

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146784.10 + phase: 0
(168 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
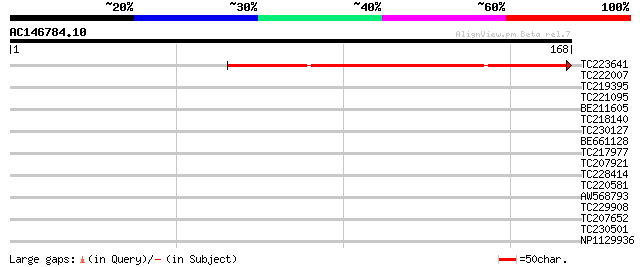
Score E
Sequences producing significant alignments: (bits) Value
TC223641 160 3e-40
TC222007 UP|Q9LKZ5 (Q9LKZ5) Receptor-like protein kinase 2, comp... 28 1.6
TC219395 UP|Q9LKZ4 (Q9LKZ4) Receptor-like protein kinase 3, part... 28 2.1
TC221095 28 2.1
BE211605 homologue to SP|O24543|AX2E_ Auxin-induced protein 22E ... 28 2.7
TC218140 28 2.7
TC230127 28 2.7
BE661128 similar to GP|5524201|gb| cryptochrome 1 {Lycopersicon ... 27 3.5
TC217977 27 3.5
TC207921 similar to UP|Q6YBV9 (Q6YBV9) Cryptochrome 1, partial (... 27 3.5
TC228414 weakly similar to UP|Q8LPR8 (Q8LPR8) AT5g22640/MDJ22_6,... 27 3.5
TC220581 weakly similar to UP|Q9LUE4 (Q9LUE4) Similarity to oxid... 27 4.6
AW568793 weakly similar to GP|20135548|gb| farnesyl pyrophosphat... 27 4.6
TC229908 similar to UP|XT33_ARATH (Q8LC45) Probable xyloglucan e... 27 4.6
TC207652 26 7.9
TC230501 26 7.9
NP1129936 CHX3 [Glycine max] 26 7.9
>TC223641
Length = 841
Score = 160 bits (405), Expect = 3e-40
Identities = 79/103 (76%), Positives = 84/103 (80%)
Frame = +3
Query: 66 PVNGAYFLIFTVIVFAVTWACCCIFKKKPRDEIPYQELEMALPESASATVVESAEGWDQG 125
P AYFLI V+VF TWACC +K R+E+PYQELEMALPESASAT VESAEGWDQ
Sbjct: 186 PSKWAYFLILMVLVFGGTWACCAC-RKNHRNEVPYQELEMALPESASATNVESAEGWDQD 362
Query: 126 WDDDWDDNVAVKSPVVRHAGSISANGLTSRSSNKDGWEDNWDD 168
WDDDWD NVAVKSP AGSISANGLTSRSSNKDGWE+NWDD
Sbjct: 363 WDDDWDVNVAVKSPAAL-AGSISANGLTSRSSNKDGWENNWDD 488
>TC222007 UP|Q9LKZ5 (Q9LKZ5) Receptor-like protein kinase 2, complete
Length = 3192
Score = 28.5 bits (62), Expect = 1.6
Identities = 12/30 (40%), Positives = 20/30 (66%)
Frame = -3
Query: 116 VESAEGWDQGWDDDWDDNVAVKSPVVRHAG 145
+E+ +G + +DDD +D+V VKSP + G
Sbjct: 3004 LEAEDGDSKAFDDDKEDSVIVKSPSLEPGG 2915
>TC219395 UP|Q9LKZ4 (Q9LKZ4) Receptor-like protein kinase 3, partial (92%)
Length = 3037
Score = 28.1 bits (61), Expect = 2.1
Identities = 12/30 (40%), Positives = 19/30 (63%)
Frame = -1
Query: 116 VESAEGWDQGWDDDWDDNVAVKSPVVRHAG 145
+E+ +G + +DDD DD+V VK P + G
Sbjct: 2755 LEAEDGDSKAFDDDKDDSVIVKFPSLESGG 2666
>TC221095
Length = 446
Score = 28.1 bits (61), Expect = 2.1
Identities = 15/33 (45%), Positives = 19/33 (57%)
Frame = +3
Query: 96 DEIPYQELEMALPESASATVVESAEGWDQGWDD 128
D ELE A PE+ S+ VVESAE ++D
Sbjct: 135 DSQSQSELEDASPENGSSGVVESAENESSQYED 233
>BE211605 homologue to SP|O24543|AX2E_ Auxin-induced protein 22E
(Indole-3-acetic acid induced protein ARG14). [Mung bean
Vigna radiata], partial (26%)
Length = 411
Score = 27.7 bits (60), Expect = 2.7
Identities = 6/18 (33%), Positives = 15/18 (83%)
Frame = +3
Query: 72 FLIFTVIVFAVTWACCCI 89
FL++ + +F+++W+C C+
Sbjct: 144 FLLYALFIFSISWSCFCL 197
>TC218140
Length = 724
Score = 27.7 bits (60), Expect = 2.7
Identities = 17/56 (30%), Positives = 28/56 (49%)
Frame = +1
Query: 110 SASATVVESAEGWDQGWDDDWDDNVAVKSPVVRHAGSISANGLTSRSSNKDGWEDN 165
S+ +T+ + E WD +D ++K + S +G +SRS N DG +DN
Sbjct: 244 SSRSTITNTEEEWDAV--GTMEDIKSLKGELDALKLSGGTSGASSRSDNNDGGKDN 405
>TC230127
Length = 884
Score = 27.7 bits (60), Expect = 2.7
Identities = 12/32 (37%), Positives = 20/32 (62%)
Frame = +1
Query: 9 CFYNIVFQVTIKQIKSESTELTLDAGKGDCVL 40
CF + + V +K+ + T + + AGKGDC+L
Sbjct: 721 CFCSFLSLVVVKKNRMF*TNV*ISAGKGDCML 816
>BE661128 similar to GP|5524201|gb| cryptochrome 1 {Lycopersicon esculentum},
partial (20%)
Length = 599
Score = 27.3 bits (59), Expect = 3.5
Identities = 14/43 (32%), Positives = 19/43 (43%), Gaps = 3/43 (6%)
Frame = +1
Query: 83 TWACCCIFKK---KPRDEIPYQELEMALPESASATVVESAEGW 122
+W CCC F + R +P+Q L M E + S E W
Sbjct: 256 SWCCCCSFHMGT*RRRTILPWQGL*MVAQEQFGSP*FLSQESW 384
>TC217977
Length = 1340
Score = 27.3 bits (59), Expect = 3.5
Identities = 13/27 (48%), Positives = 18/27 (66%), Gaps = 2/27 (7%)
Frame = -3
Query: 141 VRHAGSISANGL--TSRSSNKDGWEDN 165
+RH S S++ TSRSS KD W+D+
Sbjct: 432 IRHGYSYSSDNFIWTSRSSIKDSWDDD 352
>TC207921 similar to UP|Q6YBV9 (Q6YBV9) Cryptochrome 1, partial (41%)
Length = 1261
Score = 27.3 bits (59), Expect = 3.5
Identities = 14/43 (32%), Positives = 19/43 (43%), Gaps = 3/43 (6%)
Frame = +3
Query: 83 TWACCCIFKK---KPRDEIPYQELEMALPESASATVVESAEGW 122
+W CCC F + R +P+Q L M E + S E W
Sbjct: 348 SWCCCCSFHMGT*RRRTILPWQGL*MVAQEQFGSP*FLSQESW 476
>TC228414 weakly similar to UP|Q8LPR8 (Q8LPR8) AT5g22640/MDJ22_6, partial
(6%)
Length = 889
Score = 27.3 bits (59), Expect = 3.5
Identities = 21/57 (36%), Positives = 24/57 (41%), Gaps = 4/57 (7%)
Frame = +1
Query: 109 ESASATVVESA----EGWDQGWDDDWDDNVAVKSPVVRHAGSISANGLTSRSSNKDG 161
E A A VVE E D WDD+ DDN A S GS+ T + K G
Sbjct: 229 EKAPAKVVEVPAEVEEEDDDDWDDEEDDNSAQSS-----FGSVEQGQTTDQLKGKPG 384
>TC220581 weakly similar to UP|Q9LUE4 (Q9LUE4) Similarity to oxidoreductase,
partial (22%)
Length = 862
Score = 26.9 bits (58), Expect = 4.6
Identities = 18/74 (24%), Positives = 34/74 (45%)
Frame = +2
Query: 17 VTIKQIKSESTELTLDAGKGDCVLHVTVVTPVPEASFFLRLPSFDKILTPVNGAYFLIFT 76
+T++ ++ E T + G C + +V ++ PS+ K+L P + L+
Sbjct: 365 LTLRAMEFEPTVGRIPMGSA-CECAIAIVDSACRGDMYVTNPSWVKVLLP----WKLLCP 529
Query: 77 VIVFAVTWACCCIF 90
+V WACC +F
Sbjct: 530 ELV---DWACCLVF 562
>AW568793 weakly similar to GP|20135548|gb| farnesyl pyrophosphate synthase
{Malus x domestica}, partial (15%)
Length = 526
Score = 26.9 bits (58), Expect = 4.6
Identities = 18/51 (35%), Positives = 24/51 (46%)
Frame = +1
Query: 87 CCIFKKKPRDEIPYQELEMALPESASATVVESAEGWDQGWDDDWDDNVAVK 137
CCI KKP + P + L AS VE+ D G DD D++ V+
Sbjct: 112 CCIPAKKPDNAKP*EALGERAV*RASYGYVETGPIADGGRTDDRSDDLTVR 264
>TC229908 similar to UP|XT33_ARATH (Q8LC45) Probable xyloglucan
endotransglucosylase/hydrolase protein 33 precursor
(At-XTH33) (XTH-33) , partial (55%)
Length = 1250
Score = 26.9 bits (58), Expect = 4.6
Identities = 12/24 (50%), Positives = 14/24 (58%), Gaps = 2/24 (8%)
Frame = +3
Query: 82 VTWACCCIFKKKP--RDEIPYQEL 103
VTW+CCC K P DEI + L
Sbjct: 30 VTWSCCCFLYKFPHNHDEIDIELL 101
>TC207652
Length = 811
Score = 26.2 bits (56), Expect = 7.9
Identities = 16/58 (27%), Positives = 25/58 (42%), Gaps = 3/58 (5%)
Frame = +2
Query: 71 YFLIFTVIVFAVTW---ACCCIFKKKPRDEIPYQELEMALPESASATVVESAEGWDQG 125
+FLI + V ++W CCC KPR+++ Y A + + WD G
Sbjct: 629 HFLICLLSV--ISWHLIVCCCGIVYKPREKVQYGTSP*PSQVD*GALYIRNLYWWDSG 796
>TC230501
Length = 598
Score = 26.2 bits (56), Expect = 7.9
Identities = 13/32 (40%), Positives = 17/32 (52%)
Frame = +1
Query: 104 EMALPESASATVVESAEGWDQGWDDDWDDNVA 135
E+AL E E ++G+D D DWDD A
Sbjct: 19 ELALSED------EDSDGYDDEEDSDWDDGEA 96
>NP1129936 CHX3 [Glycine max]
Length = 1125
Score = 26.2 bits (56), Expect = 7.9
Identities = 9/25 (36%), Positives = 15/25 (60%)
Frame = +3
Query: 122 WDQGWDDDWDDNVAVKSPVVRHAGS 146
W GW W D++ + +P+V AG+
Sbjct: 123 WLMGWPT*WPDSIEIFTPIV*VAGA 197
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.319 0.135 0.427
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,176,589
Number of Sequences: 63676
Number of extensions: 104614
Number of successful extensions: 797
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 770
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 783
length of query: 168
length of database: 12,639,632
effective HSP length: 90
effective length of query: 78
effective length of database: 6,908,792
effective search space: 538885776
effective search space used: 538885776
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 55 (25.8 bits)
Medicago: description of AC146784.10


