

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146792.8 - phase: 0
(266 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
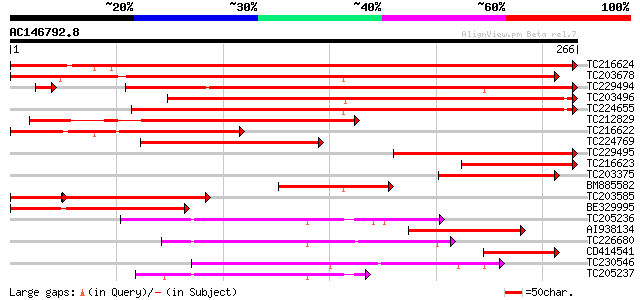
Sequences producing significant alignments: (bits) Value
TC216624 similar to UP|Q6YZ10 (Q6YZ10) 27k vesicle-associated me... 479 e-136
TC203678 similar to UP|Q6YZ10 (Q6YZ10) 27k vesicle-associated me... 360 e-100
TC229494 335 1e-92
TC203496 similar to UP|Q6YZ10 (Q6YZ10) 27k vesicle-associated me... 312 1e-85
TC224655 similar to UP|Q9LVV9 (Q9LVV9) Membrane associated prote... 310 6e-85
TC212829 similar to UP|Q8LAZ8 (Q8LAZ8) Membrane associated prote... 204 3e-53
TC216622 185 2e-47
TC224769 similar to UP|Q9LVV9 (Q9LVV9) Membrane associated prote... 153 7e-38
TC229495 similar to UP|Q6YZ10 (Q6YZ10) 27k vesicle-associated me... 132 2e-31
TC216623 similar to UP|Q6YZ10 (Q6YZ10) 27k vesicle-associated me... 99 2e-21
TC203375 homologue to UP|Q9XQB0 (Q9XQB0) Carbonic anhydrase, par... 98 3e-21
BM885582 88 3e-18
TC203585 82 6e-18
BE329995 similar to GP|1800147|gb|A membrane associated protein ... 70 8e-13
TC205236 similar to UP|Q9SC83 (Q9SC83) VAP27, partial (82%) 62 3e-10
AI938134 60 1e-09
TC226680 weakly similar to UP|SCS2_YEAST (P40075) SCS2 protein, ... 59 2e-09
CD414541 55 4e-08
TC230546 similar to GB|AAM62506.1|21553413|AY085274 VAMP (vesicl... 54 9e-08
TC205237 similar to UP|Q9SC83 (Q9SC83) VAP27, partial (58%) 49 2e-06
>TC216624 similar to UP|Q6YZ10 (Q6YZ10) 27k vesicle-associated membrane
protein-associated protein-like, partial (81%)
Length = 1108
Score = 479 bits (1234), Expect = e-136
Identities = 246/270 (91%), Positives = 254/270 (93%), Gaps = 4/270 (1%)
Frame = +3
Query: 1 MAVAEPKLHSEPKVWNFFKLPFRNSNTSSTNTTSSSVN-NNLHHHHH---HNPNSNTPLE 56
MAVA+PK HSEPKVWNFFKLPFR+S T TNTT SS N ++LHHHHH HNPN+N PLE
Sbjct: 153 MAVADPKPHSEPKVWNFFKLPFRHSAT--TNTTPSSANLHHLHHHHHNHHHNPNNNLPLE 326
Query: 57 GSTSHTSNSVSSVARSLLPTRRRLKLDPSNKLYFPYEPGKQVRSAIRIKNTSKSNVAFKF 116
GSTS+TSNSVSSVARSLLPTRRRLKLDPSNKLYFPYEPGKQVRSAIRIKNTSKSNVAFKF
Sbjct: 327 GSTSNTSNSVSSVARSLLPTRRRLKLDPSNKLYFPYEPGKQVRSAIRIKNTSKSNVAFKF 506
Query: 117 QTTAPKSCFMRPPGAILAPGESIIATVFKFVEQPENNEKPEKSGLKFKIMSLKVKGSIDY 176
QTTAPKSCFMRPPGAILAPGESIIATVFKFVEQPENNEKPEK+GLKFKIMSLKVKG IDY
Sbjct: 507 QTTAPKSCFMRPPGAILAPGESIIATVFKFVEQPENNEKPEKTGLKFKIMSLKVKGLIDY 686
Query: 177 VPELFDEQKDQVAVEQILRVVFLDPERPSPILEKLKRQLADADAALEERKKPAEDAGPKI 236
VPELFDEQKDQVAVEQILRVVFLDPERP P LEKL RQLADADAALE RKKPAEDAGPKI
Sbjct: 687 VPELFDEQKDQVAVEQILRVVFLDPERPCPALEKLNRQLADADAALEARKKPAEDAGPKI 866
Query: 237 IGEGLVIDEWKERRERYLAKQQGDVVVDSV 266
IGEGLVIDEWKERRERYLAKQQG+VVVDSV
Sbjct: 867 IGEGLVIDEWKERRERYLAKQQGEVVVDSV 956
>TC203678 similar to UP|Q6YZ10 (Q6YZ10) 27k vesicle-associated membrane
protein-associated protein-like, partial (84%)
Length = 1297
Score = 360 bits (924), Expect = e-100
Identities = 183/261 (70%), Positives = 220/261 (84%), Gaps = 3/261 (1%)
Frame = +2
Query: 1 MAVAEPKLHSEPKVWNFFKLPF-RNSNTSSTNTTSSSVNNNLHHHHHHNPNSNTPLEGST 59
M V K S+ K W+F ++PF + ++T S++TTS S +N+H + +N S ++ S+
Sbjct: 236 MEVETEKSGSDGKGWSFCRMPFWQTTHTPSSSTTSMSYMHNVHQQNQNNAQS---VDRSS 406
Query: 60 SHTSNSVSSVARSLLPTRRRLKLDPSNKLYFPYEPGKQVRSAIRIKNTSKSNVAFKFQTT 119
H+S +VSSVA+SLLPT+RRL+LDPSNKLYFPYEPGKQVRSAI IKNT KS+VAFKFQTT
Sbjct: 407 HHSSTTVSSVAKSLLPTKRRLRLDPSNKLYFPYEPGKQVRSAIAIKNTCKSHVAFKFQTT 586
Query: 120 APKSCFMRPPGAILAPGESIIATVFKFVEQPENNEK--PEKSGLKFKIMSLKVKGSIDYV 177
APKSC+MRPPG ILAPGESIIATVFKFVE PENNEK +KS +KFKIMSLKV+G +DYV
Sbjct: 587 APKSCYMRPPGGILAPGESIIATVFKFVEPPENNEKSIDKKSKVKFKIMSLKVQGEMDYV 766
Query: 178 PELFDEQKDQVAVEQILRVVFLDPERPSPILEKLKRQLADADAALEERKKPAEDAGPKII 237
PELFDEQ+DQVA+EQILRV+FLDPE+PSP+L+KLKR LA+ADAALE RKKP E+ GP +
Sbjct: 767 PELFDEQRDQVAIEQILRVIFLDPEKPSPVLDKLKRMLAEADAALEARKKPLEEKGPCVA 946
Query: 238 GEGLVIDEWKERRERYLAKQQ 258
GEGLVIDEWKERRERYLA+QQ
Sbjct: 947 GEGLVIDEWKERRERYLAQQQ 1009
>TC229494
Length = 923
Score = 335 bits (860), Expect(2) = 1e-92
Identities = 172/213 (80%), Positives = 189/213 (87%), Gaps = 1/213 (0%)
Frame = +2
Query: 55 LEGSTSHTSNSVSSVARSLLPTRRRLKLDPSNKLYFPYEPGKQVRSAIRIKNTSKSNVAF 114
++ S SHTSNSVSS+ARSLLP RRLKL PSNKLYFP PGKQ +SA+RIKNTSKSNVAF
Sbjct: 23 VQASLSHTSNSVSSLARSLLPAPRRLKLHPSNKLYFP*-PGKQAKSAVRIKNTSKSNVAF 199
Query: 115 KFQTTAPKSCFMRPPGAILAPGESIIATVFKFVEQPENNEKPEKSGLKFKIMSLKVKGSI 174
KFQT APKSCFMRPPG ILAPGESIIATVFKF+E P NNEKP+K+G+KFKI SLKVKG I
Sbjct: 200 KFQTNAPKSCFMRPPGGILAPGESIIATVFKFIEHPLNNEKPQKTGVKFKITSLKVKGPI 379
Query: 175 DYVPELFDEQKDQVAVEQILRVVFLDPERPSPILEKLKRQLADADAA-LEERKKPAEDAG 233
DYVPE+FDE KDQV VEQIL+VVFLDPER P LEKLK QLA+ADAA +E RK+ EDAG
Sbjct: 380 DYVPEMFDELKDQVTVEQILQVVFLDPERSCPALEKLKLQLAEADAAVVEARKRAPEDAG 559
Query: 234 PKIIGEGLVIDEWKERRERYLAKQQGDVVVDSV 266
PKI+GEGLVIDEWKERRERYLAKQQG+VVVDS+
Sbjct: 560 PKILGEGLVIDEWKERRERYLAKQQGEVVVDSI 658
Score = 21.9 bits (45), Expect(2) = 1e-92
Identities = 8/10 (80%), Positives = 8/10 (80%)
Frame = +3
Query: 13 KVWNFFKLPF 22
KVWN FKL F
Sbjct: 9 KVWNLFKLRF 38
>TC203496 similar to UP|Q6YZ10 (Q6YZ10) 27k vesicle-associated membrane
protein-associated protein-like, partial (82%)
Length = 871
Score = 312 bits (799), Expect = 1e-85
Identities = 158/194 (81%), Positives = 173/194 (88%), Gaps = 2/194 (1%)
Frame = +3
Query: 75 PTRRRLKLDPSNKLYFPYEPGKQVRSAIRIKNTSKSNVAFKFQTTAPKSCFMRPPGAILA 134
P RRRL LDP KLYFPYEPGKQVRSAI IKNT KS+VAFKFQTTAPKSC+MRPPG ILA
Sbjct: 3 PXRRRLXLDPPXKLYFPYEPGKQVRSAIAIKNTCKSHVAFKFQTTAPKSCYMRPPGGILA 182
Query: 135 PGESIIATVFKFVEQPENNEKP--EKSGLKFKIMSLKVKGSIDYVPELFDEQKDQVAVEQ 192
PGESIIATVFKFVE PENNEKP +KS +KFKIMSLKVKG +DYVPELFDEQ+D VA EQ
Sbjct: 183 PGESIIATVFKFVEPPENNEKPIDQKSRVKFKIMSLKVKGEMDYVPELFDEQRDHVADEQ 362
Query: 193 ILRVVFLDPERPSPILEKLKRQLADADAALEERKKPAEDAGPKIIGEGLVIDEWKERRER 252
ILRV+FLDPERPSP L+KLKRQLA+A+AALE RKKP E+ GP++ GEGLVIDEWKERRER
Sbjct: 363 ILRVIFLDPERPSPALDKLKRQLAEAEAALEARKKPPEETGPRVAGEGLVIDEWKERRER 542
Query: 253 YLAKQQGDVVVDSV 266
YLA+QQ + VVDSV
Sbjct: 543 YLARQQVE-VVDSV 581
>TC224655 similar to UP|Q9LVV9 (Q9LVV9) Membrane associated protein, partial
(67%)
Length = 1188
Score = 310 bits (793), Expect = 6e-85
Identities = 157/211 (74%), Positives = 183/211 (86%), Gaps = 2/211 (0%)
Frame = +1
Query: 58 STSHTSNSVSSVARSLLPTRRRLKLDPSNKLYFPYEPGKQVRSAIRIKNTSKSNVAFKFQ 117
+T ++ +VSSVARSLL RR+L+LDPS+ LYFPYEPGKQVRSA+R+KNTSKS+VAFKFQ
Sbjct: 217 ATQRSTKTVSSVARSLLRPRRKLRLDPSSYLYFPYEPGKQVRSAVRLKNTSKSHVAFKFQ 396
Query: 118 TTAPKSCFMRPPGAILAPGESIIATVFKFVEQPENNEK--PEKSGLKFKIMSLKVKGSID 175
TTAPKSC+MRPPG LAPGES+IATVFKFVEQPENNEK +K +KFKIMSLKVK +D
Sbjct: 397 TTAPKSCYMRPPGGTLAPGESLIATVFKFVEQPENNEKLSDQKHKVKFKIMSLKVKEGVD 576
Query: 176 YVPELFDEQKDQVAVEQILRVVFLDPERPSPILEKLKRQLADADAALEERKKPAEDAGPK 235
YVPELFDEQKD V +E+ILRVVF+DPER S +LEKLKRQLA+ADAA+E RKKP +AGP+
Sbjct: 577 YVPELFDEQKDLVTMERILRVVFIDPERQSSVLEKLKRQLAEADAAVEARKKPPVEAGPR 756
Query: 236 IIGEGLVIDEWKERRERYLAKQQGDVVVDSV 266
+ EGLVIDEWKERRE+YLA+QQ VDSV
Sbjct: 757 VAAEGLVIDEWKERREKYLARQQVQ-AVDSV 846
>TC212829 similar to UP|Q8LAZ8 (Q8LAZ8) Membrane associated protein-like
protein, partial (38%)
Length = 438
Score = 204 bits (520), Expect = 3e-53
Identities = 109/155 (70%), Positives = 117/155 (75%)
Frame = +1
Query: 10 SEPKVWNFFKLPFRNSNTSSTNTTSSSVNNNLHHHHHHNPNSNTPLEGSTSHTSNSVSSV 69
SE KVWN FKLPFR+S ++ H+HN S TSNSVSS
Sbjct: 49 SEGKVWNLFKLPFRHSTSTP---------------HNHNQVS----------TSNSVSSF 153
Query: 70 ARSLLPTRRRLKLDPSNKLYFPYEPGKQVRSAIRIKNTSKSNVAFKFQTTAPKSCFMRPP 129
A+SLLP+ RRLKLDPSNKLYFPYEPGKQ RSAIRIKNTSKSNVAFKFQT APKSCFMRPP
Sbjct: 154 AKSLLPSPRRLKLDPSNKLYFPYEPGKQGRSAIRIKNTSKSNVAFKFQTNAPKSCFMRPP 333
Query: 130 GAILAPGESIIATVFKFVEQPENNEKPEKSGLKFK 164
G ILAPGESIIATVFKF+E P NNEKPEK+GLKFK
Sbjct: 334 GGILAPGESIIATVFKFIEHPLNNEKPEKTGLKFK 438
>TC216622
Length = 455
Score = 185 bits (469), Expect = 2e-47
Identities = 96/119 (80%), Positives = 102/119 (85%), Gaps = 9/119 (7%)
Frame = +1
Query: 1 MAVAEPKLHSEPKVWNFFKLPFRNSNTSSTNTTSSSVN---------NNLHHHHHHNPNS 51
MAVA+PK HSEPKVWNFFKLPFR+S ++TNTT SS N ++ HHHHHHNPN
Sbjct: 106 MAVADPKPHSEPKVWNFFKLPFRHS--AATNTTPSSANLHHLHHHHNHHHHHHHHHNPN- 276
Query: 52 NTPLEGSTSHTSNSVSSVARSLLPTRRRLKLDPSNKLYFPYEPGKQVRSAIRIKNTSKS 110
N PLEGSTSHTSNSVSSVARSLLPTRRRLKLDPSNKLYFPYEPGKQVRSAIRIKNTSKS
Sbjct: 277 NLPLEGSTSHTSNSVSSVARSLLPTRRRLKLDPSNKLYFPYEPGKQVRSAIRIKNTSKS 453
>TC224769 similar to UP|Q9LVV9 (Q9LVV9) Membrane associated protein, partial
(34%)
Length = 481
Score = 153 bits (387), Expect = 7e-38
Identities = 74/86 (86%), Positives = 82/86 (95%)
Frame = +3
Query: 62 TSNSVSSVARSLLPTRRRLKLDPSNKLYFPYEPGKQVRSAIRIKNTSKSNVAFKFQTTAP 121
+S +VSSVARSLLP RRRL+LDPSN LYFPYEPGKQVRSA+R+KNTSKS+VAFKFQTTAP
Sbjct: 222 SSKTVSSVARSLLPPRRRLRLDPSNYLYFPYEPGKQVRSAVRLKNTSKSHVAFKFQTTAP 401
Query: 122 KSCFMRPPGAILAPGESIIATVFKFV 147
KSC+MRPPG ILAPGES+IATVFKFV
Sbjct: 402 KSCYMRPPGGILAPGESLIATVFKFV 479
>TC229495 similar to UP|Q6YZ10 (Q6YZ10) 27k vesicle-associated membrane
protein-associated protein-like, partial (33%)
Length = 564
Score = 132 bits (332), Expect = 2e-31
Identities = 65/86 (75%), Positives = 74/86 (85%)
Frame = +3
Query: 181 FDEQKDQVAVEQILRVVFLDPERPSPILEKLKRQLADADAALEERKKPAEDAGPKIIGEG 240
F +KDQV VEQIL+VVFLDPER P LEKLK QL +ADAA+E RK+ EDAGPKI+GEG
Sbjct: 3 FGTRKDQVTVEQILQVVFLDPERNCPALEKLKLQLVEADAAVEARKRAPEDAGPKILGEG 182
Query: 241 LVIDEWKERRERYLAKQQGDVVVDSV 266
LV+ EWKERRERYLAKQQG+VVVDS+
Sbjct: 183 LVVHEWKERRERYLAKQQGEVVVDSI 260
>TC216623 similar to UP|Q6YZ10 (Q6YZ10) 27k vesicle-associated membrane
protein-associated protein-like, partial (21%)
Length = 441
Score = 99.4 bits (246), Expect = 2e-21
Identities = 49/54 (90%), Positives = 51/54 (93%)
Frame = +1
Query: 213 RQLADADAALEERKKPAEDAGPKIIGEGLVIDEWKERRERYLAKQQGDVVVDSV 266
RQLADADAALE RKKPAEDAGPKIIGEGLVIDEWKERRERYLAKQQG+V +SV
Sbjct: 1 RQLADADAALEARKKPAEDAGPKIIGEGLVIDEWKERRERYLAKQQGEVARNSV 162
>TC203375 homologue to UP|Q9XQB0 (Q9XQB0) Carbonic anhydrase, partial (57%)
Length = 1427
Score = 98.2 bits (243), Expect = 3e-21
Identities = 45/57 (78%), Positives = 52/57 (90%)
Frame = -3
Query: 202 ERPSPILEKLKRQLADADAALEERKKPAEDAGPKIIGEGLVIDEWKERRERYLAKQQ 258
ERPSP+L+KLKR LA+ADAALE RKKP E+ GP++ GEGLVIDEWKERRERYLA+QQ
Sbjct: 336 ERPSPVLDKLKRMLAEADAALEARKKPPEEKGPRVAGEGLVIDEWKERRERYLAQQQ 166
>BM885582
Length = 423
Score = 88.2 bits (217), Expect = 3e-18
Identities = 45/56 (80%), Positives = 49/56 (87%), Gaps = 2/56 (3%)
Frame = +2
Query: 127 RPPGAILAPGESIIATVFKFVEQPENNEK--PEKSGLKFKIMSLKVKGSIDYVPEL 180
RPPG ILAPGESIIATVFKFVE PENNEK +KS +KFKIMSLKV+G +DYVPEL
Sbjct: 2 RPPGGILAPGESIIATVFKFVEPPENNEKSIDKKSKVKFKIMSLKVQGEMDYVPEL 169
>TC203585
Length = 363
Score = 81.6 bits (200), Expect(2) = 6e-18
Identities = 39/70 (55%), Positives = 50/70 (70%)
Frame = +1
Query: 25 SNTSSTNTTSSSVNNNLHHHHHHNPNSNTPLEGSTSHTSNSVSSVARSLLPTRRRLKLDP 84
S++SS+ TT+ +++HH HH L+ ST S +VSSVA+SLLPTRRRL+LDP
Sbjct: 154 SSSSSSTTTTIPTTSSMHHSVHHQSQILQSLDRSTHQPSTTVSSVAKSLLPTRRRLRLDP 333
Query: 85 SNKLYFPYEP 94
NKLYFPYEP
Sbjct: 334 PNKLYFPYEP 363
Score = 26.2 bits (56), Expect(2) = 6e-18
Identities = 11/27 (40%), Positives = 17/27 (62%)
Frame = +3
Query: 1 MAVAEPKLHSEPKVWNFFKLPFRNSNT 27
MAV K+ S+ KVW+F +P +N+
Sbjct: 69 MAVESDKVGSDGKVWSFCSMPLWANNS 149
>BE329995 similar to GP|1800147|gb|A membrane associated protein {Arabidopsis
thaliana}, partial (7%)
Length = 458
Score = 70.5 bits (171), Expect = 8e-13
Identities = 39/84 (46%), Positives = 51/84 (60%)
Frame = +2
Query: 1 MAVAEPKLHSEPKVWNFFKLPFRNSNTSSTNTTSSSVNNNLHHHHHHNPNSNTPLEGSTS 60
MAV K S K W+F +PF T+ ++TSS+ ++HH HH L+ ST
Sbjct: 209 MAVESDKTGSVGKGWSFCSMPFWQ--TTHASSTSSTTTFSMHHSVHHQSQILQSLDRSTH 382
Query: 61 HTSNSVSSVARSLLPTRRRLKLDP 84
S +VSSVA+SLLPTRRRL+LDP
Sbjct: 383 QPSTTVSSVAKSLLPTRRRLRLDP 454
>TC205236 similar to UP|Q9SC83 (Q9SC83) VAP27, partial (82%)
Length = 765
Score = 61.6 bits (148), Expect = 3e-10
Identities = 48/157 (30%), Positives = 83/157 (52%), Gaps = 5/157 (3%)
Frame = +2
Query: 53 TPLEGSTSHTSNSVSSVARSLLPTRRRLKLDPSNKLYFPYEPGKQVRSAIRIKNTSKSNV 112
TP+ S++S S + S + T L ++P +L FP+E KQ+ ++++ N + S V
Sbjct: 71 TPVISHNSNSSGSRLDRSISNMSTGELLNIEPL-ELKFPFELKKQISCSLQLSNKTDSYV 247
Query: 113 AFKFQTTAPKSCFMRPPGAILAPGES--IIATVFKFVEQPENNEKPEKSGLKFKIMSLK- 169
AFK +TT PK +RP I+ P + +I T+ E P + + + KF + S+K
Sbjct: 248 AFKVKTTNPKKYCVRPNTGIVTPRSTCDVIVTMQAQKEAPADMQCKD----KFLLQSVKT 415
Query: 170 VKGSI--DYVPELFDEQKDQVAVEQILRVVFLDPERP 204
V G+ D ++F+++ V E LRV+++ P P
Sbjct: 416 VDGTSPKDITADMFNKEAGHVVEECKLRVLYVSPP*P 526
>AI938134
Length = 172
Score = 59.7 bits (143), Expect = 1e-09
Identities = 33/55 (60%), Positives = 38/55 (69%)
Frame = -1
Query: 188 VAVEQILRVVFLDPERPSPILEKLKRQLADADAALEERKKPAEDAGPKIIGEGLV 242
VAV QIL+VVF+D E P P E L RQL A ALE RKKP+++ G IIG GLV
Sbjct: 166 VAVGQILQVVFIDTEGPCPA*ENLTRQLDYASVALEARKKPSQEHGSYIIGGGLV 2
>TC226680 weakly similar to UP|SCS2_YEAST (P40075) SCS2 protein, partial
(16%)
Length = 1112
Score = 59.3 bits (142), Expect = 2e-09
Identities = 41/141 (29%), Positives = 76/141 (53%), Gaps = 3/141 (2%)
Frame = +3
Query: 72 SLLPTRRRLKLDPSNKLYFPYEPGKQVRSAIRIKNTSKSNVAFKFQTTAPKSCFMRPPGA 131
S + + L + P +L FP+E KQ+ ++++ N + + VAFK +TT PK +RP
Sbjct: 120 SAMSSGELLHIQPQ-ELQFPFELRKQISCSLQLSNKTDNYVAFKVKTTNPKKYCVRPNTG 296
Query: 132 ILAPGES--IIATVFKFVEQPENNEKPEKSGLKFKIMSLKVKGSIDYVPELFDEQKDQVA 189
++ P + +I T+ E P + + +K L+ +++ + D PE+F+++
Sbjct: 297 VVMPRSTCDVIVTMQAQKEAPPDMQCKDKFLLQ-SVVASPGATTKDITPEMFNKESGHDV 473
Query: 190 VEQILRVVFL-DPERPSPILE 209
E LRVV++ P+ PSP+ E
Sbjct: 474 EECKLRVVYVAPPQPPSPVRE 536
>CD414541
Length = 563
Score = 54.7 bits (130), Expect = 4e-08
Identities = 24/36 (66%), Positives = 30/36 (82%)
Frame = -3
Query: 223 EERKKPAEDAGPKIIGEGLVIDEWKERRERYLAKQQ 258
+++KK + GP + GEGLVIDEWKERRERYLA+QQ
Sbjct: 327 KKKKKARGEKGPCVAGEGLVIDEWKERRERYLAQQQ 220
>TC230546 similar to GB|AAM62506.1|21553413|AY085274 VAMP (vesicle-associated
membrane protein)-associated protein-like {Arabidopsis
thaliana;} , partial (60%)
Length = 1019
Score = 53.5 bits (127), Expect = 9e-08
Identities = 43/154 (27%), Positives = 77/154 (49%), Gaps = 7/154 (4%)
Frame = +3
Query: 86 NKLYFPYEPGKQVRSAIRIKNTSKSNVAFKFQTTAPKSCFMRPPGAILAPGESIIATVFK 145
++L F +E KQ +++ N +++ VAFK +TT+PK F+RP ++ P + I V
Sbjct: 246 DELRFQFELEKQTYCDLKVLNNTENYVAFKVKTTSPKKYFVRPNTGVVHPWDLCIIRVTL 425
Query: 146 FVEQ--PENNEKPEKSGLKFKIMSLKVKGSIDYVPELFDEQKDQVAVEQILRVVFLDPER 203
+Q P + + +K L+ I++ D P+ F++ ++ + LRVV++ P
Sbjct: 426 QAQQEYPPDMQCKDKFLLQSTIVNPNTDVD-DLPPDTFNKDGEKSIEDMKLRVVYISPMS 602
Query: 204 PSPILE----KLKRQLADADAA-LEERKKPAEDA 232
P E K Q DA+++ +R K DA
Sbjct: 603 PQGSTEDDTVKNSTQKLDANSSETVQRLKEERDA 704
>TC205237 similar to UP|Q9SC83 (Q9SC83) VAP27, partial (58%)
Length = 696
Score = 48.9 bits (115), Expect = 2e-06
Identities = 37/115 (32%), Positives = 60/115 (52%), Gaps = 5/115 (4%)
Frame = +3
Query: 60 SHTSNSVSSVAR---SLLPTRRRLKLDPSNKLYFPYEPGKQVRSAIRIKNTSKSNVAFKF 116
SH SNS S S + T L ++P +L FP+E KQ+ ++++ N + S VAFK
Sbjct: 114 SHNSNSSGSRLDR*ISNMSTGELLNIEPL-ELKFPFELKKQISCSLQLSNKTDSYVAFKV 290
Query: 117 QTTAPKSCFMRPPGAILAPGES--IIATVFKFVEQPENNEKPEKSGLKFKIMSLK 169
+TT PK +RP ++ P + +I T+ E P + + + KF + S+K
Sbjct: 291 KTTNPKKYCVRPNTGVVTPRSTCDVIVTMQAQKEAPADMQCKD----KFLLQSVK 443
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.313 0.130 0.369
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,871,398
Number of Sequences: 63676
Number of extensions: 187016
Number of successful extensions: 4657
Number of sequences better than 10.0: 304
Number of HSP's better than 10.0 without gapping: 3004
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3949
length of query: 266
length of database: 12,639,632
effective HSP length: 96
effective length of query: 170
effective length of database: 6,526,736
effective search space: 1109545120
effective search space used: 1109545120
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 58 (26.9 bits)
Medicago: description of AC146792.8


