bits (54), Expect(2) = 2e-04
Identities = 12/35 (34%), Positives = 18/35 (51%)
Frame = -3
Query: 67 KMYAKLSKCEFWLSEVSFLGHIISDSGIVVTHQKL 101
K+ A KC F + +L H+IS G+ +KL
Sbjct: 165 KVGANRKKCSFGCKKFEYLVHMISVKGVSADPKKL 61
>NP395548 reverse transcriptase [Glycine max]
Length = 762
Score = 43.9 bits (102), Expect = 3e-04
Identities = 27/83 (32%), Positives = 41/83 (48%)
Frame = +1
Query: 19 FRCYMNKIFHAYLDWFVVVFIDDILIYSRSEEEHAEHLKIMLQVLKEKKMYAKLSKCEFW 78
F+ M IF ++ + VF+DD ++ S E + L+++LQ E + KC F
Sbjct: 409 FQMCMLAIFADIVEKSIEVFMDDFSVFVPSLESCLKKLEMVLQRCVETNLVLNWEKCHFM 588
Query: 79 LSEVSFLGHIISDSGIVVTHQKL 101
+ E LGH IS GI V K+
Sbjct: 589 VREGIVLGHKISTRGIEVDQTKI 657
>NP395547 reverse transcriptase [Glycine max]
Length = 762
Score = 43.1 bits (100), Expect = 5e-04
Identities = 27/86 (31%), Positives = 41/86 (47%)
Frame = +1
Query: 16 SDAFRCYMNKIFHAYLDWFVVVFIDDILIYSRSEEEHAEHLKIMLQVLKEKKMYAKLSKC 75
S F+ M IF ++ + VF+DD + S +L+ +LQ ++ + KC
Sbjct: 400 STTFQRCMMAIFDDMVEKCIEVFMDDFSFFGASFGNCLANLEKVLQRCEKSNLVLNWEKC 579
Query: 76 EFWLSEVSFLGHIISDSGIVVTHQKL 101
F + E LGH IS GI V +KL
Sbjct: 580 HFMVQEGIVLGHKISKRGIEVVKEKL 657
>BQ272766 weakly similar to GP|28558781|gb| pol protein {Cucumis melo},
partial (9%)
Length = 410
Score = 40.8 bits (94), Expect = 0.002
Identities = 28/73 (38%), Positives = 36/73 (48%)
Frame = -2
Query: 662 ELSRQA*ERPWISGS**CVSKSRSCDRCWSCLEIEEVDAEVYWPVSDIRKSWNSGI*SGF 721
ELSRQ ERP I G * C+ KS D WS EI + Y + +KS GI +
Sbjct: 409 ELSRQEEERPRIRGW*SCILKSHPIDWGWSSFEILKTHTSFYRSFPNSQKS*FCGISNCI 230
Query: 722 ATTSFEFA*CVSC 734
S+ + C+SC
Sbjct: 229 TPISY*PSQCLSC 191
>CF922488
Length = 741
Score = 40.8 bits (94), Expect = 0.002
Identities = 21/67 (31%), Positives = 39/67 (57%)
Frame = +3
Query: 35 VVVFIDDILIYSRSEEEHAEHLKIMLQVLKEKKMYAKLSKCEFWLSEVSFLGHIISDSGI 94
+ V++DD+++ SR+EEEH +L+ + + L++ ++ +KC F + L I S GI
Sbjct: 321 IEVYMDDMIVKSRTEEEHLVNLRKLFRRLRKYRLRLNPAKCMFEVKSRKLLDFIDS*RGI 500
Query: 95 VVTHQKL 101
V K+
Sbjct: 501 EVDSNKV 521
>BU549069
Length = 615
Score = 34.7 bits (78), Expect = 0.17
Identities = 25/74 (33%), Positives = 36/74 (47%)
Frame = -3
Query: 682 KSRSCDRCWSCLEIEEVDAEVYWPVSDIRKSWNSGI*SGFATTSFEFA*CVSCVATSEVC 741
+S S D WS EI + +Y + KS + GI + +F + C+SCV+T V
Sbjct: 613 ESHSKDWGWSSTEIPKTHTSLYRSFPNS*KSXSCGIPNCITPITF*SSQCLSCVSTPYVY 434
Query: 742 IGSIACDSEG*CAS 755
SI+C G* S
Sbjct: 433 P*SISCGQIG*RTS 392
>TC215346 homologue to UP|Q8LP18 (Q8LP18) Cullin-like protein1, partial (13%)
Length = 1050
Score = 34.3 bits (77), Expect = 0.23
Identities = 22/85 (25%), Positives = 42/85 (48%)
Frame = +3
Query: 55 HLKIMLQVLKEKKMYAKLSKCEFWLSEVSFLGHIISDSGIVVTHQKLMQ*HNGRLRSQLQ 114
H +++L+VL +Y + + LS + G +I ++ + ++ +L+S L
Sbjct: 735 HCRLLLRVLPVNALYDNSEEMDLILSHLGIFGFVILRRDRIILYFQVFF-FPAKLKSILS 911
Query: 115 RLEVFWVW*VITTDL*KDFRSWHFH 139
L V W W +I ++*+DF FH
Sbjct: 912 -LFVLWQWKLILGEI*EDFEFEQFH 983
>NP334778 reverse transcriptase [Glycine max]
Length = 431
Score = 32.3 bits (72), Expect = 0.87
Identities = 17/68 (25%), Positives = 34/68 (50%)
Frame = +3
Query: 12 LRIFSDAFRCYMNKIFHAYLDWFVVVFIDDILIYSRSEEEHAEHLKIMLQVLKEKKMYAK 71
L+ F + M +F + + ++D+++ SR EEEH +L+ + L++ ++
Sbjct: 228 LKNFGATYHRAMVALFQDMMHKEIEAYVDEMIAKSRMEEEHLVNLQNLFGQLRKYRLRLN 407
Query: 72 LSKCEFWL 79
KC F L
Sbjct: 408 PRKCVFGL 431
>BE348096
Length = 445
Score = 29.3 bits (64), Expect = 7.3
Identities = 16/33 (48%), Positives = 20/33 (60%)
Frame = -1
Query: 205 VCIKAVEDS*MELSYA*FGVGCCGFCVENLEAL 237
+C +DS +LSYA* VG G C+ LEAL
Sbjct: 169 LCSSIGQDSLKDLSYA*SRVGSHGVCLHILEAL 71
>TC215115 homologue to PIR|T02313|T02313 endoplasmic reticulum insertion
protein F13P17.9 - Arabidopsis thaliana {Arabidopsis
thaliana;} , complete
Length = 1786
Score = 28.9 bits (63), Expect = 9.6
Identities = 26/82 (31%), Positives = 39/82 (46%), Gaps = 3/82 (3%)
Frame = +3
Query: 44 IYSRSEEEHAEHLKIMLQVLK---EKKMYAKLSKCEFWLSEVSFLGHIISDSGIVVTHQK 100
++SRS +LK L + K E++ Y S C F L VSFL + T Q+
Sbjct: 168 LFSRS------YLKFRLLIGKCHLERRSYIL*SLCSFSLFAVSFL-------CMEYTQQQ 308
Query: 101 LMQ*HNGRLRSQLQRLEVFWVW 122
++ G + S LQ +E+ W W
Sbjct: 309 VLIHSIGCVLSLLQTVELLWSW 374
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.372 0.167 0.652
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 36,460,546
Number of Sequences: 63676
Number of extensions: 487421
Number of successful extensions: 6294
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 2461
Number of HSP's successfully gapped in prelim test: 187
Number of HSP's that attempted gapping in prelim test: 3748
Number of HSP's gapped (non-prelim): 2822
length of query: 818
length of database: 12,639,632
effective HSP length: 105
effective length of query: 713
effective length of database: 5,953,652
effective search space: 4244953876
effective search space used: 4244953876
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.9 bits)
S2: 63 (28.9 bits)
Medicago: description of AC146794.13


