

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146784.13 + phase: 0 /pseudo
(36 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
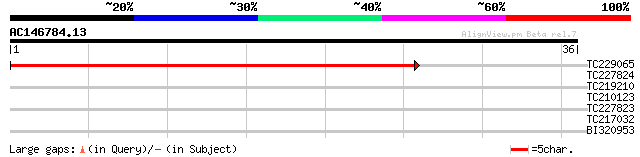
TC229065 similar to UP|Q9LZV3 (Q9LZV3) (1-4)-beta-mannan endohyd... 50 2e-07
TC227824 similar to UP|Q9FJZ3 (Q9FJZ3) Mannan endo-1,4-beta-mann... 33 0.034
TC219210 similar to UP|Q9SKU9 (Q9SKU9) (1-4)-beta-mannan endohyd... 33 0.045
TC210123 similar to UP|Q9SKU9 (Q9SKU9) (1-4)-beta-mannan endohyd... 31 0.13
TC227823 homologue to UP|Q9AT07 (Q9AT07) ADP-glucose pyrophospho... 31 0.13
TC217032 weakly similar to UP|Q9FJZ3 (Q9FJZ3) Mannan endo-1,4-be... 31 0.17
BI320953 27 2.4
>TC229065 similar to UP|Q9LZV3 (Q9LZV3) (1-4)-beta-mannan endohydrolase-like
protein (At5g01930), partial (45%)
Length = 1674
Score = 50.4 bits (119), Expect = 2e-07
Identities = 22/26 (84%), Positives = 25/26 (95%)
Frame = +1
Query: 1 MDAHIEDTEKYLGMPVVFSEFGVSSK 26
M+AHIED EKYLGMPVVF+EFGVS+K
Sbjct: 286 MEAHIEDAEKYLGMPVVFAEFGVSAK 363
>TC227824 similar to UP|Q9FJZ3 (Q9FJZ3) Mannan endo-1,4-beta-mannosidase
(At5g66460), partial (16%)
Length = 924
Score = 33.1 bits (74), Expect = 0.034
Identities = 12/26 (46%), Positives = 20/26 (76%)
Frame = +3
Query: 1 MDAHIEDTEKYLGMPVVFSEFGVSSK 26
+D HI+D + L P++F+EFG+S+K
Sbjct: 132 LDEHIQDAQNTLHKPLLFAEFGISTK 209
>TC219210 similar to UP|Q9SKU9 (Q9SKU9) (1-4)-beta-mannan endohydrolase,
partial (56%)
Length = 1116
Score = 32.7 bits (73), Expect = 0.045
Identities = 14/24 (58%), Positives = 19/24 (78%)
Frame = +1
Query: 1 MDAHIEDTEKYLGMPVVFSEFGVS 24
M +HIED +K L PV+FSE+G+S
Sbjct: 565 MLSHIEDGDKVLNKPVLFSEYGLS 636
>TC210123 similar to UP|Q9SKU9 (Q9SKU9) (1-4)-beta-mannan endohydrolase,
partial (19%)
Length = 640
Score = 31.2 bits (69), Expect = 0.13
Identities = 13/22 (59%), Positives = 18/22 (81%)
Frame = +2
Query: 3 AHIEDTEKYLGMPVVFSEFGVS 24
+HIED ++ L PV+FSEFG+S
Sbjct: 2 SHIEDGDEVLKKPVLFSEFGLS 67
>TC227823 homologue to UP|Q9AT07 (Q9AT07) ADP-glucose pyrophosphorylase large
subunit CagpL2, partial (25%)
Length = 1272
Score = 31.2 bits (69), Expect = 0.13
Identities = 11/26 (42%), Positives = 20/26 (76%)
Frame = +3
Query: 1 MDAHIEDTEKYLGMPVVFSEFGVSSK 26
++ HI+D + L P++F+EFG+S+K
Sbjct: 243 LNEHIQDAQNTLHKPLLFAEFGISTK 320
>TC217032 weakly similar to UP|Q9FJZ3 (Q9FJZ3) Mannan endo-1,4-beta-mannosidase
(At5g66460), partial (75%)
Length = 1606
Score = 30.8 bits (68), Expect = 0.17
Identities = 12/32 (37%), Positives = 18/32 (55%)
Frame = +2
Query: 1 MDAHIEDTEKYLGMPVVFSEFGVSSKDLATTQ 32
+ HI+D + LG P+V EFG S K + +
Sbjct: 1040 LQTHIQDAKNVLGKPIVVGEFGKSLKSYSVVE 1135
>BI320953
Length = 133
Score = 26.9 bits (58), Expect = 2.4
Identities = 10/22 (45%), Positives = 14/22 (63%)
Frame = +2
Query: 1 MDAHIEDTEKYLGMPVVFSEFG 22
+ HI+D + LG P+V EFG
Sbjct: 65 LQTHIQDAKIVLGKPIVVGEFG 130
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.314 0.130 0.358
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,018,900
Number of Sequences: 63676
Number of extensions: 4730
Number of successful extensions: 18
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 18
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 18
length of query: 36
length of database: 12,639,632
effective HSP length: 12
effective length of query: 24
effective length of database: 11,875,520
effective search space: 285012480
effective search space used: 285012480
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 53 (25.0 bits)
Medicago: description of AC146784.13


