

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
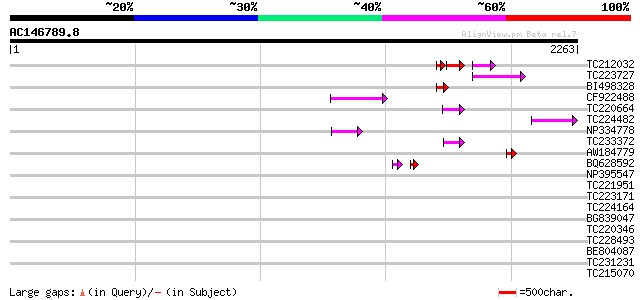
Query= AC146789.8 + phase: 0 /pseudo
(2263 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC212032 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 81 1e-29
TC223727 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 73 2e-12
BI498328 69 2e-11
CF922488 69 2e-11
TC220664 weakly similar to UP|Q84TE0 (Q84TE0) At5g51080, partial... 59 3e-08
TC224482 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 58 5e-08
NP334778 reverse transcriptase [Glycine max] 56 2e-07
TC233372 weakly similar to UP|Q84TE0 (Q84TE0) At5g51080, partial... 52 3e-06
AW184779 46 2e-04
BQ628592 33 3e-04
NP395547 reverse transcriptase [Glycine max] 44 0.001
TC221951 UP|NO75_SOYBN (P08297) Early nodulin 75 precursor (N-75... 42 0.004
TC223171 similar to UP|Q8RW43 (Q8RW43) TCP1 protein, partial (9%) 39 0.020
TC224164 similar to UP|Q9M6E8 (Q9M6E8) 9-cis-epoxycarotenoid dio... 37 0.13
BG839047 35 0.49
TC220346 similar to UP|Q7XII4 (Q7XII4) Remorin-like protein, par... 35 0.49
TC228493 similar to UP|Q9ZTW1 (Q9ZTW1) Lipase (Fragment), partia... 35 0.49
BE804087 34 0.64
TC231231 weakly similar to UP|Q88TJ4 (Q88TJ4) Cell surface prote... 34 0.64
TC215070 similar to UP|Q40336 (Q40336) Proline-rich cell wall pr... 34 0.64
>TC212032 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (3%)
Length = 803
Score = 81.3 bits (199), Expect(3) = 1e-29
Identities = 36/72 (50%), Positives = 54/72 (75%)
Frame = +2
Query: 1742 IEEAIDLRIKNIEIYGDSALVINQIKGKWETLHAGLIPYRDYARRLLTFFNKVELHHIPR 1801
++ AID +K +++YGDSALVI+Q++G+ ET LIPY+ Y + L FF+++ HH+
Sbjct: 206 VQAAIDSNVKLLKVYGDSALVIHQLRGECETRDPNLIPYQAYIKELAGFFDEISFHHVA* 385
Query: 1802 DENQMADALATV 1813
+ENQMADALAT+
Sbjct: 386 EENQMADALATL 421
Score = 56.2 bits (134), Expect(3) = 1e-29
Identities = 23/37 (62%), Positives = 30/37 (80%)
Frame = +1
Query: 1704 NGIGAVLLTPKGAHIPFTARLRFDCTNNIAEYEACIM 1740
+G+GA+L++P IPFT RL FDCTNN+AEYEAC +
Sbjct: 91 HGVGAILVSPDNQCIPFTTRLGFDCTNNMAEYEACAL 201
Score = 33.1 bits (74), Expect(3) = 1e-29
Identities = 31/93 (33%), Positives = 47/93 (50%)
Frame = +1
Query: 1847 Q*TLVS*Y*GVSPNSRVSSWSIQQR*EDSQKII**FLPEWGYLVQKKL*HGLAQMCRQI* 1906
+* LV *Y + RV + +QR ++ ++I L E + VQ+K *HG A + +
Sbjct: 523 R*ALVF*YQAIR*KQRVPTGGFRQRQKEKEEIGSRLLHERKHTVQEKP*HGSATLRER*G 702
Query: 1907 S*FIDT*NTRRIFRNSS*WAHNG*KDIASRLLL 1939
S +F + *WA +G*KD+ LLL
Sbjct: 703 SRKHAGGGP*GLFWYAQ*WARHG*KDLKGGLLL 801
>TC223727 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (9%)
Length = 843
Score = 72.8 bits (177), Expect = 2e-12
Identities = 64/210 (30%), Positives = 103/210 (48%)
Frame = +3
Query: 1847 Q*TLVS*Y*GVSPNSRVSSWSIQQR*EDSQKII**FLPEWGYLVQKKL*HGLAQMCRQI* 1906
+* LV + + R+ + +Q *ED +K+ FL E + V +K H + +C
Sbjct: 129 R*ALVFRHQAICHKQRIPARDCRQ**EDIEKVGGRFLHERKHTV*EKPRHETSAVCGCQG 308
Query: 1907 S*FIDT*NTRRIFRNSS*WAHNG*KDIASRLLLDDNGIRLLQAYQKVSQVSDIRRQDSYA 1966
D + + N+ A G +D SRLLL +G LL +++ Q+S +RR
Sbjct: 309 GKSHDRGSP*GLVWNARQRACYGQEDPKSRLLLAYHGK*LLCPCEEMPQMSSVRR*CQCP 488
Query: 1967 VNYT*FAFLPVAFLYVGH*HDRKD*AQSFKWTPFYLSGY*LLH*VG*GRIICQCDQASSG 2026
+ LP+AFL+VG+ R *AQ +W+ + L H VG G + + + G
Sbjct: 489 TASSECHVLPLAFLHVGNRCHRGH*AQGLEWSSLHPRSDRLFHQVGRGSFLYRRHEGCGG 668
Query: 2027 QIYQESYHLSLWHPQPDHHRQRNKSKQQDD 2056
Q++ E HL +W + D+H QR++ K QDD
Sbjct: 669 QVH*ERDHLPIWFAKEDYHGQRHQPK*QDD 758
>BI498328
Length = 335
Score = 69.3 bits (168), Expect = 2e-11
Identities = 30/48 (62%), Positives = 39/48 (80%)
Frame = +1
Query: 1704 NGIGAVLLTPKGAHIPFTARLRFDCTNNIAEYEACIMGIEEAIDLRIK 1751
+G+GAVL++P IPFTARL FDCTNN+AEYEAC +G++ AID +K
Sbjct: 190 HGVGAVLVSPDDQCIPFTARLGFDCTNNMAEYEACALGVQAAIDFDVK 333
>CF922488
Length = 741
Score = 69.3 bits (168), Expect = 2e-11
Identities = 76/228 (33%), Positives = 105/228 (45%), Gaps = 1/228 (0%)
Frame = +1
Query: 1279 RKMVRSECVLTTVI*TKLAQKTTFLYLILT-CWLTVLPSLKSSHSWMDSPVIIKSRWHPK 1337
++M R EC T I*TK Q+ +FLY I T W+T P +S SWMD VI + R H +
Sbjct: 10 KRMGRCECAWTIEI*TKPVQRISFLYRISTFSWIT-RPVFPNSPSWMDFQVITR*R*HQR 186
Query: 1338 IERKHHLLHLGALFATK*CRSV*SMQELLIRGV*LLFSTI*YIKRLRSMWMI**SSRLLK 1397
I ++ L G A + CR * M G +S * +R RS WM * ++ +
Sbjct: 187 IWKRQLSLLYGEPSAIRLCRLG*RMLGQHTSGPWWHYSRT*CTRR*RSTWMT*S*NQERR 366
Query: 1398 KTMSSIYRRCSSA*GSTNFV*IPTNALLVSDPESF*VSLSARKASK*TLTKSKPSERCLL 1457
+ SI C +T *IP + L +PES L AR+ + T+ K S R
Sbjct: 367 RNTLSICESCLGDYVNTG*D*IPQSVCLR*NPESCSTLLIAREE*RWIRTR*K*SLRWPS 546
Query: 1458 REQKKK*GVFLDD*ITSPGSFLI*LQLVGRYSNYSAKNKVLCGPKIAR 1505
Q+ K V * TS S+ *L L +S A+ + G I +
Sbjct: 547 HIQRSKSKVSWGG*TTS*DSYHS*LPLASLFSYCCARISLSNGTMIVK 690
>TC220664 weakly similar to UP|Q84TE0 (Q84TE0) At5g51080, partial (31%)
Length = 724
Score = 58.5 bits (140), Expect = 3e-08
Identities = 32/85 (37%), Positives = 46/85 (53%)
Frame = +1
Query: 1729 TNNIAEYEACIMGIEEAIDLRIKNIEIYGDSALVINQIKGKWETLHAGLIPYRDYARRLL 1788
TNN AEY A I+G++ A+ I I GDS LV QI G W+ + L + A+ L
Sbjct: 64 TNNAAEYRAMILGMKYALKKGFTGIRIQGDSKLVCMQIDGSWKVKNENLSTLYNVAKELK 243
Query: 1789 TFFNKVELHHIPRDENQMADALATV 1813
F+ ++ H+ R+ N ADA A +
Sbjct: 244 DKFSSFQISHVLRNFNSDADAQANL 318
>TC224482 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (6%)
Length = 669
Score = 57.8 bits (138), Expect = 5e-08
Identities = 53/181 (29%), Positives = 84/181 (46%)
Frame = +3
Query: 2083 SCK*EYKEDCPEDGGDL*RLA*NASLCITWISYFCTHINRGNPFLPGIWHGGSTSCRSRN 2142
S +*EY++D P+D + LA +A + +T + F ++N GN L GIW GG + R+
Sbjct: 3 SGQ*EYQKDYPKDDRVIQGLARDAPIRVTRLPDFSANVNWGNAVLIGIWDGGCVTV*GRS 182
Query: 2143 SIIKSPNGG*LIRS*MGSE*IRPVESD*RKAHDSPMSRTVISKKDETGF*QESSSP*V*R 2202
+IK + +GS + + * A + S ++ K+E QE V
Sbjct: 183 PVIKDLGRIRIKGIRVGSNAL*SAQPH*G*ALNGHESWALVPAKNEECIRQEGMLAQVP* 362
Query: 2203 R*SCAQKDFLFPTRLPRQVGS*L*RSVCC*KSLLRWCHDSTNHGR*RTSTTCEHRCSQEI 2262
CA+++ R++G L R+ CC + R + HG *R + T E C Q I
Sbjct: 363 GRPCAKENVPCCQGPSREMGPELRRAFCCEEGFFRRSPGAYQHGW*RVTFTHEF*CRQAI 542
Query: 2263 L 2263
L
Sbjct: 543 L 545
>NP334778 reverse transcriptase [Glycine max]
Length = 431
Score = 56.2 bits (134), Expect = 2e-07
Identities = 47/123 (38%), Positives = 62/123 (50%), Gaps = 1/123 (0%)
Frame = +1
Query: 1286 CVLTTVI*TKLAQKTTFLYLILT-CWLTVLPSLKSSHSWMDSPVIIKSRWHPKIERKHHL 1344
CV T I*T+ Q+ FLY T WLT P L S SWMD VII+ RWH +I ++
Sbjct: 7 CVWTIEI*TEPVQRIIFLYRTSTFSWLT-WPVLPYSLSWMDFRVIIR*RWHQRIWKRQLS 183
Query: 1345 LHLGALFATK*CRSV*SMQELLIRGV*LLFSTI*YIKRLRSMWMI**SSRLLKKTMSSIY 1404
L G A +*CR * + G +S * +R R MWM * ++ ++ SI
Sbjct: 184 LLYGEPSAIR*CRLG*RILGQPTIGPWWHYSRT*CTRRSRPMWMK*SRNQEWRRNT*SIC 363
Query: 1405 RRC 1407
+ C
Sbjct: 364 KIC 372
>TC233372 weakly similar to UP|Q84TE0 (Q84TE0) At5g51080, partial (12%)
Length = 561
Score = 52.0 bits (123), Expect = 3e-06
Identities = 29/81 (35%), Positives = 42/81 (51%)
Frame = +1
Query: 1733 AEYEACIMGIEEAIDLRIKNIEIYGDSALVINQIKGKWETLHAGLIPYRDYARRLLTFFN 1792
AEY + I+G+ K I + GDS LV NQI+G W+ + + A+ L F
Sbjct: 1 AEYRSLILGLXHXXKKGYKXIIVQGDSLLVCNQIQGLWKIKNQNMGTLCXEAKELKDKFL 180
Query: 1793 KVELHHIPRDENQMADALATV 1813
++ HIPR+ N ADA A +
Sbjct: 181 SFKISHIPREYNSEADAQANL 243
>AW184779
Length = 432
Score = 45.8 bits (107), Expect = 2e-04
Identities = 22/41 (53%), Positives = 25/41 (60%)
Frame = +2
Query: 1981 YVGH*HDRKD*AQSFKWTPFYLSGY*LLH*VG*GRIICQCD 2021
YVGH DR *AQ FKWT + S *LLH +G +C CD
Sbjct: 290 YVGHRRDRSH*AQGFKWTSLHFSRN*LLHQMGRSSFVC*CD 412
>BQ628592
Length = 423
Score = 33.1 bits (74), Expect(2) = 3e-04
Identities = 19/40 (47%), Positives = 24/40 (59%)
Frame = -2
Query: 1526 KEDL*SCT*QC*KIPWVVCSDNKMKPEEKSTPFII*ARSL 1565
+EDL SCT* C* W C + +++ PF I*ARSL
Sbjct: 320 QEDLFSCT*LC*TSLWDACWFSTTTLGKRNKPFTI*ARSL 201
Score = 31.2 bits (69), Expect(2) = 3e-04
Identities = 17/34 (50%), Positives = 21/34 (61%)
Frame = -1
Query: 1598 RGWYLKWIQSSTYLRSPL*QEGLHGGRCYYLNMI 1631
RG + KWI +T LRS ++ GGR YYLN I
Sbjct: 105 RGLFPKWIL*NTSLRSRPSRDESLGGRYYYLNSI 4
>NP395547 reverse transcriptase [Glycine max]
Length = 762
Score = 43.5 bits (101), Expect = 0.001
Identities = 43/133 (32%), Positives = 59/133 (44%), Gaps = 3/133 (2%)
Frame = +2
Query: 1278 QRKMVRSECVLTTVI*TKLAQKTTFLYLILTCWLTVLPSLK---SSHSWMDSPVIIKSRW 1334
Q + ECVL K +KT + W+ L L+ S+ SW D+ V I+ +W
Sbjct: 143 QEESPDGECVLIIGSSMKPQEKTITHF---PSWIKCLRDLQGNPSTVSWTDTQVTIRLQW 313
Query: 1335 HPKIERKHHLLHLGALFATK*CRSV*SMQELLIRGV*LLFSTI*YIKRLRSMWMI**SSR 1394
+I++K L L CRSV M LL R V* F L+S+WMI S
Sbjct: 314 ILRIKKKQLLHVLSVFLLIAACRSVYVMPLLLFRDV*WQFLMTW*RNVLKSLWMIFRSLV 493
Query: 1395 LLKKTMSSIYRRC 1407
L + I R+C
Sbjct: 494 HLLEIA*QI*RKC 532
>TC221951 UP|NO75_SOYBN (P08297) Early nodulin 75 precursor (N-75) (NGM-75),
complete
Length = 1179
Score = 41.6 bits (96), Expect = 0.004
Identities = 26/102 (25%), Positives = 45/102 (43%)
Frame = +2
Query: 423 HNILNSHKILHHKILNNHTNNNHTNNSHTNNTLNKISNNNLINNAHNNLDHQECRLTQYQ 482
H + + H+ ++H + N+H N NH SH NT N+L+ N H N +H Q
Sbjct: 383 HLMRSHHQSMNHLMRNHHQNTNHLMRSHHQNT------NHLMRNHHQNTNHLMRSHHQST 544
Query: 483 SPMQNYFPAYSRKTLSKLEQLLLSQKNYHLGIDLTRLVTSMK 524
+ + + S S + + +N+H +L + MK
Sbjct: 545 NHLMRSHQSTSHLMRSHHQSINHLMRNHHQNTNLLKKSHHMK 670
Score = 33.5 bits (75), Expect = 1.1
Identities = 20/70 (28%), Positives = 33/70 (46%)
Frame = +2
Query: 404 QITILKYRNILKYRNILKYHNILNSHKILHHKILNNHTNNNHTNNSHTNNTLNKISNNNL 463
Q T L RN + N+L ++ + H + ++ +H + NH SH S N+L
Sbjct: 260 QNTYLLMRNRHQNTNLLMRNHPMRIHHRSTNHLMRSHQSTNHLMRSHHQ------SMNHL 421
Query: 464 INNAHNNLDH 473
+ N H N +H
Sbjct: 422 MRNHHQNTNH 451
Score = 31.2 bits (69), Expect = 5.4
Identities = 16/65 (24%), Positives = 33/65 (50%)
Frame = +2
Query: 423 HNILNSHKILHHKILNNHTNNNHTNNSHTNNTLNKISNNNLINNAHNNLDHQECRLTQYQ 482
++++ SH+ H + ++H + NH +H NT NL+ +H+ +H++ +
Sbjct: 545 NHLMRSHQSTSHLMRSHHQSINHLMRNHHQNT-------NLLKKSHHMKNHRQNTNLLMK 703
Query: 483 SPMQN 487
S QN
Sbjct: 704 SHHQN 718
>TC223171 similar to UP|Q8RW43 (Q8RW43) TCP1 protein, partial (9%)
Length = 411
Score = 39.3 bits (90), Expect = 0.020
Identities = 22/55 (40%), Positives = 30/55 (54%), Gaps = 7/55 (12%)
Frame = +3
Query: 419 ILKYHNILNSHKIL-------HHKILNNHTNNNHTNNSHTNNTLNKISNNNLINN 466
IL +HN N+ +L + NN++NNNH NN+H NN N NNN +N
Sbjct: 237 ILLHHNHNNNEHVLFAGTAFDNMVAWNNNSNNNHNNNNHNNNNNN---NNNTASN 392
Score = 31.2 bits (69), Expect = 5.4
Identities = 18/58 (31%), Positives = 26/58 (44%), Gaps = 12/58 (20%)
Frame = +3
Query: 431 ILHHKILNNHTNNNHTN------------NSHTNNTLNKISNNNLINNAHNNLDHQEC 476
+LHH NH NN H N+++NN N ++NN NN +N + C
Sbjct: 240 LLHH----NHNNNEHVLFAGTAFDNMVAWNNNSNNNHNNNNHNNNNNNNNNTASNDTC 401
>TC224164 similar to UP|Q9M6E8 (Q9M6E8) 9-cis-epoxycarotenoid dioxygenase,
partial (22%)
Length = 557
Score = 36.6 bits (83), Expect = 0.13
Identities = 23/76 (30%), Positives = 42/76 (55%), Gaps = 4/76 (5%)
Frame = +1
Query: 391 LPPSHPPLTLFN-KQITILKYRNILKYRNILKYHNILNSHKILHHKILNNHTNNNHTNNS 449
L +H P + +N K+ L+++ ++ L+ H++ +S+ L K+ N N+NH NN+
Sbjct: 97 LDQNHTPFSSYNLKEKVSLQFQ----FQFQLQRHHMFSSNTPLPQKVPTNLHNHNHNNNN 264
Query: 450 HT---NNTLNKISNNN 462
H+ N T N+ S N
Sbjct: 265 HSSKRNQTSNRFSLRN 312
>BG839047
Length = 610
Score = 34.7 bits (78), Expect = 0.49
Identities = 16/36 (44%), Positives = 25/36 (69%), Gaps = 2/36 (5%)
Frame = -2
Query: 424 NILN--SHKILHHKILNNHTNNNHTNNSHTNNTLNK 457
NI+N H I ++K+ NN+ N+N +NS++N LNK
Sbjct: 573 NIINHLKHNI*YNKLHNNNNNSNSNSNSNSNKRLNK 466
>TC220346 similar to UP|Q7XII4 (Q7XII4) Remorin-like protein, partial (24%)
Length = 909
Score = 34.7 bits (78), Expect = 0.49
Identities = 16/35 (45%), Positives = 22/35 (62%)
Frame = +3
Query: 437 LNNHTNNNHTNNSHTNNTLNKISNNNLINNAHNNL 471
LN N NS+TN+ N +NNN+ NN++NNL
Sbjct: 546 LNTMHGNEEGVNSYTNSNNNYNNNNNINNNSYNNL 650
>TC228493 similar to UP|Q9ZTW1 (Q9ZTW1) Lipase (Fragment), partial (51%)
Length = 1219
Score = 34.7 bits (78), Expect = 0.49
Identities = 23/62 (37%), Positives = 29/62 (46%), Gaps = 4/62 (6%)
Frame = -2
Query: 16 HKRKHWHK----RKLKLRP*QKRRPDHKHLHLLLSELRPRHLLLGHYVLTLQRSPLRNVP 71
H + WH +KL Q+ P H HL LLL + PRH H LQ P+ VP
Sbjct: 822 HPPRPWHAIFLLQKLTRLIHQRHTPLHPHLKLLLLPIPPRH----HMQHRLQVMPITRVP 655
Query: 72 RL 73
R+
Sbjct: 654 RV 649
>BE804087
Length = 160
Score = 34.3 bits (77), Expect = 0.64
Identities = 20/51 (39%), Positives = 27/51 (52%)
Frame = +1
Query: 2083 SCK*EYKEDCPEDGGDL*RLA*NASLCITWISYFCTHINRGNPFLPGIWHG 2133
S +*EY E ED + RLA*+ C ++S T+ GN L G+W G
Sbjct: 4 SSQ*EY*EKYLEDDSVVQRLA*DVFFCPAYVSNLGTNFYWGNTLLLGLWEG 156
>TC231231 weakly similar to UP|Q88TJ4 (Q88TJ4) Cell surface protein, partial
(10%)
Length = 1077
Score = 34.3 bits (77), Expect = 0.64
Identities = 23/83 (27%), Positives = 41/83 (48%), Gaps = 6/83 (7%)
Frame = -1
Query: 392 PPSHPPLTLFNKQITILKYRNI----LKYRNILKYHNILNSHKILHHKILNNHTNNNHTN 447
P S+P N Q+T+ LK + +H+++N + ++H+ N H+ N+H+
Sbjct: 807 PNSYPLTQPNNYQLTLHNLHPS*CFHLK*KGHYCHHHLMN-RRSMNHR*TNRHSMNHHSK 631
Query: 448 NSHTNNT--LNKISNNNLINNAH 468
N H N +N+ S N + N H
Sbjct: 630 NLHWTNRHWMNRHSTNRHLMNRH 562
>TC215070 similar to UP|Q40336 (Q40336) Proline-rich cell wall protein,
partial (82%)
Length = 1597
Score = 34.3 bits (77), Expect = 0.64
Identities = 27/103 (26%), Positives = 48/103 (46%), Gaps = 6/103 (5%)
Frame = +3
Query: 387 PINTLPPSHPPLTLFNKQITILK--YRNILKYRNILKYHNILNSHKI--LHHKILNNHT- 441
P+ TLP LT N Q + +N+++ + H + +H++ LHH + NH
Sbjct: 222 PMYTLPMFQNLLTTLNHQFIHIHPMSQNLIQSLQCIPIHLMSQNHQLCQLHHHMFQNHQL 401
Query: 442 -NNNHTNNSHTNNTLNKISNNNLINNAHNNLDHQECRLTQYQS 483
+ N + IS N+ + + H +L+HQ C+L + S
Sbjct: 402 *TLPMSQNRLLCQLHHLISLNHQL*HHHMSLNHQWCQLHHHMS 530
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.357 0.157 0.557
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 105,393,543
Number of Sequences: 63676
Number of extensions: 1586110
Number of successful extensions: 21282
Number of sequences better than 10.0: 67
Number of HSP's better than 10.0 without gapping: 11005
Number of HSP's successfully gapped in prelim test: 861
Number of HSP's that attempted gapping in prelim test: 8173
Number of HSP's gapped (non-prelim): 14003
length of query: 2263
length of database: 12,639,632
effective HSP length: 112
effective length of query: 2151
effective length of database: 5,507,920
effective search space: 11847535920
effective search space used: 11847535920
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.7 bits)
S2: 67 (30.4 bits)
Medicago: description of AC146789.8


