

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
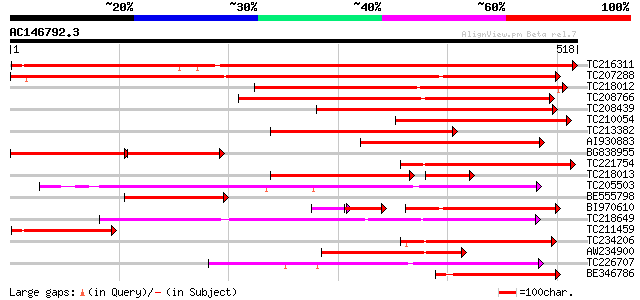
Query= AC146792.3 + phase: 0
(518 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC216311 UP|Q7XA52 (Q7XA52) Monosaccharide transporter, complete 832 0.0
TC207288 UP|Q7XA51 (Q7XA51) Monosaccharide transporter, complete 528 e-150
TC218012 similar to UP|Q6VEF2 (Q6VEF2) Monosaccharide transporte... 350 7e-97
TC208766 similar to UP|STA_RICCO (Q10710) Sugar carrier protein ... 340 1e-93
TC208439 weakly similar to UP|O04078 (O04078) Monosaccharid tran... 323 1e-88
TC210054 homologue to UP|Q7XA52 (Q7XA52) Monosaccharide transpor... 290 8e-79
TC213382 homologue to UP|Q7XA52 (Q7XA52) Monosaccharide transpor... 270 1e-72
AI930883 weakly similar to GP|4138724|emb hexose transporter {Vi... 223 2e-58
BG838955 similar to GP|5734440|emb hexose transporter {Lycopersi... 135 6e-58
TC221754 similar to UP|Q06312 (Q06312) H(+)/MONOSACCHARIDE cotra... 207 1e-53
TC218013 similar to UP|Q6VEF2 (Q6VEF2) Monosaccharide transporte... 154 1e-51
TC205503 UP|Q7XA50 (Q7XA50) Sorbitol-like transporter, complete 198 5e-51
BE555798 homologue to GP|1353516|gb|A sugar transporter {Medicag... 184 6e-47
BI970610 153 4e-44
TC218649 similar to UP|Q39416 (Q39416) Integral membrane protein... 172 4e-43
TC211459 homologue to UP|Q7XA52 (Q7XA52) Monosaccharide transpor... 165 5e-41
TC234206 weakly similar to GB|AAO64833.1|29028908|BT005898 At1g3... 152 3e-37
AW234900 weakly similar to GP|16945177|em STP5 protein {Arabidop... 147 9e-36
TC226707 similar to UP|Q7XA50 (Q7XA50) Sorbitol-like transporter... 132 3e-31
BE346786 similar to GP|5734440|emb hexose transporter {Lycopersi... 127 2e-29
>TC216311 UP|Q7XA52 (Q7XA52) Monosaccharide transporter, complete
Length = 1925
Score = 832 bits (2150), Expect = 0.0
Identities = 424/523 (81%), Positives = 461/523 (88%), Gaps = 6/523 (1%)
Frame = +1
Query: 2 AGGGIPIGGGNKEYPGNLTPFVTITCIVAAMGGLIFGYDIGISGGVTSMDPFLKKFFPAV 61
A GGI GGG KEYPG+LTPFVT+TCIVAAMGGLIFGYDIGISGGVTSMDPFL KFFP+V
Sbjct: 103 AVGGISNGGG-KEYPGSLTPFVTVTCIVAAMGGLIFGYDIGISGGVTSMDPFLLKFFPSV 279
Query: 62 YRKKNKDKSTNQYCQYDSQTLTMFTSSLYLAALLSSLVASTITRRFGRKLSMLFGGLLFL 121
+RKKN DK+ NQYCQYDSQTLTMFTSSLYLAALLSSLVAST+TRRFGRKLSMLFGGLLFL
Sbjct: 280 FRKKNSDKTVNQYCQYDSQTLTMFTSSLYLAALLSSLVASTVTRRFGRKLSMLFGGLLFL 459
Query: 122 VGALINGFANHVWMLIVGRILLGFGIGFANQA---VPLYLSEMAPYKYRGAL--NIGFQL 176
VGALINGFA HVWMLIVGRILLGFGIGFANQ +P+ + + G L N Q
Sbjct: 460 VGALINGFAQHVWMLIVGRILLGFGIGFANQVCATLPI*NGFIQI*RSIGTLAFNCQSQF 639
Query: 177 SITIGILVANVLNYFFAKIKGGWGWRLSLG-GAMVPALIITIGSLVLPDTPNSMIERGDR 235
I G V +L + K WGW++ + GAMVPALIIT+GSLVLPDTPNSMIERGDR
Sbjct: 640 GIPRGQCVELLL---WLKSMVAWGWKIEVWEGAMVPALIITVGSLVLPDTPNSMIERGDR 810
Query: 236 DGAKAQLKRIRGIEDVDEEFNDLVAASEASMQVENPWRNLLQRKYRPQLTMAVLIPFFQQ 295
+ AKAQL+RIRGI++VDEEFNDLVAASE+S QVE+PWRNLLQRKYRP LTMAVLIPFFQQ
Sbjct: 811 EKAKAQLQRIRGIDNVDEEFNDLVAASESSSQVEHPWRNLLQRKYRPHLTMAVLIPFFQQ 990
Query: 296 FTGINVIMFYAPVLFNSIGFKDDASLMSAVITGVVNVVATCVSIYGVDKWGRRALFLEGG 355
TGINVIMFYAPVLF+SIGFKDDA+LMSAVITGVVNVVATCVSIYGVDKWGRRALFLEGG
Sbjct: 991 LTGINVIMFYAPVLFSSIGFKDDAALMSAVITGVVNVVATCVSIYGVDKWGRRALFLEGG 1170
Query: 356 AQMLICQVAVAAAIGAKFGTSGNPGNLPEWYAIVVVLFICIYVAGFAWSWGPLGWLVPSE 415
QMLICQ VAAAIGAKFGT GNPG+LP+WYAIVVVLFICIYV+ FAWSWGPLGWLVPSE
Sbjct: 1171VQMLICQAVVAAAIGAKFGTDGNPGDLPKWYAIVVVLFICIYVSAFAWSWGPLGWLVPSE 1350
Query: 416 IFPLEIRSAAQSVNVSVNMLFTFLVAQVFLIMLCHMKFGLFLFFAFFVLVMSIYVFFLLP 475
IFPLEIRSAAQS+NVSVNMLFTFL+AQVFL MLCHMKFGLFLFFAFFVL+M+ +V+F LP
Sbjct: 1351IFPLEIRSAAQSINVSVNMLFTFLIAQVFLTMLCHMKFGLFLFFAFFVLIMTFFVYFFLP 1530
Query: 476 ETKGIPIEEMDRVWKSHPFWSRFVEHGDHGNGVEMGKGAPKNV 518
ETKGIPIEEM +VW++HPFWSRFVEH D+GNGVEMGKGA K V
Sbjct: 1531ETKGIPIEEMGQVWQAHPFWSRFVEHDDYGNGVEMGKGAIKEV 1659
>TC207288 UP|Q7XA51 (Q7XA51) Monosaccharide transporter, complete
Length = 1813
Score = 528 bits (1361), Expect = e-150
Identities = 270/507 (53%), Positives = 355/507 (69%), Gaps = 4/507 (0%)
Frame = +1
Query: 1 MAGGGIPIGGGNKE---YPGNLTPFVTITCIVAAMGGLIFGYDIGISGGVTSMDPFLKKF 57
MAGGG+ GG K Y T + TC+V A+GG +FGYD+G+SGGV SMD FLK+F
Sbjct: 28 MAGGGLTNGGPGKRAHLYEHKFTAYFAFTCVVGALGGSLFGYDLGVSGGVPSMDDFLKEF 207
Query: 58 FPAVYRKKNKDKSTNQYCQYDSQTLTMFTSSLYLAALLSSLVASTITRRFGRKLSMLFGG 117
FP VYR+K YC+YD Q LT+FTSSLY +AL+ + AS +TR+ GRK S++ G
Sbjct: 208 FPKVYRRKQMHLHETDYCKYDDQVLTLFTSSLYFSALVMTFFASFLTRKKGRKASIIVGA 387
Query: 118 LLFLVGALINGFANHVWMLIVGRILLGFGIGFANQAVPLYLSEMAPYKYRGALNIGFQLS 177
L FL GA++N A ++ MLI+GR+LLG GIGF NQAVPLYLSEMAP K RGA+N FQ +
Sbjct: 388 LSFLAGAILNAAAKNIAMLIIGRVLLGGGIGFGNQAVPLYLSEMAPAKNRGAVNQLFQFT 567
Query: 178 ITIGILVANVLNYFFAKIKGGWGWRLSLGGAMVPALIITIGSLVLPDTPNSMIERGDRDG 237
GIL+AN++NYF KI +GWR+SLG A +PA + +G + +TPNS++E+G D
Sbjct: 568 TCAGILIANLVNYFTEKIH-PYGWRISLGLAGLPAFAMLVGGICCAETPNSLVEQGRLDK 744
Query: 238 AKAQLKRIRGIEDVDEEFNDLVAASEASMQVENPWRNLLQRKYRPQLTMAVL-IPFFQQF 296
AK L+RIRG E+V+ EF DL ASE + V++P+R LL+RKYRPQL + L IP FQQ
Sbjct: 745 AKQVLQRIRGTENVEAEFEDLKEASEEAQAVKSPFRTLLKRKYRPQLIIGALGIPAFQQL 924
Query: 297 TGINVIMFYAPVLFNSIGFKDDASLMSAVITGVVNVVATCVSIYGVDKWGRRALFLEGGA 356
TG N I+FYAPV+F S+GF +ASL S+ IT +VAT +S++ VDK+GRR FLE G
Sbjct: 925 TGNNSILFYAPVIFQSLGFGANASLFSSFITNGALLVATVISMFLVDKYGRRKFFLEAGF 1104
Query: 357 QMLICQVAVAAAIGAKFGTSGNPGNLPEWYAIVVVLFICIYVAGFAWSWGPLGWLVPSEI 416
+M+ C + A + FG G + +VV+ ++V + SWGPLGWLVPSE+
Sbjct: 1105EMICCMIITGAVLAVNFGHGKEIGKGVSAFLVVVIF---LFVLAYGRSWGPLGWLVPSEL 1275
Query: 417 FPLEIRSAAQSVNVSVNMLFTFLVAQVFLIMLCHMKFGLFLFFAFFVLVMSIYVFFLLPE 476
FPLEIRS+AQS+ V VNM+FT LVAQ+FL+ LCH+KFG+FL FA ++ MS +VFFLLPE
Sbjct: 1276FPLEIRSSAQSIVVCVNMIFTALVAQLFLMSLCHLKFGIFLLFASLIIFMSFFVFFLLPE 1455
Query: 477 TKGIPIEEMDRVWKSHPFWSRFVEHGD 503
TK +PIEE+ ++++H FW RFV D
Sbjct: 1456TKKVPIEEIYLLFENHWFWRRFVTDQD 1536
>TC218012 similar to UP|Q6VEF2 (Q6VEF2) Monosaccharide transporter 4, partial
(54%)
Length = 1142
Score = 350 bits (899), Expect = 7e-97
Identities = 172/295 (58%), Positives = 223/295 (75%), Gaps = 9/295 (3%)
Frame = +3
Query: 224 DTPNSMIERGDRDGAKAQLKRIRGIEDVDEEFNDLVAASEASMQVENPWRNLLQRKYRPQ 283
DTPNS+IERG + K LK+IRG ++++ EF +LV AS + +V++P+RNLL+R+ RPQ
Sbjct: 3 DTPNSLIERGRLEEGKTVLKKIRGTDNIELEFQELVEASRVAKEVKHPFRNLLKRRNRPQ 182
Query: 284 LTMAVLIPFFQQFTGINVIMFYAPVLFNSIGFKDDASLMSAVITGVVNVVATCVSIYGVD 343
L +++ + FQQFTGIN IMFYAPVLFN++GFK+DASL SAVITG VNV++T VSIY VD
Sbjct: 183 LVISIALQIFQQFTGINAIMFYAPVLFNTLGFKNDASLYSAVITGAVNVLSTVVSIYSVD 362
Query: 344 KWGRRALFLEGGAQMLICQVAVAAAIGAKFGTSGNPGNLPEWYAIVVVLFICIYVAGFAW 403
K GRR L LE G QM + QV +A +G K + + +L + AI+VV+ +C +V+ FAW
Sbjct: 363 KLGRRMLLLEAGVQMFLSQVVIAIILGIK--VTDHSDDLSKGIAILVVVMVCTFVSSFAW 536
Query: 404 SWGPLGWLVPSEIFPLEIRSAAQSVNVSVNMLFTFLVAQVFLIMLCHMKFGLFLFFAFFV 463
SWGPLGWL+PSE FPLE RSA QSV V VN+LFTF++AQ FL MLCH KFG+FLFF+ +V
Sbjct: 537 SWGPLGWLIPSETFPLETRSAGQSVTVCVNLLFTFVIAQAFLSMLCHFKFGIFLFFSGWV 716
Query: 464 LVMSIYVFFLLPETKGIPIEEM-DRVWKSHPFWSRFVE--------HGDHGNGVE 509
LVMS++V FLLPETK +PIEEM +RVWK H FW RF++ H +GNG +
Sbjct: 717 LVMSVFVLFLLPETKNVPIEEMTERVWKQHWFWKRFIDDAADEKVAHVSNGNGFD 881
>TC208766 similar to UP|STA_RICCO (Q10710) Sugar carrier protein A, partial
(55%)
Length = 1202
Score = 340 bits (871), Expect = 1e-93
Identities = 159/288 (55%), Positives = 216/288 (74%)
Frame = +1
Query: 210 VPALIITIGSLVLPDTPNSMIERGDRDGAKAQLKRIRGIEDVDEEFNDLVAASEASMQVE 269
VPAL++T+G + LPDTPNS+IERG + + L++IRG ++VD EF D+V ASE + ++
Sbjct: 4 VPALLMTVGGIFLPDTPNSLIERGLAEKGRKLLEKIRGTKEVDAEFQDMVDASELAKSIK 183
Query: 270 NPWRNLLQRKYRPQLTMAVLIPFFQQFTGINVIMFYAPVLFNSIGFKDDASLMSAVITGV 329
+P+RN+L+R+YRP+L MA+ +P FQ TGIN I+FYAPVLF S+GF DASL+S+ +TG
Sbjct: 184 HPFRNILERRYRPELVMAIFMPTFQILTGINSILFYAPVLFQSMGFGGDASLISSALTGG 363
Query: 330 VNVVATCVSIYGVDKWGRRALFLEGGAQMLICQVAVAAAIGAKFGTSGNPGNLPEWYAIV 389
V +T +SI VD+ GRR L + GG QM+ CQ+ VA +G KFG L + ++I+
Sbjct: 364 VLASSTFISIATVDRLGRRVLLVSGGLQMITCQIIVAIILGVKFGAD---QELSKGFSIL 534
Query: 390 VVLFICIYVAGFAWSWGPLGWLVPSEIFPLEIRSAAQSVNVSVNMLFTFLVAQVFLIMLC 449
VV+ IC++V F WSWGPLGW VPSEIFPLEIRSA Q + V+VN+LFTF++AQ FL +LC
Sbjct: 535 VVVVICLFVVAFGWSWGPLGWTVPSEIFPLEIRSAGQGITVAVNLLFTFIIAQAFLALLC 714
Query: 450 HMKFGLFLFFAFFVLVMSIYVFFLLPETKGIPIEEMDRVWKSHPFWSR 497
KFG+FLFFA ++ +M+I+V+ LPETKGIPIEEM +W+ H FW R
Sbjct: 715 SFKFGIFLFFAGWITIMTIFVYLFLPETKGIPIEEMSFMWRRHWFWKR 858
>TC208439 weakly similar to UP|O04078 (O04078) Monosaccharid transport
protein, partial (43%)
Length = 936
Score = 323 bits (828), Expect = 1e-88
Identities = 150/220 (68%), Positives = 177/220 (80%)
Frame = +1
Query: 281 RPQLTMAVLIPFFQQFTGINVIMFYAPVLFNSIGFKDDASLMSAVITGVVNVVATCVSIY 340
RPQL A+ IPFFQQFTG+NVI FYAP+LF +IGF ASLMSAVI G V+T VSI
Sbjct: 1 RPQLVFAICIPFFQQFTGLNVITFYAPILFRTIGFGSGASLMSAVIIGSFKPVSTLVSIL 180
Query: 341 GVDKWGRRALFLEGGAQMLICQVAVAAAIGAKFGTSGNPGNLPEWYAIVVVLFICIYVAG 400
VDK+GRR LFLEGGAQMLICQ+ + AI FGT+GNPG LP+WYAIVVV IC+YV+G
Sbjct: 181 LVDKFGRRTLFLEGGAQMLICQIIMTIAIAVTFGTNGNPGTLPKWYAIVVVGIICVYVSG 360
Query: 401 FAWSWGPLGWLVPSEIFPLEIRSAAQSVNVSVNMLFTFLVAQVFLIMLCHMKFGLFLFFA 460
FAWSWGPLGWL+PSEIFPLEIR AAQS+ V VNM+ TF +AQ F MLCHMKFGLF+FF
Sbjct: 361 FAWSWGPLGWLIPSEIFPLEIRPAAQSITVGVNMISTFFIAQFFTSMLCHMKFGLFIFFG 540
Query: 461 FFVLVMSIYVFFLLPETKGIPIEEMDRVWKSHPFWSRFVE 500
FV++M+ +++ LLPETKGIP+EEM VW+ HP W +F+E
Sbjct: 541 CFVVIMTTFIYKLLPETKGIPLEEMSMVWQKHPIWGKFLE 660
>TC210054 homologue to UP|Q7XA52 (Q7XA52) Monosaccharide transporter, partial
(31%)
Length = 620
Score = 290 bits (743), Expect = 8e-79
Identities = 130/161 (80%), Positives = 149/161 (91%)
Frame = +2
Query: 353 EGGAQMLICQVAVAAAIGAKFGTSGNPGNLPEWYAIVVVLFICIYVAGFAWSWGPLGWLV 412
EGG QM+ICQ VAAAIGAKFG GNPG+LP+WYA+VVVLFICIYV+ FAWSWGPLGWLV
Sbjct: 2 EGGVQMVICQAVVAAAIGAKFGIDGNPGDLPKWYAVVVVLFICIYVSAFAWSWGPLGWLV 181
Query: 413 PSEIFPLEIRSAAQSVNVSVNMLFTFLVAQVFLIMLCHMKFGLFLFFAFFVLVMSIYVFF 472
PSEIFPLEIRSAAQS+NVSVNM FTFL+AQVFL MLCHMKFGLF+ FAFFVL+M+ +++F
Sbjct: 182 PSEIFPLEIRSAAQSINVSVNMFFTFLIAQVFLTMLCHMKFGLFILFAFFVLIMTFFIYF 361
Query: 473 LLPETKGIPIEEMDRVWKSHPFWSRFVEHGDHGNGVEMGKG 513
LPETKGIPIEEM++VWK+HPFWSRFVE+ D+GNGVEMG+G
Sbjct: 362 FLPETKGIPIEEMNQVWKAHPFWSRFVENDDYGNGVEMGRG 484
>TC213382 homologue to UP|Q7XA52 (Q7XA52) Monosaccharide transporter, partial
(35%)
Length = 624
Score = 270 bits (690), Expect = 1e-72
Identities = 137/172 (79%), Positives = 150/172 (86%), Gaps = 1/172 (0%)
Frame = +1
Query: 239 KAQLKRIRGIEDVDEEFNDLVAASEASMQVENPWRNLLQRKYRPQLTMAVLIPFFQQFTG 298
KAQL+R+RGI+DV+EEFNDLVAASE+S +VE+PWRNLLQRKYRP LTMAVLIPFFQQ TG
Sbjct: 1 KAQLRRVRGIDDVEEEFNDLVAASESSRKVEHPWRNLLQRKYRPHLTMAVLIPFFQQLTG 180
Query: 299 INVIMFYAPVLFNSIGFKDDASLMSAVITGVVNVVATCVSIYGVDKWGRRALFLEGGAQM 358
INVIMFYAPVLF+SIGFKDD++LMSAVITGVVNVVATCVSIYGVDKWGRRALFLEGG QM
Sbjct: 181 INVIMFYAPVLFSSIGFKDDSALMSAVITGVVNVVATCVSIYGVDKWGRRALFLEGGVQM 360
Query: 359 LICQVAVAAAIGAKFGT-SGNPGNLPEWYAIVVVLFICIYVAGFAWSWGPLG 409
+ICQ VAAAIGAKFG G+L +WYA VVVLFICIYV G LG
Sbjct: 361 VICQAVVAAAIGAKFGNLMETLGDLAKWYAXVVVLFICIYVFSIWLVMGSLG 516
>AI930883 weakly similar to GP|4138724|emb hexose transporter {Vitis
vinifera}, partial (31%)
Length = 505
Score = 223 bits (568), Expect = 2e-58
Identities = 113/168 (67%), Positives = 129/168 (76%)
Frame = +2
Query: 321 LMSAVITGVVNVVATCVSIYGVDKWGRRALFLEGGAQMLICQVAVAAAIGAKFGTSGNPG 380
LMSAVITGVVN ATCVSIY VDKW RRALFLEGG QMLICQ V AIGA GT GNP
Sbjct: 2 LMSAVITGVVNADATCVSIYSVDKWCRRALFLEGGVQMLICQAVVVTAIGA*LGTDGNPC 181
Query: 381 NLPEWYAIVVVLFICIYVAGFAWSWGPLGWLVPSEIFPLEIRSAAQSVNVSVNMLFTFLV 440
+LP+W AIVV L+I IYV+ FAW W L WLVPSEIF EIRSAAQS+ VSVNML+TFL+
Sbjct: 182 DLPQWNAIVVALYIGIYVSAFAW*WRTLCWLVPSEIFHFEIRSAAQSITVSVNMLYTFLI 361
Query: 441 AQVFLIMLCHMKFGLFLFFAFFVLVMSIYVFFLLPETKGIPIEEMDRV 488
A VFL MLCHM + LF+AFF+L+M+ +V+F +T GI IE + V
Sbjct: 362 AHVFLTMLCHMTDRMDLFYAFFLLIMTFFVYFF*LDTHGISIEHIGHV 505
>BG838955 similar to GP|5734440|emb hexose transporter {Lycopersicon
esculentum}, partial (36%)
Length = 631
Score = 135 bits (340), Expect(2) = 6e-58
Identities = 66/110 (60%), Positives = 81/110 (73%), Gaps = 1/110 (0%)
Frame = -3
Query: 1 MAGGGIPIGGGNKEYPGNLTPFVTITCIVAAMGGLIFGYDIGISGGVTSMDPFLKKFFPA 60
MAGGG GGG ++ +TP V I+CI+AA GGL+FGYD+G+SGGVTSM PFL KFFP
Sbjct: 593 MAGGGFTSGGGAGDFEAKITPIVIISCIMAATGGLMFGYDVGVSGGVTSMPPFLXKFFPT 414
Query: 61 VYRKKNKDKS-TNQYCQYDSQTLTMFTSSLYLAALLSSLVASTITRRFGR 109
VYRK ++K + YC+YD+Q L +FTSSLYLA L S+ AS TRR GR
Sbjct: 413 VYRKTVEEKGLDSNYCKYDNQGLQLFTSSLYLAGLTSTFFASYTTRRLGR 264
Score = 107 bits (268), Expect(2) = 6e-58
Identities = 54/89 (60%), Positives = 70/89 (77%)
Frame = -2
Query: 108 GRKLSMLFGGLLFLVGALINGFANHVWMLIVGRILLGFGIGFANQAVPLYLSEMAPYKYR 167
G+KL+ML G+ F+ G ++N A + MLIVGRILLG G+GFANQAVP++LSE+AP + R
Sbjct: 270 GKKLTMLIAGVFFICGVVLNAAAQDLAMLIVGRILLGCGVGFANQAVPVFLSEIAPSRIR 91
Query: 168 GALNIGFQLSITIGILVANVLNYFFAKIK 196
ALNI FQL++TIGIL AN++NY KIK
Sbjct: 90 RALNILFQLNVTIGILFANLVNYGTNKIK 4
>TC221754 similar to UP|Q06312 (Q06312) H(+)/MONOSACCHARIDE cotransporter,
partial (24%)
Length = 582
Score = 207 bits (526), Expect = 1e-53
Identities = 98/160 (61%), Positives = 118/160 (73%)
Frame = +3
Query: 358 MLICQVAVAAAIGAKFGTSGNPGNLPEWYAIVVVLFICIYVAGFAWSWGPLGWLVPSEIF 417
MLICQ+A I KFG SG G+ A +++ FIC +VA FAWSWGPLGWLVPSEI
Sbjct: 3 MLICQIATGVMIAMKFGVSGE-GSFSSGEANLILFFICAFVAAFAWSWGPLGWLVPSEIC 179
Query: 418 PLEIRSAAQSVNVSVNMLFTFLVAQVFLIMLCHMKFGLFLFFAFFVLVMSIYVFFLLPET 477
PLE+RSA Q++NV+VNMLFTF +AQVFL+MLCH+KFGLF FFA FVL+M+I++ LLPET
Sbjct: 180 PLEVRSAGQAINVAVNMLFTFAIAQVFLVMLCHLKFGLFFFFAAFVLIMTIFIAMLLPET 359
Query: 478 KGIPIEEMDRVWKSHPFWSRFVEHGDHGNGVEMGKGAPKN 517
K IPIEEM VW+SH FWS+ V H D E + KN
Sbjct: 360 KNIPIEEMHTVWRSHWFWSKIVPHADDDRKPEAAQS*IKN 479
>TC218013 similar to UP|Q6VEF2 (Q6VEF2) Monosaccharide transporter 4, partial
(33%)
Length = 568
Score = 154 bits (388), Expect(2) = 1e-51
Identities = 76/132 (57%), Positives = 100/132 (75%)
Frame = +2
Query: 239 KAQLKRIRGIEDVDEEFNDLVAASEASMQVENPWRNLLQRKYRPQLTMAVLIPFFQQFTG 298
K LKR + +++ EF +L+ AS + +V++P+RNLL+R+ RPQL ++V + FQQFTG
Sbjct: 14 KQCLKRFGALYNIELEFQELLEASRVAKEVKHPFRNLLKRRNRPQLVISVALQIFQQFTG 193
Query: 299 INVIMFYAPVLFNSIGFKDDASLMSAVITGVVNVVATCVSIYGVDKWGRRALFLEGGAQM 358
IN IMFYAPVLFN++GFK+DASL SAVITG VNV++T VSIY VDK GRR L LE G QM
Sbjct: 194 INAIMFYAPVLFNTLGFKNDASLYSAVITGAVNVLSTVVSIYSVDKVGRRILLLEAGVQM 373
Query: 359 LICQVAVAAAIG 370
+ QV +A +G
Sbjct: 374 FLSQVVIAIILG 409
Score = 68.2 bits (165), Expect(2) = 1e-51
Identities = 28/44 (63%), Positives = 36/44 (81%)
Frame = +3
Query: 381 NLPEWYAIVVVLFICIYVAGFAWSWGPLGWLVPSEIFPLEIRSA 424
+L + AI+VV+ +C +V+ FAWSWGPLGWL+PSE FPLE RSA
Sbjct: 435 DLSKGIAILVVVMVCTFVSSFAWSWGPLGWLIPSETFPLETRSA 566
>TC205503 UP|Q7XA50 (Q7XA50) Sorbitol-like transporter, complete
Length = 1928
Score = 198 bits (503), Expect = 5e-51
Identities = 132/478 (27%), Positives = 230/478 (47%), Gaps = 19/478 (3%)
Frame = +1
Query: 28 IVAAMGGLIFGYDIGISGGVTSMDPFLKKFFPAVYRKKNKDKSTNQYCQYDSQTLTMFTS 87
++A+M ++ GYDIG+ G A+Y K++ S Q + +
Sbjct: 271 MLASMTSILLGYDIGVMSGA------------AIYIKRDLKVSDEQ--------IEILLG 390
Query: 88 SLYLAALLSSLVASTITRRFGRKLSMLFGGLLFLVGALINGFANHVWMLIVGRILLGFGI 147
+ L +L+ S +A + GR+ ++ GG +FLVG+ + GF H L+ GR + G GI
Sbjct: 391 IINLYSLIGSCLAGRTSDWIGRRYTIGLGGAIFLVGSTLMGFYPHYSFLMCGRFVAGIGI 570
Query: 148 GFANQAVPLYLSEMAPYKYRGALNIGFQLSITIGILVANVLNYFFAKIKGGWGWRLSLGG 207
G+A P+Y +E++P RG L ++ I GIL+ + NY F+K+ GWR+ LG
Sbjct: 571 GYALMIAPVYTAEVSPASSRGFLTSFPEVFINGGILLGYISNYGFSKLTLKVGWRMMLGV 750
Query: 208 AMVPALIITIGSLVLPDTPNSMIERG--------------DRDGAKAQLKRIRGIEDVDE 253
+P++++T G L +P++P ++ RG ++ A+ +L I+ + E
Sbjct: 751 GAIPSVVLTEGVLAMPESPRWLVMRGRLGEARKVLNKTSDSKEEAQLRLAEIKQAAGIPE 930
Query: 254 EFNDLVAASEASMQVENPWRNLL---QRKYRPQLTMAVLIPFFQQFTGINVIMFYAPVLF 310
ND V E W+ L R + A+ I FFQQ +G++ ++ Y+P +F
Sbjct: 931 SCNDDVVQVNKQSNGEGVWKELFLYPTPAIRHIVIAALGIHFFQQASGVDAVVLYSPRIF 1110
Query: 311 NSIGFKDDA-SLMSAVITGVVNVVATCVSIYGVDKWGRRALFLEGGAQMLICQVAVAAAI 369
G +D L++ V G V V + + +D+ GRR L L M++ + +A ++
Sbjct: 1111EKAGITNDTHKLLATVAVGFVKTVFILAATFTLDRVGRRPLLLSSVGGMVLSLLTLAISL 1290
Query: 370 GAKFGTSGNPGNLPEWYAIVVVLFICIYVAGFAWSWGPLGWLVPSEIFPLEIRSAAQSVN 429
+ W + + YVA F+ GP+ W+ SEIFPL +R+ +
Sbjct: 1291----TVIDHSERKLMWAVGSSIAMVLAYVATFSIGAGPITWVYSSEIFPLRLRAQGAAAG 1458
Query: 430 VSVNMLFTFLVAQVFLIMLCHMKF-GLFLFFAFFVLVMSIYVFFLLPETKGIPIEEMD 486
V+VN + +V+ FL + + G F + V I+ + +LPET+G +E+M+
Sbjct: 1459VAVNRTTSAVVSMTFLSLTRAITIGGAFFLYCGIATVGWIFFYTVLPETRGKTLEDME 1632
>BE555798 homologue to GP|1353516|gb|A sugar transporter {Medicago
truncatula}, partial (18%)
Length = 312
Score = 184 bits (468), Expect = 6e-47
Identities = 90/95 (94%), Positives = 93/95 (97%)
Frame = +2
Query: 106 RFGRKLSMLFGGLLFLVGALINGFANHVWMLIVGRILLGFGIGFANQAVPLYLSEMAPYK 165
+FGRKLSMLF GLLFLVGALINGFA HVWMLIVGRILLGFGIGFANQ+VPLYLSEMAPYK
Sbjct: 8 KFGRKLSMLF*GLLFLVGALINGFAQHVWMLIVGRILLGFGIGFANQSVPLYLSEMAPYK 187
Query: 166 YRGALNIGFQLSITIGILVANVLNYFFAKIKGGWG 200
YRGALNIGFQLSIT+GILVANVLNYFFAKIKGGWG
Sbjct: 188 YRGALNIGFQLSITVGILVANVLNYFFAKIKGGWG 292
>BI970610
Length = 742
Score = 153 bits (387), Expect(3) = 4e-44
Identities = 74/142 (52%), Positives = 99/142 (69%)
Frame = +1
Query: 362 QVAVAAAIGAKFGTSGNPGNLPEWYAIVVVLFICIYVAGFAWSWGPLGWLVPSEIFPLEI 421
Q+ A + FG G + +VV+ ++V + SWGPLGWLVPSE+FPLEI
Sbjct: 208 QIITGAVLAVNFGHGKEIGKGVSAFLVVVIF---LFVLAYGRSWGPLGWLVPSELFPLEI 378
Query: 422 RSAAQSVNVSVNMLFTFLVAQVFLIMLCHMKFGLFLFFAFFVLVMSIYVFFLLPETKGIP 481
RS+AQS+ V VNM+FT LVAQ+FL+ LCH+KFG+FL FA ++ MS +VFFLLPETK +P
Sbjct: 379 RSSAQSIVVCVNMIFTALVAQLFLMSLCHLKFGIFLLFASLIIFMSFFVFFLLPETKKVP 558
Query: 482 IEEMDRVWKSHPFWSRFVEHGD 503
IEE+ ++++H FW RFV D
Sbjct: 559 IEEIYLLFENHWFWRRFVTDQD 624
Score = 34.3 bits (77), Expect(3) = 4e-44
Identities = 18/38 (47%), Positives = 24/38 (62%)
Frame = +2
Query: 307 PVLFNSIGFKDDASLMSAVITGVVNVVATCVSIYGVDK 344
P F S GF +ASL S+ IT +VAT +S++ VDK
Sbjct: 98 PCHFQSXGFXXNASLFSSXITNGALLVATVISMFLVDK 211
Score = 29.6 bits (65), Expect(3) = 4e-44
Identities = 17/38 (44%), Positives = 22/38 (57%), Gaps = 1/38 (2%)
Frame = +3
Query: 276 LQRKYRPQLTMAVL-IPFFQQFTGINVIMFYAPVLFNS 312
L+RK L + L I FQQ N I+FYAPV+F +
Sbjct: 3 LKRKXXXXLMIGALGIXAFQQLXXNNSILFYAPVIFRA 116
>TC218649 similar to UP|Q39416 (Q39416) Integral membrane protein, partial
(93%)
Length = 1695
Score = 172 bits (435), Expect = 4e-43
Identities = 114/405 (28%), Positives = 203/405 (49%), Gaps = 2/405 (0%)
Frame = +2
Query: 83 TMFTSSLYLAALLSSLVASTITRRFGRKLSMLFGGLLFLVGALINGFANHVWMLIVGRIL 142
++F S + A++ ++ + I GRK S++ + ++G L FA L +GR+L
Sbjct: 293 SLFGSLSNVGAMVGAIASGQIAEYIGRKGSLMIASIPNIIGWLAISFAKDSSFLYMGRLL 472
Query: 143 LGFGIGFANQAVPLYLSEMAPYKYRGALNIGFQLSITIGILVANVLNYFFAKIKGGWGWR 202
GFG+G + VP+Y++E++P RG L QLS+TIGI++A +L F WR
Sbjct: 473 EGFGVGIISYTVPVYIAEISPPNLRGGLVSVNQLSVTIGIMLAYLLGIFVE-------WR 631
Query: 203 LSLGGAMVPALIITIGSLVLPDTPNSMIERGDRDGAKAQLKRIRGIE-DVDEEFNDLV-A 260
+ ++P I+ +P++P + + G + + L+ +RG + D+ E N++ A
Sbjct: 632 ILAIIGILPCTILIPALFFIPESPRWLAKMGMTEEFETSLQVLRGFDTDISVEVNEIKRA 811
Query: 261 ASEASMQVENPWRNLLQRKYRPQLTMAVLIPFFQQFTGINVIMFYAPVLFNSIGFKDDAS 320
+ + ++ + +L QR+Y L + + + QQ +GIN ++FY+ +F + G +
Sbjct: 812 VASTNTRITVRFADLKQRRYWLPLMIGIGLLILQQLSGINGVLFYSSTIFRNAGISSSDA 991
Query: 321 LMSAVITGVVNVVATCVSIYGVDKWGRRALFLEGGAQMLICQVAVAAAIGAKFGTSGNPG 380
V G V V+AT ++++ DK GRR L + M + VA K S
Sbjct: 992 ATFGV--GAVQVLATSLTLWLADKSGRRLLLIVSATGMSFSLLVVAITFYIKASIS-ETS 1162
Query: 381 NLPEWYAIVVVLFICIYVAGFAWSWGPLGWLVPSEIFPLEIRSAAQSVNVSVNMLFTFLV 440
+L + + ++ + V F+ G + W++ SEI P+ I+ A SV N LF++LV
Sbjct: 1163SLYGILSTLSLVGVVAMVIAFSLGMGAMPWIIMSEILPINIKGLAGSVATLANWLFSWLV 1342
Query: 441 AQVFLIMLCHMKFGLFLFFAFFVLVMSIYVFFLLPETKGIPIEEM 485
++L G F +A + ++V +PETKG IEE+
Sbjct: 1343TLTANMLLDWSSGGTFTIYAVVCALTVVFVTIWVPETKGKTIEEI 1477
>TC211459 homologue to UP|Q7XA52 (Q7XA52) Monosaccharide transporter, partial
(19%)
Length = 416
Score = 165 bits (417), Expect = 5e-41
Identities = 82/96 (85%), Positives = 88/96 (91%)
Frame = +3
Query: 2 AGGGIPIGGGNKEYPGNLTPFVTITCIVAAMGGLIFGYDIGISGGVTSMDPFLKKFFPAV 61
A GGI GGG KEYPG+LT FVT+TCIVAAMGGLIFGYDIGISGGVTSMDPFL KFFP+V
Sbjct: 132 AVGGINTGGG-KEYPGSLTLFVTVTCIVAAMGGLIFGYDIGISGGVTSMDPFLLKFFPSV 308
Query: 62 YRKKNKDKSTNQYCQYDSQTLTMFTSSLYLAALLSS 97
+RKKN DK+ NQYCQYDSQTLTMFTS+LYLAALLSS
Sbjct: 309 FRKKNSDKTVNQYCQYDSQTLTMFTSTLYLAALLSS 416
>TC234206 weakly similar to GB|AAO64833.1|29028908|BT005898 At1g34580
{Arabidopsis thaliana;} , partial (23%)
Length = 740
Score = 152 bits (385), Expect = 3e-37
Identities = 69/146 (47%), Positives = 99/146 (67%), Gaps = 4/146 (2%)
Frame = +1
Query: 358 MLIC----QVAVAAAIGAKFGTSGNPGNLPEWYAIVVVLFICIYVAGFAWSWGPLGWLVP 413
++IC ++AV+A + G G ++ + A++V++ +C Y AGF WSWGPL WL+P
Sbjct: 136 IIICVGMLKIAVSALLAMVTGVHGTK-DISKGNAMLVLVLLCFYDAGFGWSWGPLTWLIP 312
Query: 414 SEIFPLEIRSAAQSVNVSVNMLFTFLVAQVFLIMLCHMKFGLFLFFAFFVLVMSIYVFFL 473
SEIFPL+IR+ QS+ V V + F ++Q FL MLCH KFG FLF+ ++ VM++++ F
Sbjct: 313 SEIFPLKIRTTGQSIAVGVQFIALFALSQTFLTMLCHFKFGAFLFYTVWIAVMTLFIMFF 492
Query: 474 LPETKGIPIEEMDRVWKSHPFWSRFV 499
LPETKGIP+E M +W H FW RFV
Sbjct: 493 LPETKGIPLESMYTIWGKHWFWGRFV 570
>AW234900 weakly similar to GP|16945177|em STP5 protein {Arabidopsis
thaliana}, partial (24%)
Length = 405
Score = 147 bits (372), Expect = 9e-36
Identities = 70/132 (53%), Positives = 93/132 (70%)
Frame = +2
Query: 286 MAVLIPFFQQFTGINVIMFYAPVLFNSIGFKDDASLMSAVITGVVNVVATCVSIYGVDKW 345
MA+ IPFFQQ TGIN++ FYAP +F S+G DA+L+SA+I G VN+V+ VS VD++
Sbjct: 11 MAIAIPFFQQMTGINIVAFYAPNIFQSVGLGHDAALLSAIILGAVNLVSLLVSTAIVDRF 190
Query: 346 GRRALFLEGGAQMLICQVAVAAAIGAKFGTSGNPGNLPEWYAIVVVLFICIYVAGFAWSW 405
GRR LF+ GG ML+CQ+AV+ + G G ++ AIVV++ +C Y AGF WSW
Sbjct: 191 GRRFLFVTGGICMLVCQIAVSILLAVVTGVHGTK-DMSNGSAIVVLVLLCCYTAGFGWSW 367
Query: 406 GPLGWLVPSEIF 417
GPL WL+PSEIF
Sbjct: 368 GPLTWLIPSEIF 403
Score = 28.5 bits (62), Expect = 7.5
Identities = 16/38 (42%), Positives = 24/38 (63%), Gaps = 2/38 (5%)
Frame = +2
Query: 87 SSLYLAA--LLSSLVASTITRRFGRKLSMLFGGLLFLV 122
S++ L A L+S LV++ I RFGR+ + GG+ LV
Sbjct: 122 SAIILGAVNLVSLLVSTAIVDRFGRRFLFVTGGICMLV 235
>TC226707 similar to UP|Q7XA50 (Q7XA50) Sorbitol-like transporter, partial
(62%)
Length = 2338
Score = 132 bits (333), Expect = 3e-31
Identities = 89/325 (27%), Positives = 156/325 (47%), Gaps = 19/325 (5%)
Frame = +3
Query: 182 ILVANVLNYFFAKIKGGWGWRLSLGGAMVPALIITIGSLVLPDTPNSMIERGDRDGAKAQ 241
IL+ + NY F+K+ GWRL LG +P+++I + L +P++P ++ +G AK
Sbjct: 3 ILIGYISNYGFSKLALRLGWRLMLGVGAIPSILIGVAVLAMPESPRWLVAKGRLGEAKRV 182
Query: 242 LKRIRGIED--------------VDEEFNDLVAASEASMQVENPWRNLLQRK---YRPQL 284
L +I E+ + ++ +D V WR L R
Sbjct: 183 LYKISESEEEARLRLADIKDTAGIPQDCDDDVVLVSKQTHGHGVWRELFLHPTPAVRHIF 362
Query: 285 TMAVLIPFFQQFTGINVIMFYAPVLFNSIGFK-DDASLMSAVITGVVNVVATCVSIYGVD 343
++ I FF Q TGI+ ++ Y+P +F G K D+ L++ V G V V+ V+ + +D
Sbjct: 363 IASLGIHFFAQATGIDAVVLYSPRIFEKAGIKSDNYRLLATVAVGFVKTVSILVATFFLD 542
Query: 344 KWGRRALFLEGGAQMLICQVAVAAAIGAKFGTSGNPGNLPEWYAIVVVLFICIYVAGFAW 403
+ GRR L L + +++ + +G + W + + + YVA F+
Sbjct: 543 RAGRRVLLLCSVSGLILSLLT----LGLSLTVVDHSQTTLNWAVGLSIAAVLSYVATFSI 710
Query: 404 SWGPLGWLVPSEIFPLEIRSAAQSVNVSVNMLFTFLVAQVFLIMLCHMKF-GLFLFFAFF 462
GP+ W+ SEIFPL +R+ ++ +VN + + ++A FL + + G F FA
Sbjct: 711 GSGPITWVYSSEIFPLRLRAQGVAIGAAVNRVTSGVIAMTFLSLQKAITIGGAFFLFAGV 890
Query: 463 VLVMSIYVFFLLPETKGIPIEEMDR 487
V I+ + LLPET+G +EE+++
Sbjct: 891 AAVAWIFHYTLLPETRGKTLEEIEK 965
>BE346786 similar to GP|5734440|emb hexose transporter {Lycopersicon
esculentum}, partial (19%)
Length = 435
Score = 127 bits (318), Expect = 2e-29
Identities = 59/115 (51%), Positives = 77/115 (66%), Gaps = 1/115 (0%)
Frame = -1
Query: 390 VVLFICIYVAGFAWSWGPLGWLVPSEIFPLEIRSAAQSVNVSVNMLFTFLVAQVFLIMLC 449
VV+ +CI + GPLGW +PSEIFPLE RS Q ++V VN LFTF++ Q MLC
Sbjct: 435 VVVMVCICMV-----MGPLGWFIPSEIFPLETRSVGQGLSVCVNFLFTFVIGQAVFSMLC 271
Query: 450 HMKFGLFLFFAFFVLVMSIYVFFLLPETKGIPIEEM-DRVWKSHPFWSRFVEHGD 503
+FG+F FF ++L+MS +V FL PET +PIEEM +RVWK H W RF++ D
Sbjct: 270 LFRFGIFFFFYGWILIMSTFVLFLFPETNNVPIEEMAERVWKQHWLWKRFIDEDD 106
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.327 0.143 0.444
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 23,827,254
Number of Sequences: 63676
Number of extensions: 349667
Number of successful extensions: 3251
Number of sequences better than 10.0: 125
Number of HSP's better than 10.0 without gapping: 3136
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3184
length of query: 518
length of database: 12,639,632
effective HSP length: 102
effective length of query: 416
effective length of database: 6,144,680
effective search space: 2556186880
effective search space used: 2556186880
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 61 (28.1 bits)
Medicago: description of AC146792.3


