

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146790.7 + phase: 0
(546 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
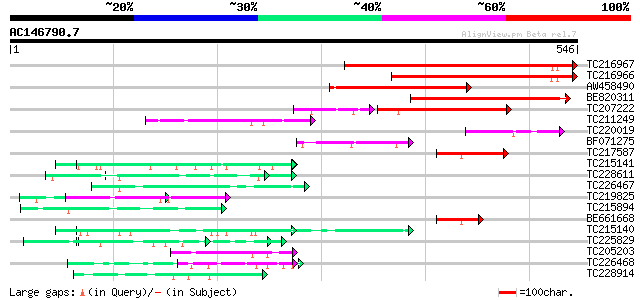
Score E
Sequences producing significant alignments: (bits) Value
TC216967 278 4e-75
TC216966 weakly similar to UP|Q6PHL0 (Q6PHL0) Wu:fd15e07 protein... 212 4e-55
AW458490 weakly similar to GP|20147195|gb| AT5g66100/K2A18_18 {A... 197 1e-50
BE820311 176 2e-44
TC207222 130 1e-41
TC211249 similar to UP|FXE3_MOUSE (Q9QY14) Forkhead box protein ... 132 3e-31
TC220019 66 3e-11
BF071275 similar to PIR|S14981|S149 extensin class I (clone w1-8... 65 6e-11
TC217587 similar to UP|Q94A38 (Q94A38) AT5g46250/MPL12_3, partia... 55 6e-08
TC215141 similar to GB|AAL32703.1|17065098|AY062625 nucleotide s... 50 3e-06
TC228611 weakly similar to GB|CAA72487.1|1890317|ATP26A peroxida... 50 3e-06
TC226467 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 pr... 49 4e-06
TC219825 weakly similar to UP|MID2_YEAST (P36027) Mating process... 49 4e-06
TC215894 similar to UP|Q96569 (Q96569) L-lactate dehydrogenase ... 49 6e-06
BE661668 similar to GP|10177706|dbj contains similarity to RNA-b... 48 1e-05
TC215140 similar to GB|AAL32703.1|17065098|AY062625 nucleotide s... 48 1e-05
TC225829 weakly similar to UP|Q9C5Q6 (Q9C5Q6) Fasciclin-like ara... 48 1e-05
TC205203 similar to UP|Q39763 (Q39763) Proline-rich cell wall pr... 47 3e-05
TC226468 weakly similar to UP|O65450 (O65450) Glycine-rich prote... 46 4e-05
TC228914 similar to UP|DAH1_MOUSE (Q9QYB2) Dachshund homolog 1 (... 46 4e-05
>TC216967
Length = 1117
Score = 278 bits (711), Expect = 4e-75
Identities = 137/237 (57%), Positives = 174/237 (72%), Gaps = 13/237 (5%)
Frame = +1
Query: 323 DRGNYANTRDAHAPQPRMPPRGILRPPPPSTAAFLGPQTIGPFPAPVAYPDFYYFPTVPL 382
DR YANT DAH Q RMPPRG++RPPPP+ A F+GPQ + P P +P++YYFPT+
Sbjct: 1 DRVGYANTTDAHVNQQRMPPRGLMRPPPPNPAPFMGPQPMRPLANPAGFPEYYYFPTLQF 180
Query: 383 DPFRGMPVFPHMPSPATFFPAAESSLSNVIVNQIDYYFSDINLANDEFLKSNMDEEGWVP 442
+PF GMP F H P PA F+P AE+ L+N I NQIDYYFSD NL DE+L+SNMDE+GWVP
Sbjct: 181 EPFGGMPFFTHAPPPAMFYPVAETPLTNTIANQIDYYFSDANLVKDEYLRSNMDEQGWVP 360
Query: 443 ITLIANFPRVKNLTSNIQLILDSMRNSNVVDVQGDKLRRRN-WIDRLTSAQVQADAGSVS 501
ITLIA+FPRV++LTSNI+LILDS+R S VV+VQGDKLRR N W+ L SAQ++A++GS+S
Sbjct: 361 ITLIASFPRVRSLTSNIKLILDSLRTSTVVEVQGDKLRRCNEWMKWLPSAQLRANSGSIS 540
Query: 502 PIESRDNSFTADFQTITLDK----------TTKDE--GESSRQSQLSNGSDVAGNIN 546
P S N+ ADF+ IT+D+ TT ++ GESS QSQ+ N + V+ N N
Sbjct: 541 PSGSSSNNLAADFEKITVDEAISHKVNTEPTTNEDAAGESSTQSQVPNDATVSSN*N 711
>TC216966 weakly similar to UP|Q6PHL0 (Q6PHL0) Wu:fd15e07 protein (Fragment),
partial (10%)
Length = 1027
Score = 212 bits (539), Expect = 4e-55
Identities = 108/190 (56%), Positives = 137/190 (71%), Gaps = 11/190 (5%)
Frame = +1
Query: 368 PVAYPDFYYFPTVPLDPFRGMPVFPHMPSPATFFPAAESSLSNVIVNQIDYYFSDINLAN 427
P +P++YYFPT+ +PF GMP F H P PA F+P AE+ L+N I NQIDYYFSD NL
Sbjct: 13 PAGFPEYYYFPTLQFEPFGGMPFFTHAPPPAMFYPVAETPLTNTIANQIDYYFSDANLVK 192
Query: 428 DEFLKSNMDEEGWVPITLIANFPRVKNLTSNIQLILDSMRNSNVVDVQGDKLRRR-NWID 486
DE+L+SNMDE+GWVPI+LIA+FPRV++LTSNI+LILDS+R S V+VQGDKLRRR W+
Sbjct: 193 DEYLRSNMDEQGWVPISLIASFPRVRSLTSNIKLILDSLRTSTFVEVQGDKLRRRTEWMK 372
Query: 487 RLTSAQVQADAGSVSPIESRDNSFTADFQTITLD-------KTTKDE--GESSRQSQLSN 537
L S Q+ A +GS+SP ES N+ ADF+ IT+D T D+ GESS QSQ+ N
Sbjct: 373 WLPSVQLLAKSGSISPSESSSNNLAADFEKITVDDAKVNAKPTANDDAAGESSTQSQVPN 552
Query: 538 -GSDVAGNIN 546
+ V+ N N
Sbjct: 553 DATTVSSN*N 582
>AW458490 weakly similar to GP|20147195|gb| AT5g66100/K2A18_18 {Arabidopsis
thaliana}, partial (7%)
Length = 411
Score = 197 bits (501), Expect = 1e-50
Identities = 91/137 (66%), Positives = 107/137 (77%), Gaps = 1/137 (0%)
Frame = +3
Query: 309 GSYHQNSYSSRREH-DRGNYANTRDAHAPQPRMPPRGILRPPPPSTAAFLGPQTIGPFPA 367
GSYH NSY SRR+H DRGNY N RDA PQPRMPPRG+LR PPP+TAAF+GPQ IGPF
Sbjct: 3 GSYH-NSYGSRRDHQDRGNYVNARDAFLPQPRMPPRGLLRHPPPNTAAFVGPQPIGPFAN 179
Query: 368 PVAYPDFYYFPTVPLDPFRGMPVFPHMPSPATFFPAAESSLSNVIVNQIDYYFSDINLAN 427
P+++P+FYY+ V ++ F GMP F H P P FF AAES LSN+IV QI+YYFSD NL
Sbjct: 180 PISFPEFYYYQPVTVEQFTGMPFFTHSPPPTPFFSAAESPLSNMIVYQIEYYFSDANLVK 359
Query: 428 DEFLKSNMDEEGWVPIT 444
D FL+S MDE+GWVP+T
Sbjct: 360 DAFLRSKMDEQGWVPVT 410
>BE820311
Length = 680
Score = 176 bits (447), Expect = 2e-44
Identities = 93/155 (60%), Positives = 113/155 (72%), Gaps = 1/155 (0%)
Frame = -3
Query: 387 GMPVFPHMPSPATFFPAAESSLSNVIVNQIDYYFSDINLANDEFLKSNMDEEGWVPITLI 446
GMP F H P AAES LSN+IV QI+YYFSD NL D FL+S MDE+GWVP+TLI
Sbjct: 672 GMPFFTHSPXXTPXXXAAESPLSNMIVYQIEYYFSDANLVKDAFLRSKMDEQGWVPVTLI 493
Query: 447 ANFPRVKNLTSNIQLILDSMRNSNVVDVQGDKLRRRN-WIDRLTSAQVQADAGSVSPIES 505
A+FPRVKNLT+NIQLILDS+R S VV+VQGDKLRR N W+ L SAQ ++++GS+SP S
Sbjct: 492 ADFPRVKNLTTNIQLILDSLRTSTVVEVQGDKLRRLNEWMRWLPSAQRRSNSGSISPSGS 313
Query: 506 RDNSFTADFQTITLDKTTKDEGESSRQSQLSNGSD 540
N+ TA+FQTI L++TT DE + NG D
Sbjct: 312 THNNLTANFQTIILEETTADESPTQ-----PNGGD 223
>TC207222
Length = 2191
Score = 130 bits (326), Expect(2) = 1e-41
Identities = 64/135 (47%), Positives = 87/135 (64%), Gaps = 6/135 (4%)
Frame = +1
Query: 355 AFLGPQTIGPFPAPVAYPD------FYYFPTVPLDPFRGMPVFPHMPSPATFFPAAESSL 408
+F P + P +P+ + + F P P D RG+P P MP FF + L
Sbjct: 499 SFFHPSPMRPIGSPIGFHELAPPLVFVAAPPPPPDSLRGVPFVPPMPHHPLFFTGPDPQL 678
Query: 409 SNVIVNQIDYYFSDINLANDEFLKSNMDEEGWVPITLIANFPRVKNLTSNIQLILDSMRN 468
N IVNQ+DYYFS+ NL D FL+ NMD++GWVPI LIA F +V +LT NIQ+ILD+++
Sbjct: 679 HNKIVNQVDYYFSNENLVKDTFLRQNMDDQGWVPIKLIAGFNKVMHLTDNIQVILDAIQT 858
Query: 469 SNVVDVQGDKLRRRN 483
S+VV+VQGDK+RR+N
Sbjct: 859 SSVVEVQGDKIRRQN 903
Score = 58.9 bits (141), Expect(2) = 1e-41
Identities = 34/82 (41%), Positives = 48/82 (58%), Gaps = 4/82 (4%)
Frame = +3
Query: 274 DSSPRDHHRNNNWDAR--PFVGGSSRRGHFGSHPRGDGSYHQNSYSSRREHDRGNYAN-- 329
+SSP+DH + + + + P S R + G H RGDGS+H N Y +RR+ + N +
Sbjct: 243 NSSPKDHTQRSGFASNDHPQQRNSFRNRNGGQHQRGDGSHHHN-YGNRRDQEWNNNXSFG 419
Query: 330 TRDAHAPQPRMPPRGILRPPPP 351
+RD H P PR+ PR I PPPP
Sbjct: 420 SRDTHVP-PRVAPRFIRPPPPP 482
>TC211249 similar to UP|FXE3_MOUSE (Q9QY14) Forkhead box protein E3, partial
(5%)
Length = 536
Score = 132 bits (333), Expect = 3e-31
Identities = 76/172 (44%), Positives = 101/172 (58%), Gaps = 8/172 (4%)
Frame = +1
Query: 131 KPAWNKQPLNGVAVEIGPVMGAESWPALSESAKIPGKLPPESSSSKIAPPAAVDGSPSTS 190
KPAW K NGV+ E+GPVMGA SWP LSES K KL +S S + A + +T
Sbjct: 4 KPAWKKPESNGVS-EVGPVMGAHSWPDLSESTKPTAKL---TSDSSVKTAADEESLTTTP 171
Query: 191 QGPIISHSPQRQG-STSNTKSSSMANNNLPNRPRPMRRISGS---NIGPCPAQSSLS--- 243
Q P+ S SPQ+Q +N + N + NR R M++ GS N+G P Q++ S
Sbjct: 172 QDPVNSDSPQKQAIGNANPSPTPAMNYGMANRQRSMKQRGGSNGYNVGSGPVQNNFSGAT 351
Query: 244 -NPPTPPPLPPYPVYQLPPAVSYPNMLPSIPDSSPRDHHRNNNWDARPFVGG 294
N P PPP PP+PV +PP ++ + +P +P +PRD +RNNNWDARP GG
Sbjct: 352 QNLPPPPPPPPFPVLPMPPR-TFAHGIPGVPVPTPRDPYRNNNWDARPPHGG 504
Score = 30.4 bits (67), Expect = 2.1
Identities = 34/131 (25%), Positives = 50/131 (37%), Gaps = 1/131 (0%)
Frame = +1
Query: 41 VRGGSGWDTESQSQSPTGIHQSLPS-SSSSSSSSLTTVDQPPPSDDSPKAAVVSSSPKAV 99
V G W S+S PT S S +++ SLTT Q P V S SP+
Sbjct: 55 VMGAHSWPDLSESTKPTAKLTSDSSVKTAADEESLTTTPQDP---------VNSDSPQKQ 207
Query: 100 VVSSPLPAPMEKNSNTIASDGGDGGNAGGSKKPAWNKQPLNGVAVEIGPVMGAESWPALS 159
+ + P+P + +A+ GGS NG V GPV ++ +
Sbjct: 208 AIGNANPSPTPAMNYGMANRQRSMKQRGGS----------NGYNVGSGPVQ--NNFSGAT 351
Query: 160 ESAKIPGKLPP 170
++ P PP
Sbjct: 352 QNLPPPPPPPP 384
>TC220019
Length = 582
Score = 66.2 bits (160), Expect = 3e-11
Identities = 42/99 (42%), Positives = 55/99 (55%), Gaps = 4/99 (4%)
Frame = +2
Query: 440 WVPITLIANFPRVKNLTSNIQLILDSMRNSNVVDVQGDKLRRRN----WIDRLTSAQVQA 495
WV I LIA F +VK LT NIQ++LD++R S+VV+VQGDK+RRRN WI + + QV
Sbjct: 2 WVTINLIAGFKKVKYLTENIQIVLDAVRTSSVVEVQGDKIRRRNDWRRWI--MPAGQVPN 175
Query: 496 DAGSVSPIESRDNSFTADFQTITLDKTTKDEGESSRQSQ 534
GS + Q ITL+ T + SQ
Sbjct: 176 VRGSQTV-----GQLAEQVQNITLETTNNNNAGILDDSQ 277
>BF071275 similar to PIR|S14981|S149 extensin class I (clone w1-8 L) - tomato
(fragment), partial (10%)
Length = 389
Score = 65.5 bits (158), Expect = 6e-11
Identities = 45/127 (35%), Positives = 57/127 (44%), Gaps = 14/127 (11%)
Frame = +1
Query: 277 PRDH--HRNNNWDARPFVGGSSRRGHFGSHPRGDGSYHQNSYSSRREHDRGNY------- 327
P DH HRN+ R G H RGDG +H N+Y RR+ D GN
Sbjct: 25 PNDHPPHRNS----------FRHRNGGGPHQRGDGHHHHNNYGGRRDQDPGNQDWNNHRN 174
Query: 328 ANTRDAHAPQPRMPPRGILRPPPPSTA-AFLGPQTIGPFPAPVAY----PDFYYFPTVPL 382
N RD + PR PR I PPPP+ A F P + P+ + + P Y P PL
Sbjct: 175 FNGRD-NFMSPRFGPRFIRPPPPPNPAQLFPPPPPLRPYGGSIGFTELPPQMVYVPPPPL 351
Query: 383 DPFRGMP 389
+ RG+P
Sbjct: 352 ESMRGVP 372
>TC217587 similar to UP|Q94A38 (Q94A38) AT5g46250/MPL12_3, partial (27%)
Length = 724
Score = 55.5 bits (132), Expect = 6e-08
Identities = 27/71 (38%), Positives = 47/71 (66%), Gaps = 2/71 (2%)
Frame = +2
Query: 412 IVNQIDYYFSDINLANDEFLKS--NMDEEGWVPITLIANFPRVKNLTSNIQLILDSMRNS 469
I+ Q +YYFSD NL D++L ++EG+VP+++IA+F ++K LT + I +++ S
Sbjct: 362 IIKQAEYYFSDENLPTDKYLLGFVKRNKEGFVPVSVIASFRKIKKLTRDHAFIAAALKES 541
Query: 470 NVVDVQGDKLR 480
+++ V GD R
Sbjct: 542 SLLVVSGDGRR 574
>TC215141 similar to GB|AAL32703.1|17065098|AY062625 nucleotide sugar
epimerase-like protein {Arabidopsis thaliana;} , partial
(89%)
Length = 1743
Score = 50.1 bits (118), Expect = 3e-06
Identities = 70/263 (26%), Positives = 94/263 (35%), Gaps = 30/263 (11%)
Frame = +2
Query: 45 SGWDTESQSQSPTGIHQSLP----SSSSSSSSSLTTVDQPPP----SDDSPKAAVVSSSP 96
SG T S +P S P SSSSSSS TT+ PPP + +P S P
Sbjct: 71 SGDSTAPSSSTPPPNSSSAPLSWSHSSSSSSSPSTTLPSPPPTPPTATSTPTPTTSSPPP 250
Query: 97 KAVVVSSPLPAPMEKNSNTIASDGGDGGNAGGSKKPAW------------------NKQP 138
+ S P + T + G A + W P
Sbjct: 251 PSAPGRSRSATPPPPAAPTASPSLSLGLRASWAATAPWP*RSAATVSWGLTTSTTTTTLP 430
Query: 139 LNGVAVEIGPVMGAESWPALSESAKIPGKLPPESSSSKIAPPAAVDGSPSTSQGP--IIS 196
LN + SW K+ PP S SS + P SP++S P S
Sbjct: 431 LNAPGRQCSGNTKCSSW-------KVTSTTPPCSRSSSTSSP-----SPTSSTWPHRPAS 574
Query: 197 HSPQRQGSTSN--TKSSSMANNNLPNRPRPMRRISGSNIGPCPAQSSLSNPPTPPPLPPY 254
+P R + ++ T +S + LPN P P +SG PA S+ S P T P
Sbjct: 575 ATPCRTPNPTSPPTSLASSTFSRLPNPPTPNPPLSGLR----PAPSTASTPKTLSPSSTA 742
Query: 255 PVYQLPPAVSYPNMLPSIPDSSP 277
P PPA + P P+ +P
Sbjct: 743 PT--SPPASTPPPRRPARKSPTP 805
Score = 43.9 bits (102), Expect = 2e-04
Identities = 56/236 (23%), Positives = 91/236 (37%), Gaps = 24/236 (10%)
Frame = +2
Query: 65 SSSSSSSSSLTTVDQPPPSDDS-----PKAAVVSSSPKAVVVSSPLPAPMEKNSNTIASD 119
++++S S+ + PPP+ S ++ SSSP + S P P ++ T +
Sbjct: 59 TTATSGDSTAPSSSTPPPNSSSAPLSWSHSSSSSSSPSTTLPSPPPTPPTATSTPTPTTS 238
Query: 120 GGDGGNAGGSKKPAWNKQPLNGVAVEIGPVMGAESWPALS-----ESAKIPGKLPPESSS 174
+A G + A P A + SW A + +A + L +++
Sbjct: 239 SPPPPSAPGRSRSATPPPPAAPTASPSLSLGLRASWAATAPWP*RSAATVSWGLTTSTTT 418
Query: 175 SKI---APPAAVDGSPSTSQGPIISHSPQRQGSTSNTKSSSMANNNLPNRPRPMRRISGS 231
+ + AP G+ S + S +P S S+T S S ++ P+RP
Sbjct: 419 TTLPLNAPGRQCSGNTKCSSWKVTSTTPPCSRS-SSTSSPSPTSSTWPHRPASATPCRTP 595
Query: 232 NIGPCPAQ------SSLSNPPTP-PPL----PPYPVYQLPPAVSYPNMLPSIPDSS 276
N P S L NPPTP PPL P P +S + P+ P +S
Sbjct: 596 NPTSPPTSLASSTFSRLPNPPTPNPPLSGLRPAPSTASTPKTLSPSSTAPTSPPAS 763
>TC228611 weakly similar to GB|CAA72487.1|1890317|ATP26A peroxidase ATP26a
{Arabidopsis thaliana;} , partial (49%)
Length = 731
Score = 49.7 bits (117), Expect = 3e-06
Identities = 50/188 (26%), Positives = 70/188 (36%), Gaps = 4/188 (2%)
Frame = +3
Query: 93 SSSPKAVVVSSP--LPAPMEKNSNTIASDGGDGGNAG-GSKKPAWNKQPLNGVAVEIGPV 149
SSSP + S+P +P + S G S+ PA P P
Sbjct: 42 SSSPHCYLFSAPPTRDSPWTSTRTLVRSSARS*GTPSQASRSPAPPPPP---------PR 194
Query: 150 MGAESWPALSESAKIP-GKLPPESSSSKIAPPAAVDGSPSTSQGPIISHSPQRQGSTSNT 208
+ S A S +A P PP S A P + SP+T P S S R +S
Sbjct: 195 SASSSTTASSPTAVTPPSSSPPRPSPGPSATPTSTSPSPAT---PSTSSSAPRPPLSSPA 365
Query: 209 KSSSMANNNLPNRPRPMRRISGSNIGPCPAQSSLSNPPTPPPLPPYPVYQLPPAVSYPNM 268
+ S A P P S + P + ++ + PP+PPP PP P++ PN
Sbjct: 366 PTPSPAPTFSPPPPATSSPCSAAPSSPSSSAAATAEPPSPPPYPPTSPPPPCPSLKSPNS 545
Query: 269 LPSIPDSS 276
P+ S
Sbjct: 546 SPNAASPS 569
Score = 42.4 bits (98), Expect = 5e-04
Identities = 52/226 (23%), Positives = 83/226 (36%), Gaps = 10/226 (4%)
Frame = +3
Query: 35 SPWAQ---VVRGGS-GWDTESQSQSPTGIHQSLPSSSSSSSSSLTTVDQPPPSDDSPKAA 90
SPW +VR + T SQ+ P S+SSS+++ + PPS P+
Sbjct: 90 SPWTSTRTLVRSSARS*GTPSQASRSPAPPPPPPRSASSSTTASSPTAVTPPSSSPPR-- 263
Query: 91 VVSSSPKAVVVSSPLPAPMEKNSNTIASDGGDGGNAGGSKKPAWNKQPLNGVAVEIGPVM 150
P P P ++T S ++ + P + P A P
Sbjct: 264 -------------PSPGPSATPTSTSPSPATPSTSSSAPRPPLSSPAPTPSPAPTFSPPP 404
Query: 151 GAESWPALSESAKIPGKLPPESSSSKIAPPAAVDGSPSTSQGPIISHSPQRQGSTSNTKS 210
A S P + + P SS++ A P + P TS P P + S+ +
Sbjct: 405 PATSSPCSAAPSS------PSSSAAATAEPPSPPPYPPTSPPP---PCPSLKSPNSSPNA 557
Query: 211 SSMANNNLPNRPRPMRRISGSNIGPCPA------QSSLSNPPTPPP 250
+S + N+ P ++GP P+ SS ++P T PP
Sbjct: 558 ASPSKNSSP------------SVGPTPSDSLTAPSSSRTSPTTLPP 659
Score = 39.3 bits (90), Expect = 0.005
Identities = 49/184 (26%), Positives = 69/184 (36%), Gaps = 14/184 (7%)
Frame = +3
Query: 4 TQNNHSPTSVPSPAPPSAETNNPNFIRKNLPSPWAQVVRGGSGWDTESQSQSPTGIHQSL 63
+Q + SP P P P SA ++ + P + R G S SP+ +
Sbjct: 150 SQASRSPAP-PPPPPRSASSSTTASSPTAVTPPSSSPPRPSPGPSATPTSTSPS---PAT 317
Query: 64 PSSSSSSS-----------SSLTTVDQPPPSDDSPKAAVVS--SSPKAVVVSSPLPAPME 110
PS+SSS+ S T PPP+ SP +A S SS A P P P
Sbjct: 318 PSTSSSAPRPPLSSPAPTPSPAPTFSPPPPATSSPCSAAPSSPSSSAAATAEPPSPPPYP 497
Query: 111 KNSNTIASDGGDGGNAG-GSKKPAWNKQPLNGVAVEIGPVMGAESWPALSESAKIPGKLP 169
S N+ + P+ N P +GP ++S A S S P LP
Sbjct: 498 PTSPPPPCPSLKSPNSSPNAASPSKNSSP------SVGPT-PSDSLTAPSSSRTSPTTLP 656
Query: 170 PESS 173
P ++
Sbjct: 657 PPTT 668
>TC226467 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 protein
precursor, partial (6%)
Length = 1017
Score = 49.3 bits (116), Expect = 4e-06
Identities = 51/213 (23%), Positives = 72/213 (32%), Gaps = 3/213 (1%)
Frame = +3
Query: 79 QPPPSDDSPKAAVVSSSPKAVVVSSP---LPAPMEKNSNTIASDGGDGGNAGGSKKPAWN 135
Q P + +P A +S P +SP P + +AS + P
Sbjct: 30 QLPSNAQAPATAPANSPPTTPAATSPSSGTPLAVTLPPTVVASPPT-------TSTPPTT 188
Query: 136 KQPLNGVAVEIGPVMGAESWPALSESAKIPGKLPPESSSSKIAPPAAVDGSPSTSQGPII 195
QP VA + PV A + S K P P+ K++P P P+
Sbjct: 189 SQPPAIVAPKPAPVTPPAPKVAPASSPKAPPPQTPQPQPPKVSPVVTPTSPPPIPPPPVS 368
Query: 196 SHSPQRQGSTSNTKSSSMANNNLPNRPRPMRRISGSNIGPCPAQSSLSNPPTPPPLPPYP 255
+ P + T + + P P P + P PA + PTP P+PP P
Sbjct: 369 TPPPLPPPKIAPTPAKT------PPAPAPAKAT------PAPAPAPTKPAPTPAPVPPPP 512
Query: 256 VYQLPPAVSYPNMLPSIPDSSPRDHHRNNNWDA 288
P V P PS R HR+ A
Sbjct: 513 TLAPTPVVEVPAPAPS-SHKHRRHRHRHRRHQA 608
Score = 31.2 bits (69), Expect = 1.2
Identities = 24/75 (32%), Positives = 32/75 (42%), Gaps = 4/75 (5%)
Frame = +3
Query: 333 AHAPQPRMPPRGILRPPPPSTAAFLGPQTIGPF-PAPVAYPDFYYFPTVPLDPFRGMPV- 390
A A P+ PP +P PP + + P + P P PV+ P PL P + P
Sbjct: 252 APASSPKAPPPQTPQPQPPKVSPVVTPTSPPPIPPPPVSTPP-------PLPPPKIAPTP 410
Query: 391 --FPHMPSPATFFPA 403
P P+PA PA
Sbjct: 411 AKTPPAPAPAKATPA 455
Score = 29.3 bits (64), Expect = 4.7
Identities = 23/71 (32%), Positives = 29/71 (40%), Gaps = 3/71 (4%)
Frame = +3
Query: 336 PQPRMPPRGILRPPPPSTAAFLGPQTIGPFPAPVAYPDFYYFPTVPLDPFRGMPVFPH-- 393
P P++ P + PPP T PQ P +PV P P +P P P P
Sbjct: 237 PAPKVAPASSPKAPPPQT-----PQPQPPKVSPVVTPTSP--PPIPPPPVSTPPPLPPPK 395
Query: 394 -MPSPATFFPA 403
P+PA PA
Sbjct: 396 IAPTPAKTPPA 428
>TC219825 weakly similar to UP|MID2_YEAST (P36027) Mating process protein
MID2 (Serine-rich protein SMS1) (Protein kinase A
interference protein), partial (19%)
Length = 862
Score = 49.3 bits (116), Expect = 4e-06
Identities = 45/160 (28%), Positives = 69/160 (43%), Gaps = 1/160 (0%)
Frame = +2
Query: 54 QSPTGIHQSLPSSSSSSSSSLTTVDQPPPSDDSPKAAVVSSSPKAVVVSSPLPAPMEKNS 113
Q+PT SSSS+SSSS + PPP SP P SP P+P S
Sbjct: 2 QAPTSGPSFF*SSSSTSSSSSPSPSSPPPPPSSPSPPSTPPRPS----PSPPPSPPSPAS 169
Query: 114 NTIASDGGDGGNAGGSKKPAWNKQPLNGVAVEIGPVMGAESWPALSESAKIPGKLPPESS 173
++ +S G + S + P + + P + S +LS S+ + P +S
Sbjct: 170 SSASSSPSYGSPSS*SSTTPSSSSPWSS*SSPSTPTTPSSS-SSLSSSSSLSF*SPTSTS 346
Query: 174 -SSKIAPPAAVDGSPSTSQGPIISHSPQRQGSTSNTKSSS 212
S +PP++ SPST+ P S + + + + SSS
Sbjct: 347 PPSGTSPPSSPSSSPSTASPP*RSPTTSSRAGSGSPLSSS 466
Score = 43.9 bits (102), Expect = 2e-04
Identities = 51/157 (32%), Positives = 62/157 (39%), Gaps = 11/157 (7%)
Frame = +2
Query: 10 PTSVPSPAPPSAETN----NPNFIRKNLPSPWAQVVRGGSGWDTESQSQSPTGIHQSLPS 65
P S PSPA SA ++ +P+ PS S W + S +PT S
Sbjct: 143 PPSPPSPASSSASSSPSYGSPSS*SSTTPS-------SSSPWSS*SSPSTPTTPSSSSSL 301
Query: 66 SSSSSSSSLTTVDQPPPSDDSPKAAVVSSSPKAVVVSSPLPAPMEKNSNTIASDGGDGGN 125
SSSSS S + PPS SP SSP SSP A S T +S G G
Sbjct: 302 SSSSSLSF*SPTSTSPPSGTSP-----PSSPS----SSPSTASPP*RSPTTSSRAGSGSP 454
Query: 126 AGGSKKPAWNKQPLNGVA-----VEIGPVMG--AESW 155
S P W+ L+ V +G MG ESW
Sbjct: 455 L-SSSPPIWSPAGLSPVFSAWLWCTVGRTMGFSPESW 562
Score = 28.1 bits (61), Expect(2) = 0.58
Identities = 17/52 (32%), Positives = 23/52 (43%)
Frame = +2
Query: 204 STSNTKSSSMANNNLPNRPRPMRRISGSNIGPCPAQSSLSNPPTPPPLPPYP 255
S+S+T SSS + + P P P P + P+PPP PP P
Sbjct: 35 SSSSTSSSSSPSPSSPPPPPS---------SPSPPSTPPRPSPSPPPSPPSP 163
Score = 22.7 bits (47), Expect(2) = 0.58
Identities = 14/35 (40%), Positives = 14/35 (40%)
Frame = +3
Query: 248 PPPLPPYPVYQLPPAVSYPNMLPSIPDSSPRDHHR 282
PPPLP YP P S PR HHR
Sbjct: 282 PPPLPRYP------------HRPHSLFSRPRLHHR 350
>TC215894 similar to UP|Q96569 (Q96569) L-lactate dehydrogenase , complete
Length = 1675
Score = 48.9 bits (115), Expect = 6e-06
Identities = 56/204 (27%), Positives = 79/204 (38%), Gaps = 6/204 (2%)
Frame = +1
Query: 11 TSVPSPAPPSAETNNPNFIRKNLPSPWAQVVRGGSGWDTESQSQS-----PTGIHQSLPS 65
T+ P P PSA T +P+ + P+ W R S T S S PT +
Sbjct: 235 TTPPLPPRPSATTRSPS----SAPATWEWPSRRPSSLRTSPTSSSSSTPTPTSSAARCST 402
Query: 66 SSSSSSSSLTTVDQPPPSDDSPKAAVVSSSPKAVVVSSPLPAPMEKNSNTIASDGGDGGN 125
SS+ SS PPP+ SP A +SSP P PA + + +S G +
Sbjct: 403 SSTPPPSSPAPRSTPPPTPPSPPAPTSASSP-------PAPARSPASHASTSSRGTSPSS 561
Query: 126 AGGSKKPAWNKQPLNGVAVEIGPVMGAESWPALSESAKIPGKLPPESSSSKIAPPAAVDG 185
A PL ++ P + S+P S S+ P SS +PP A
Sbjct: 562 A-----------PLFPLSSVTPPTQLSSSFPTPSTSS------PTSRGSSPASPPTASSA 690
Query: 186 -SPSTSQGPIISHSPQRQGSTSNT 208
+P+ + +S SP ST T
Sbjct: 691 PAPTWTLPASVSSSPITLTSTPRT 762
Score = 37.7 bits (86), Expect = 0.013
Identities = 34/123 (27%), Positives = 49/123 (39%)
Frame = +1
Query: 168 LPPESSSSKIAPPAAVDGSPSTSQGPIISHSPQRQGSTSNTKSSSMANNNLPNRPRPMRR 227
LPP S++ +P +A P+T + P S R TS++ S+ P P
Sbjct: 247 LPPRPSATTRSPSSA----PATWEWPSRRPSSLRTSPTSSSSST----------PTPTSS 384
Query: 228 ISGSNIGPCPAQSSLSNPPTPPPLPPYPVYQLPPAVSYPNMLPSIPDSSPRDHHRNNNWD 287
+ + P SS + TPPP PP P P + S P P SP H ++
Sbjct: 385 AARCSTSSTPPPSSPAPRSTPPPTPPSP--PAPTSAS----SPPAPARSPASHASTSSRG 546
Query: 288 ARP 290
P
Sbjct: 547 TSP 555
Score = 33.5 bits (75), Expect = 0.25
Identities = 31/100 (31%), Positives = 42/100 (42%)
Frame = +1
Query: 9 SPTSVPSPAPPSAETNNPNFIRKNLPSPWAQVVRGGSGWDTESQSQSPTGIHQSLPSSSS 68
+P S P P PPS + SP A S T S+ SP+ P SS
Sbjct: 430 APRSTPPPTPPSPPAPT------SASSPPAPARSPASHASTSSRGTSPSSA-PLFPLSSV 588
Query: 69 SSSSSLTTVDQPPPSDDSPKAAVVSSSPKAVVVSSPLPAP 108
+ + L++ P PS SP + SSP + +S PAP
Sbjct: 589 TPPTQLSS-SFPTPSTSSPTSR--GSSPASPPTASSAPAP 699
Score = 29.3 bits (64), Expect = 4.7
Identities = 43/177 (24%), Positives = 56/177 (31%), Gaps = 12/177 (6%)
Frame = +1
Query: 243 SNPPTPPPLPPYPVYQLPPAVSYPNMLPSIPDSSPRDHHRNNNWDARPFVGGSSRRGHFG 302
S P T PPLPP PS SP W +R R
Sbjct: 223 SGP*TTPPLPP---------------RPSATTRSPSSAPATWEWPSR-------RPSSLR 336
Query: 303 SHPRGDGSYHQNSYSSRREHDRGNYANTRDAHAPQPRMPPRGILRPPPPSTAAFLGPQTI 362
+ P S SS R + ++T +P PR P P PPS A P +
Sbjct: 337 TSPTSSSSSTPTPTSSAA---RCSTSSTPPPSSPAPRSTP----PPTPPSPPA---PTSA 486
Query: 363 GPFPAPVAYPDFYYFPTVPLDPFRGMPVFP------------HMPSPATFFPAAESS 407
PAP P + + P+FP P+P+T P + S
Sbjct: 487 SSPPAPARSPASHASTSSRGTSPSSAPLFPLSSVTPPTQLSSSFPTPSTSSPTSRGS 657
>BE661668 similar to GP|10177706|dbj contains similarity to RNA-binding
protein~gene_id:MPL12.3 {Arabidopsis thaliana}, partial
(13%)
Length = 554
Score = 48.1 bits (113), Expect = 1e-05
Identities = 21/47 (44%), Positives = 35/47 (73%), Gaps = 2/47 (4%)
Frame = +1
Query: 412 IVNQIDYYFSDINLANDEFLKS--NMDEEGWVPITLIANFPRVKNLT 456
I+ Q++YYFSD NL D++L ++EG+VP+++IA+F ++K LT
Sbjct: 355 IIKQVEYYFSDENLPTDKYLLGFVKRNKEGFVPVSVIASFRKIKKLT 495
>TC215140 similar to GB|AAL32703.1|17065098|AY062625 nucleotide sugar
epimerase-like protein {Arabidopsis thaliana;} , partial
(91%)
Length = 1885
Score = 47.8 bits (112), Expect = 1e-05
Identities = 69/271 (25%), Positives = 101/271 (36%), Gaps = 39/271 (14%)
Frame = +1
Query: 45 SGWDTESQSQSPTGIHQSLPSS--SSSSSSS--------------------LTTVDQPPP 82
SG T S +P S+P S SSSSSS T PPP
Sbjct: 115 SGDSTAPNSSTPPPNSSSVPLS*SHSSSSSSSPSTTLPSPQTSPPTATSTPTPTSSPPPP 294
Query: 83 SDDSPKAAVVSSSPKAVVVSSP--LPAPMEKNSNT--IASDGGDGGNAGGSKKPAWNKQP 138
++ + P A +SP PAP+ ++ T + S +G + A P
Sbjct: 295 PSPPGRSRSATPPPLAAPTASPSSSPAPLASSAATAPLPSKNAATAFSGSTTSTATTIPP 474
Query: 139 LNGVAVEIGPVMGAESWPALSESAKIPGKLPPESSSSKIAPPAAVDGSPSTSQGP--IIS 196
N + SW K+ PP SS P SP++S P + S
Sbjct: 475 *NAPVRQCSGNTRFSSW-------KVTLTTPPCLRSSSTLSP-----SPTSSTSPLRLAS 618
Query: 197 HSPQRQGSTSN--TKSSSMANNNLPNRPRPMRRISGSNIGP---------CPAQSSLSNP 245
+P R + ++ T +S ++ LPN P P SG + P P+ ++ +NP
Sbjct: 619 ATPCRTPNPTSPPTLLASSTSSRLPNPPIPNPP*SGLHPAPSTASTPKTLSPSSTAPTNP 798
Query: 246 PTPPPLPPYPVYQLPPAVSYPNMLPSIPDSS 276
P P P P + P + PS PDS+
Sbjct: 799 PASTPPPRKPARRSPTPTTTSTASPS-PDSA 888
Score = 42.4 bits (98), Expect = 5e-04
Identities = 76/342 (22%), Positives = 114/342 (33%), Gaps = 17/342 (4%)
Frame = +1
Query: 65 SSSSSSSSSLTTVDQPPPSDDSPKAAVVSSSPKAVVVSSPLPAPMEKNSNTIASDGGDGG 124
++++S S+ PPP+ S + SS + S+ LP+P
Sbjct: 103 TTATSGDSTAPNSSTPPPNSSSVPLS*SHSSSSSSSPSTTLPSPQTSPPT---------- 252
Query: 125 NAGGSKKPAWNKQPLNGVAVEIGPVMGAESWPALSESAKIPGKLPPESSSSKIAPPAAVD 184
A S P + S P P S S+ P AA
Sbjct: 253 ---------------------------ATSTPTPTSSPPPPPSPPGRSRSATPPPLAAPT 351
Query: 185 GSPSTSQGPIISHS-----PQRQGSTSNTKSSSMANNNLPNRPRPMRRISGS-------- 231
SPS+S P+ S + P + +T+ + S++ +P P+R+ SG+
Sbjct: 352 ASPSSSPAPLASSAATAPLPSKNAATAFSGSTTSTATTIPP*NAPVRQCSGNTRFSSWKV 531
Query: 232 --NIGPC-PAQSSLSNPPTPPPLPPYPVYQLPPAVSYPNMLPSIPDSSPRDHHRNNNWDA 288
PC + S+LS PT P P P P++ SS N
Sbjct: 532 TLTTPPCLRSSSTLSPSPTSSTSPLRLASATPCRTPNPTSPPTLLASSTSSRLPNPPIPN 711
Query: 289 RPFVGGSSRRGHFGSHPRGDGSYHQNSYSSRREHDRGNYANTR-DAHAPQPRMPPRGILR 347
P G HP ++ S+ + + A T A P PR P R
Sbjct: 712 PP---------*SGLHPA------PSTASTPKTLSPSSTAPTNPPASTPPPRKPARRSPT 846
Query: 348 PPPPSTAAFLGPQTIGPFPAPVAYPDFYYFPTVPLDPFRGMP 389
P STA+ P AP P + F + P +G P
Sbjct: 847 PTTTSTASPSPDSASSPSTAPGGDPTWLTFSS-PRTSSKGKP 969
Score = 28.9 bits (63), Expect = 6.2
Identities = 41/188 (21%), Positives = 73/188 (38%), Gaps = 14/188 (7%)
Frame = +1
Query: 9 SPTSVPSPAPPSAETNNPNFIRKNLPSPWAQVVRGGSGWDTESQSQSP-------TGIHQ 61
S T PSP ++ + P+P + S T S+ +P +G+H
Sbjct: 562 SSTLSPSPTSSTSPLRLASATPCRTPNPTSPPTLLASS--TSSRLPNPPIPNPP*SGLHP 735
Query: 62 SLPSSSSSSS--SSLTTVDQPPPSDDSPKAAVVSSSPKAVVVSSPLPAPMEKNSNTIASD 119
+ PS++S+ S +T PP+ P SP S+ P+P +S + A
Sbjct: 736 A-PSTASTPKTLSPSSTAPTNPPASTPPPRKPARRSPTPTTTSTASPSPDSASSPSTAPG 912
Query: 120 GGDGGNAGGSKKPAWNKQPLNGVAVEIG---PVMGAESWPALSESAK--IPGKLPPESSS 174
G S + + +P G PV S +L + + P + P +++
Sbjct: 913 GDPTWLTFSSPRTSSKGKPSTCTRRRRGSRWPVTSPTSTTSLKAAWEPWTPPRRAPGAAA 1092
Query: 175 SKIAPPAA 182
+ APP++
Sbjct: 1093RRRAPPSS 1116
>TC225829 weakly similar to UP|Q9C5Q6 (Q9C5Q6) Fasciclin-like
arabinogalactan-protein 2, partial (57%)
Length = 1652
Score = 47.8 bits (112), Expect = 1e-05
Identities = 54/211 (25%), Positives = 75/211 (34%), Gaps = 9/211 (4%)
Frame = +2
Query: 65 SSSSSSSSSLTTVDQPPPSDDSPKAAVVSSSPKAVVVSSPLPAPMEKNSNTIASDGGDGG 124
SSS SSSSS PPS + A +P + ++ P+P +T A
Sbjct: 17 SSSLSSSSSS*PPPPSPPSPPTTSPACWPHTPASPPSTTTSPSPTWPKRSTAARPS---- 184
Query: 125 NAGGSKKPAWNKQ----PLNGVAVEIGPVMGAESWPALSESAKIP-----GKLPPESSSS 175
P+W P + + P S S + P P SS
Sbjct: 185 -------PSWPSTMPPCPPSSTSTSPSPPSKTSSPSTSSSTTSAPKSSTRSTTAPPSSPP 343
Query: 176 KIAPPAAVDGSPSTSQGPIISHSPQRQGSTSNTKSSSMANNNLPNRPRPMRRISGSNIGP 235
PPA P+TS P + P R S T ++ + P+ P R+ P
Sbjct: 344 CSRPPAPPPAPPATSTSP--TSRPARSASPPKTTTAP----STPSTSNPSRKC------P 487
Query: 236 CPAQSSLSNPPTPPPLPPYPVYQLPPAVSYP 266
+ SS S PP PP P +P PP+ P
Sbjct: 488 TTSPSSKSAPPLAPPTPRHPPPPPPPST*SP 580
Score = 43.9 bits (102), Expect = 2e-04
Identities = 52/187 (27%), Positives = 65/187 (33%)
Frame = +2
Query: 67 SSSSSSSLTTVDQPPPSDDSPKAAVVSSSPKAVVVSSPLPAPMEKNSNTIASDGGDGGNA 126
+SSS SS ++ PPPS SP + P P S T
Sbjct: 14 ASSSLSSSSSS*PPPPSPPSPPTTSPACWPHT---------PASPPSTTT---------- 136
Query: 127 GGSKKPAWNKQPLNGVAVEIGPVMGAESWPALSESAKIPGKLPPESSSSKIAPPAAVDGS 186
S P W P A P SWP+ PP S+S+ +PP+ S
Sbjct: 137 --SPSPTW---PKRSTAARPSP-----SWPSTMPPC------PPSSTSTSPSPPSKTS-S 265
Query: 187 PSTSQGPIISHSPQRQGSTSNTKSSSMANNNLPNRPRPMRRISGSNIGPCPAQSSLSNPP 246
PSTS +TS KSS+ + P+ P PC S PP
Sbjct: 266 PSTSSS-----------TTSAPKSSTRSTTAPPSSP------------PC------SRPP 358
Query: 247 TPPPLPP 253
PPP PP
Sbjct: 359 APPPAPP 379
Score = 43.1 bits (100), Expect = 3e-04
Identities = 50/184 (27%), Positives = 69/184 (37%), Gaps = 4/184 (2%)
Frame = +2
Query: 14 PSPAPPSAETNNPNFIRKNLPSPWAQVVRGGSGWDTESQSQSPTGIHQSLPSSSSSSSSS 73
P P+PPS T +P SP + W S + P+ S PS+ S
Sbjct: 53 PPPSPPSPPTTSPACWPHTPASPPSTTTSPSPTWPKRSTAARPS---PSWPSTMPPCPPS 223
Query: 74 LTTVDQPPPSDDS--PKAAVVSSSPKAVVVSSPLPAPMEKNSNTIASDGGDGGNAGGSKK 131
T+ PPS S ++ +S+PK+ S+ P S A
Sbjct: 224 STSTSPSPPSKTSSPSTSSSTTSAPKSSTRSTTAPPSSPPCSRPPAPP---------PAP 376
Query: 132 PAWNKQPLNGVAVEIGP--VMGAESWPALSESAKIPGKLPPESSSSKIAPPAAVDGSPST 189
PA + P + A P A S P+ S ++ K P S SSK APP A P T
Sbjct: 377 PATSTSPTSRPARSASPPKTTTAPSTPSTSNPSR---KCPTTSPSSKSAPPLA----PPT 535
Query: 190 SQGP 193
+ P
Sbjct: 536 PRHP 547
>TC205203 similar to UP|Q39763 (Q39763) Proline-rich cell wall protein,
partial (48%)
Length = 1076
Score = 46.6 bits (109), Expect = 3e-05
Identities = 42/132 (31%), Positives = 56/132 (41%), Gaps = 10/132 (7%)
Frame = +3
Query: 156 PALSESAKIPGKLPPESSSSKIAPPAAVDGSPSTSQGPIISHSPQRQGSTSNTKSSSMAN 215
P+ S + P PP SS P V SP SQ P + +P +S SS +
Sbjct: 276 PSTSPAPPTPQASPPPVQSS----PPPVPSSPPPSQSPPPASTPPPSPLSSPPPSSPPPS 443
Query: 216 NNLPNRPRPMRRISGSN--IGPCPAQSSLSNPP-------TPPPLPPYPVYQLPPAVSYP 266
+ P+ P P S P P+ S+PP TPPP P PPAV P
Sbjct: 444 SPPPSSPPPASPPPSSPPPASPPPSSPPPSSPPPFSPPPATPPPATP------PPAVPPP 605
Query: 267 NMLPSI-PDSSP 277
++ P++ P SSP
Sbjct: 606 SLTPTVTPLSSP 641
Score = 38.9 bits (89), Expect = 0.006
Identities = 47/182 (25%), Positives = 60/182 (32%)
Frame = +3
Query: 15 SPAPPSAETNNPNFIRKNLPSPWAQVVRGGSGWDTESQSQSPTGIHQSLPSSSSSSSSSL 74
SPAPP+ + + P P P + SQSP PS SS S
Sbjct: 285 SPAPPTPQASPPPVQSSPPPVP-----------SSPPPSQSPPPASTPPPSPLSSPPPSS 431
Query: 75 TTVDQPPPSDDSPKAAVVSSSPKAVVVSSPLPAPMEKNSNTIASDGGDGGNAGGSKKPAW 134
PPPS P + SS P A SP P+ + S P +
Sbjct: 432 PPPSSPPPSSPPPASPPPSSPPPA----SPPPS-----------------SPPPSSPPPF 548
Query: 135 NKQPLNGVAVEIGPVMGAESWPALSESAKIPGKLPPESSSSKIAPPAAVDGSPSTSQGPI 194
+ P P + S P P S SK+ PA SPS + GP
Sbjct: 549 SPPPATPPPATPPPAVPPPSLTPTVTPLSSPPVSSPAPSPSKVFAPAL---SPSLAPGPS 719
Query: 195 IS 196
+S
Sbjct: 720 LS 725
Score = 35.0 bits (79), Expect = 0.086
Identities = 33/99 (33%), Positives = 43/99 (43%), Gaps = 4/99 (4%)
Frame = +3
Query: 183 VDGSPSTSQGPIISHSPQR-QGSTSNTKSSSMANNNLPNRPRPMRRISGSNIGPCPAQSS 241
V G + +QGP S +P Q S +SS +P+ P P + ++ P S
Sbjct: 246 VAGVGAQAQGPSTSPAPPTPQASPPPVQSSPPP---VPSSPPPSQSPPPAST---PPPSP 407
Query: 242 LSNPPTPPPLPPYPVYQLPPAVSYPNMLP---SIPDSSP 277
LS+PP P P P PP S P P S P SSP
Sbjct: 408 LSSPPPSSPPPSSPPPSSPPPASPPPSSPPPASPPPSSP 524
>TC226468 weakly similar to UP|O65450 (O65450) Glycine-rich protein, partial
(20%)
Length = 1041
Score = 46.2 bits (108), Expect = 4e-05
Identities = 40/133 (30%), Positives = 59/133 (44%), Gaps = 17/133 (12%)
Frame = -2
Query: 162 AKIPGKLPPESSSSKIAPPAAVDGSPSTS------QGPIISHSPQRQGSTSNTKSSSMAN 215
A +P KLPP + + P SPS+S Q P + SP +TS ++S
Sbjct: 977 ATVPVKLPPTT----LPPTTPTATSPSSSTPLAVTQPPTVVASPP---TTSTPPTTSQPP 819
Query: 216 NNLPNRPRPMRRISGSNIGPC------PAQSSLSNPP-----TPPPLPPYPVYQLPPAVS 264
N+ +P P++ S + P P Q + PP +PPPLPP PV PP +
Sbjct: 818 ANVAPKPAPVKP-SAPKVAPASSPKAPPPQIPIPQPPKPSPVSPPPLPPPPVAS-PPPLP 645
Query: 265 YPNMLPSIPDSSP 277
P + P+ + P
Sbjct: 644 PPKIAPTPAKTPP 606
Score = 43.9 bits (102), Expect = 2e-04
Identities = 61/236 (25%), Positives = 90/236 (37%), Gaps = 8/236 (3%)
Frame = -2
Query: 56 PTGIHQSLPSSSSSSSSSLTTVDQPPPSDDSPKAAVVSSSPKAVVVSSPLPAPMEKNSNT 115
PT + + P+++S SSS+ V QPP VV+S P +S P + +N
Sbjct: 953 PTTLPPTTPTATSPSSSTPLAVTQPP--------TVVASPP----TTSTPPTTSQPPANV 810
Query: 116 IASDGGDGGNAGGSKKPAWNKQPLNGVAVEIGPVMGAESWPALSESAKIPGKLPPESS-- 173
+ KPA P+ A ++ P ++ P +IP PP+ S
Sbjct: 809 -------------APKPA----PVKPSAPKVAPASSPKAPPP-----QIPIPQPPKPSPV 696
Query: 174 SSKIAPPAAVDGSPSTSQGPIISHSPQRQGSTSNTKSSSMANNNLPNRPRPMRRISGSNI 233
S PP V SP P I+ +P + P P P +
Sbjct: 695 SPPPLPPPPV-ASPPPLPPPKIAPTPAKT----------------PPSPAPAKAT----- 582
Query: 234 GPCPAQSSLSNPPTPPPLPPYPVYQLPPA--VSYPNMLPS----IPDSSPRDHHRN 283
P PA + PTP P+PP P PA + P P+ +P +P HHR+
Sbjct: 581 -PAPAPAPTKPAPTPSPVPPPPAATPAPAPVIEVPAPAPAPVIEVPAPAPA-HHRH 420
>TC228914 similar to UP|DAH1_MOUSE (Q9QYB2) Dachshund homolog 1 (Dach1),
partial (3%)
Length = 784
Score = 46.2 bits (108), Expect = 4e-05
Identities = 52/199 (26%), Positives = 74/199 (37%), Gaps = 12/199 (6%)
Frame = +3
Query: 62 SLPSSSSSSSSSLTTVDQPPPSDDSPKAAVVSSSPKAVVVSSPLPAPMEKNSNTIASDGG 121
+LP +S SSS PP S SP SS+P SSP S + G
Sbjct: 123 TLPPASPRSSS-------PPSSPPSPSTTPTSSAPGNAPHSSP------SASKRRPTPSG 263
Query: 122 DGGNAGGSKKPAWNKQPLNGVAVEIGPVMGAESWP--ALSESAKIPGKLPP------ESS 173
A S KP + P +WP +L S P PP SS
Sbjct: 264 PSSAASTSPKP-------TSTSSRAAPSKSPFTWPSASLGTSMSSPASPPPPAPNASTSS 422
Query: 174 SSKIAPPAAVDGSPSTSQ---GPIISHSPQRQGSTSNTKSSSMANNNLPNRPRPM-RRIS 229
++ A PA + +T+ P + P R T+ + S +N P +R
Sbjct: 423 TTSAASPALASSAANTASATTAPSLPSIPSRTTPTTARSTPSSSNPTSSTSPTATPKRTR 602
Query: 230 GSNIGPCPAQSSLSNPPTP 248
S+ P + +S S+PP+P
Sbjct: 603 VSSPTPSSSSTSRSSPPSP 659
Score = 36.2 bits (82), Expect = 0.039
Identities = 54/223 (24%), Positives = 83/223 (37%), Gaps = 2/223 (0%)
Frame = +3
Query: 3 TTQNNHSPTSVPSPAPPSAETNNPNFIRKNLPSPWAQVVRGGSGWDTESQSQSPTGIHQS 62
TT + P S S +PPS+ PSP T S +P S
Sbjct: 111 TTTSTLPPASPRSSSPPSSP-----------PSP-----------STTPTSSAPGNAPHS 224
Query: 63 LPSSSSSSSSSLTTVDQPPPSDDSPKAAVVSSSPKAVVVSSPLPAPMEKNSNTIASDGGD 122
PS+S +P PS S A S+SPK SS AP + ++ G
Sbjct: 225 SPSASKR---------RPTPSGPSSAA---STSPKPTSTSSRA-APSKSPFTWPSASLGT 365
Query: 123 GGNAGGSKKPAWNKQPLNGVAVEIGPVMGAESWPALSESAKIPGKLP--PESSSSKIAPP 180
++ S P + P + + + A + SA LP P ++ A
Sbjct: 366 SMSSPASPPPPAPNASTSSTTSAASPALASSA--ANTASATTAPSLPSIPSRTTPTTARS 539
Query: 181 AAVDGSPSTSQGPIISHSPQRQGSTSNTKSSSMANNNLPNRPR 223
+P++S P + +P+R +S T SSS + + P P+
Sbjct: 540 TPSSSNPTSSTSP--TATPKRTRVSSPTPSSSSTSRSSPPSPK 662
Score = 30.4 bits (67), Expect = 2.1
Identities = 46/192 (23%), Positives = 70/192 (35%), Gaps = 31/192 (16%)
Frame = +3
Query: 170 PESSSSKIAPPAAVDGSPSTSQG--PIISHSPQRQGSTSNTKSSSMANNNLPNRPRPMRR 227
P SSS +PP+ S++ G P S S ++ T + SS+ + + P
Sbjct: 141 PRSSSPPSSPPSPSTTPTSSAPGNAPHSSPSASKRRPTPSGPSSAASTSPKPTSTSSRAA 320
Query: 228 ISGSNIGPCPAQ--SSLSNPPTPPPLPPYPVYQLPPAVSYPNM---------------LP 270
S S A +S+S+P +PPP P + + P + LP
Sbjct: 321 PSKSPFTWPSASLGTSMSSPASPPPPAPNASTSSTTSAASPALASSAANTASATTAPSLP 500
Query: 271 SIPDSSPRDHHRNNNWDARPFVGGS----SRRGHFGSHPRGDGSYHQNSYSSRRE----- 321
SIP + R+ + P S +R S S ++S S +E
Sbjct: 501 SIPSRTTPTTARSTPSSSNPTSSTSPTATPKRTRVSSPTPSSSSTSRSSPPSPKEPTATV 680
Query: 322 ---HDRGNYANT 330
H RG ANT
Sbjct: 681 TVSHTRGEEANT 716
Score = 30.0 bits (66), Expect = 2.8
Identities = 30/101 (29%), Positives = 42/101 (40%), Gaps = 5/101 (4%)
Frame = +3
Query: 9 SPTSVPSPAPPSAETNNPNFIRK-NLPSPWAQVV----RGGSGWDTESQSQSPTGIHQSL 63
S TS SPA S+ N + +LPS ++ R + S SPT +
Sbjct: 420 STTSAASPALASSAANTASATTAPSLPSIPSRTTPTTARSTPSSSNPTSSTSPTATPKRT 599
Query: 64 PSSSSSSSSSLTTVDQPPPSDDSPKAAVVSSSPKAVVVSSP 104
SS + SSS T+ PPS P A V S + ++P
Sbjct: 600 RVSSPTPSSSSTS-RSSPPSPKEPTATVTVSHTRGEEANTP 719
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.309 0.128 0.384
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 29,401,013
Number of Sequences: 63676
Number of extensions: 694615
Number of successful extensions: 18086
Number of sequences better than 10.0: 1106
Number of HSP's better than 10.0 without gapping: 8421
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 13730
length of query: 546
length of database: 12,639,632
effective HSP length: 102
effective length of query: 444
effective length of database: 6,144,680
effective search space: 2728237920
effective search space used: 2728237920
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 61 (28.1 bits)
Medicago: description of AC146790.7


