

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146789.1 + phase: 0
(431 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
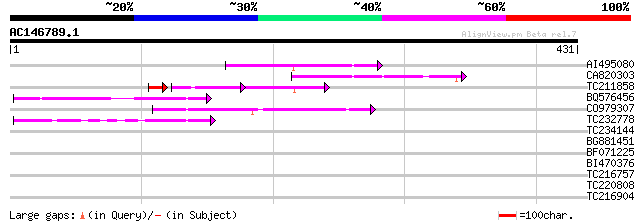
Score E
Sequences producing significant alignments: (bits) Value
AI495080 96 4e-20
CA820303 84 1e-16
TC211858 59 1e-15
BQ576456 69 4e-12
CO979307 55 5e-08
TC232778 similar to UP|Q9LNG5 (Q9LNG5) F21D18.16, partial (5%) 54 1e-07
TC234144 40 0.002
BG881451 weakly similar to PIR|D96521|D96 protein F21D18.16 [imp... 39 0.003
BF071225 weakly similar to PIR|D96521|D96 protein F21D18.16 [imp... 37 0.013
BI470376 weakly similar to GP|14090338|dbj P0638D12.5 {Oryza sat... 31 1.2
TC216757 similar to PIR|T05768|T05768 subtilisin-like proteinase... 30 1.6
TC220808 weakly similar to UP|Q9SK32 (Q9SK32) Expressed protein,... 30 1.6
TC216904 UP|P93166 (P93166) SCOF-1, complete 28 8.0
>AI495080
Length = 407
Score = 95.5 bits (236), Expect = 4e-20
Identities = 52/122 (42%), Positives = 68/122 (55%), Gaps = 3/122 (2%)
Frame = +2
Query: 165 RALGGSVTLLTTWFLKHFPGFFSVDLNTDYLEKYPVAARWKLEKGHGEGI---TYRSLLD 221
R L G +TLL W +HFP + DY E P A RW K + I +YR L+
Sbjct: 2 RQLSGYITLLQCWIYEHFPSVAESTAD*DYDEASPRACRWIATKKTVKSIRTPSYRERLN 181
Query: 222 RIQLDDVCWRPYEEHREIQDFEEVFWYSGWIMCGVRRVYRHLPERVLRQYGYVQTIPRHP 281
R+++ DVCW Y EHRE+QDF YSG + G+ VY + PE+V+RQ+GY IP P
Sbjct: 182 RLRISDVCWILYGEHREVQDFHVRSCYSGLLHWGLVAVY-YRPEKVVRQFGYTPAIPAPP 358
Query: 282 TD 283
D
Sbjct: 359 VD 364
>CA820303
Length = 421
Score = 84.0 bits (206), Expect = 1e-16
Identities = 53/139 (38%), Positives = 73/139 (52%), Gaps = 6/139 (4%)
Frame = -1
Query: 215 TYRSLLDRIQLDDVCWRPYEEHREIQDFEEVFWYSGWIMCGVRRVYRHLPERVLRQYGYV 274
+YR LDR+++ DVCW PY EHRE+QDF YSG + G VY + ERV+RQ+GY
Sbjct: 400 SYRERLDRLRIPDVCWIPYGEHREVQDFHVRSCYSGLLCWGPVVVY-YRSERVVRQFGYT 224
Query: 275 QTIPRHPTDVRDLPPPSIVQMFIDFRTHTLKADARGVQAGEDTRRVADGYVLWYTRVSHP 334
QTIP D + I ++ + H + A V G + Y+ W+ R SHP
Sbjct: 223 QTIPASTVD-SWVSYDDIHDRWMHYEDHIVPAGEVCVVPG----ACSSDYIDWFFRTSHP 59
Query: 335 QILP-----PIP-GDLPRP 347
+ P P+P G P+P
Sbjct: 58 FMTPGHAVDPLPHGHAPQP 2
>TC211858
Length = 524
Score = 58.5 bits (140), Expect(3) = 1e-15
Identities = 28/70 (40%), Positives = 38/70 (54%), Gaps = 3/70 (4%)
Frame = -3
Query: 177 WFLKHFPGFFSVDLNTDYLEKYPVAARWKLEKGHGEGIT---YRSLLDRIQLDDVCWRPY 233
W +HFP ++ Y E P A+RW K H +GIT YR+ D + + DV W PY
Sbjct: 231 WIYEHFPSVHQCVIDDTY*ETSPRASRWLTSKAHMKGITGAPYRARCDALTVIDVSWLPY 52
Query: 234 EEHREIQDFE 243
EHR ++ FE
Sbjct: 51 TEHRGVRAFE 22
Score = 35.4 bits (80), Expect(3) = 1e-15
Identities = 19/57 (33%), Positives = 31/57 (54%), Gaps = 1/57 (1%)
Frame = -1
Query: 124 IELIYLTTMAD-GYAGMRNYSWGAMTLAYLYGELADACRPGHRALGGSVTLLTTWFL 179
+ ++YL D G +G Y+WG L ++Y +L +A R R + G +TLL F+
Sbjct: 470 VHVVYLDAFRDLGQSG--GYAWGVAALVHMYNQLDEASRTTTRQIVGYLTLLQVNFV 306
Score = 26.2 bits (56), Expect(3) = 1e-15
Identities = 10/15 (66%), Positives = 13/15 (86%)
Frame = -2
Query: 106 YLLYLVGCLLFGDRS 120
YLL+LVGC LF ++S
Sbjct: 523 YLLHLVGCTLFANKS 479
>BQ576456
Length = 424
Score = 68.9 bits (167), Expect = 4e-12
Identities = 47/150 (31%), Positives = 68/150 (45%)
Frame = +1
Query: 4 GEMTVTLDDVACLMHLPIEGRMLAHGKKMPKHEGAALLMTYLGVAQHEAEKICNQEYGGY 63
GE+T+TLDD+ACL+HLPI G L + + E LLM L V++ EA + + Y
Sbjct: 31 GELTITLDDMACLLHLPITG-ALHRFEPLGVDEAVLLLMELLEVSREEARAKIVRAHRAY 207
Query: 64 ISYPRLRDFYTSYLGRANVLVGTEDPEEVEELERVRTYCVRCYLLYLVGCLLFGDRSNKR 123
+ LR+ Y S R VR YL +LV C LF ++S
Sbjct: 208 VRLSWLREVYQSRC-----------------QARCWIVAVRAYLFHLVDCTLFANKSATH 336
Query: 124 IELIYLTTMADGYAGMRNYSWGAMTLAYLY 153
+ +++L D Y WG L ++Y
Sbjct: 337 VHVVHLKGF*D-LC*SGGYGWGVAALVHMY 423
>CO979307
Length = 553
Score = 55.5 bits (132), Expect = 5e-08
Identities = 43/172 (25%), Positives = 80/172 (46%), Gaps = 2/172 (1%)
Frame = -3
Query: 109 YLVGCLLFGDRSNKRIELIYLTTMADGYAGMRNYSWGAMTLAYLYGELADACRPGHRALG 168
+L+ ++F D+S + + YL +++ G L YLY L+ ++ G
Sbjct: 551 HLISSMIFADKSLAHVHVAYL*YLSN-LTACHK*V*GVAALVYLYDHLSYVSLDDNKQCG 375
Query: 169 GSVTLLTTWFLKHFP--GFFSVDLNTDYLEKYPVAARWKLEKGHGEGITYRSLLDRIQLD 226
+TLL +H P G+ + + DY + P+ RW G + + ++ I ++
Sbjct: 374 DYMTLLMA*VFEHLPSVGYSNPE---DYSDGDPLNTRWNPLGGTRHALLVKERMNNIHMN 204
Query: 227 DVCWRPYEEHREIQDFEEVFWYSGWIMCGVRRVYRHLPERVLRQYGYVQTIP 278
PYE ++ ++ F E G+ C + + +L +RV RQ+ +VQTIP
Sbjct: 203 VFI*CPYE*NKLVRPFVEQTL*FGYFKCRI-DMRSYLLKRVRRQFCHVQTIP 51
>TC232778 similar to UP|Q9LNG5 (Q9LNG5) F21D18.16, partial (5%)
Length = 746
Score = 54.3 bits (129), Expect = 1e-07
Identities = 46/153 (30%), Positives = 69/153 (45%)
Frame = -1
Query: 4 GEMTVTLDDVACLMHLPIEGRMLAHGKKMPKHEGAALLMTYLGVAQHEAEKICNQEYGGY 63
GE T+TL DV L+ L +G L + + A L LGV E E G
Sbjct: 422 GEATITLQDVLILLGLRTDGAPLIGSTNL---DWADLCEELLGVRPQEGEI-----EGSV 267
Query: 64 ISYPRLRDFYTSYLGRANVLVGTEDPEEVEELERVRTYCVRCYLLYLVGCLLFGDRSNKR 123
+ L Y N+ D VE+L+R +R ++L +G ++F ++S+ R
Sbjct: 266 VKLSWL----AHYFSHINI-----DEGNVEQLQRF----IRAWILRFIGGVIFVNKSSSR 126
Query: 124 IELIYLTTMADGYAGMRNYSWGAMTLAYLYGEL 156
+ L YL + D + Y+WG LAYLY E+
Sbjct: 125 VSLRYLQFLRD-FE*CSTYAWGPAMLAYLYREM 30
>TC234144
Length = 468
Score = 40.0 bits (92), Expect = 0.002
Identities = 16/39 (41%), Positives = 23/39 (58%)
Frame = +3
Query: 217 RSLLDRIQLDDVCWRPYEEHREIQDFEEVFWYSGWIMCG 255
R LD + D+ W PY HRE + F ++ +SG+I CG
Sbjct: 6 RPALDML*TKDILWMPYTVHREHRPFHDISLFSGYIRCG 122
>BG881451 weakly similar to PIR|D96521|D96 protein F21D18.16 [imported] -
Arabidopsis thaliana, partial (1%)
Length = 421
Score = 39.3 bits (90), Expect = 0.003
Identities = 24/70 (34%), Positives = 37/70 (52%), Gaps = 5/70 (7%)
Frame = +3
Query: 214 ITYRSLLDRIQLDDVCWRPYE--EHREIQDFEE---VFWYSGWIMCGVRRVYRHLPERVL 268
+ YR LD ++ DV W PY + I ++ + + S ++ + RHLP R L
Sbjct: 6 VFYRKSLDSLKPCDVEWLPYRNMDSMVIPEYIKSTLILGRSKTMLICFDKAERHLPNRCL 185
Query: 269 RQYGYVQTIP 278
RQYG +Q+IP
Sbjct: 186 RQYGMLQSIP 215
>BF071225 weakly similar to PIR|D96521|D96 protein F21D18.16 [imported] -
Arabidopsis thaliana, partial (4%)
Length = 359
Score = 37.4 bits (85), Expect = 0.013
Identities = 25/84 (29%), Positives = 39/84 (45%), Gaps = 5/84 (5%)
Frame = +2
Query: 209 GHGEGITYRSLLDRIQLDDVCWRPYEEH-----REIQDFEEVFWYSGWIMCGVRRVYRHL 263
G+ + +R LD ++ + W PY +I V W++ + V H
Sbjct: 29 GNDDLRVFRRKLDLMKRHEFVWEPYTPTVMAALPQICVVGSVVWFAVVPLICFHVVEWHQ 208
Query: 264 PERVLRQYGYVQTIPRHPTDVRDL 287
P+RVLRQYG Q IP P+ ++L
Sbjct: 209 PDRVLRQYGLQQPIPGCPSQPQNL 280
>BI470376 weakly similar to GP|14090338|dbj P0638D12.5 {Oryza sativa
(japonica cultivar-group)}, partial (5%)
Length = 429
Score = 30.8 bits (68), Expect = 1.2
Identities = 12/23 (52%), Positives = 18/23 (78%)
Frame = -1
Query: 4 GEMTVTLDDVACLMHLPIEGRML 26
GE T+TL DV+ L+ +P++GR L
Sbjct: 273 GEATITLQDVSVLLGIPVDGRPL 205
>TC216757 similar to PIR|T05768|T05768 subtilisin-like proteinase -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(20%)
Length = 1126
Score = 30.4 bits (67), Expect = 1.6
Identities = 13/24 (54%), Positives = 14/24 (58%)
Frame = -1
Query: 405 REQIAPILTRRRAQRPRRRHHHQD 428
R IAP+L R R PRR H H D
Sbjct: 517 RHHIAPLLHRLREHEPRRLHRHAD 446
>TC220808 weakly similar to UP|Q9SK32 (Q9SK32) Expressed protein, partial
(9%)
Length = 869
Score = 30.4 bits (67), Expect = 1.6
Identities = 44/176 (25%), Positives = 69/176 (39%), Gaps = 10/176 (5%)
Frame = +1
Query: 261 RHLPERVLRQYGYVQTIPRHPTDVRDLPPPS-IVQMFIDFRTHTLKA----DARGVQAGE 315
RHLP+R LRQ+ QTIP+ DV S IV +D A R V E
Sbjct: 58 RHLPDRCLRQFAMHQTIPK---DVERWERKSRIVDHGVDLMGKMDLALKEWSERWVHVVE 228
Query: 316 DTRRVADG-YVLWYTRVSHPQILPPIPGDLPRPANEEQIIAEQWQR--YEARSSPDTYDM 372
V +G Y+ WY +++ I + ++QR R + D+
Sbjct: 229 GGDIVDEGEYMQWYQKITRKYI-----------GRVTSSLESEYQRTVTAMREIANIADI 375
Query: 373 VS--GAVAYADAQLGQEEVMSMTPQQWYEAMTHMREQIAPILTRRRAQRPRRRHHH 426
VS G +Y E++ ++ +T E+I +++ R RR+ H
Sbjct: 376 VSAEGLDSY------NRELLDEVKNIVHKCLTEQFEEIPNEKVKKKGNRKRRQKDH 525
>TC216904 UP|P93166 (P93166) SCOF-1, complete
Length = 1114
Score = 28.1 bits (61), Expect = 8.0
Identities = 26/97 (26%), Positives = 38/97 (38%), Gaps = 2/97 (2%)
Frame = +1
Query: 332 SHPQILPPIPGDLPRPANEEQIIAEQWQRYEARSSPDTYDMVSGAVAYADAQ-LGQEEVM 390
S P +P LP A +R+ R P + A A AD + Q ++
Sbjct: 307 SRPSF*RRVPRPLPHHARS--------RRHHHRQQPPRQPSAATATATADTRSFHQAQLQ 462
Query: 391 SMTPQQWYEAMTHMR-EQIAPILTRRRAQRPRRRHHH 426
+ +Q + R Q TRRR +RP +HHH
Sbjct: 463 MLRLRQELPLLPSARWTQGQSPETRRRRRRPTPQHHH 573
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.323 0.139 0.442
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,604,363
Number of Sequences: 63676
Number of extensions: 321829
Number of successful extensions: 1860
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 1808
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1844
length of query: 431
length of database: 12,639,632
effective HSP length: 100
effective length of query: 331
effective length of database: 6,272,032
effective search space: 2076042592
effective search space used: 2076042592
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 60 (27.7 bits)
Medicago: description of AC146789.1


