

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146776.6 - phase: 0
(388 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
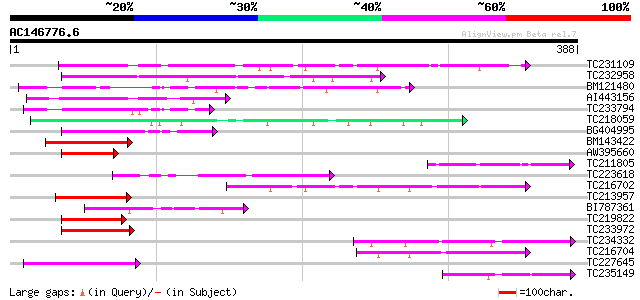
Score E
Sequences producing significant alignments: (bits) Value
TC231109 weakly similar to UP|Q9LUP7 (Q9LUP7) Gb|AAD25583.1, par... 117 7e-27
TC232958 97 9e-21
BM121480 77 2e-14
AI443156 74 1e-13
TC233794 similar to GB|AAP37713.1|30725382|BT008354 At3g23880 {A... 70 2e-12
TC218059 weakly similar to UP|Q84KQ9 (Q84KQ9) F-box, partial (7%) 68 6e-12
BG404995 similar to GP|22831068|dbj OJ1131_E05.32 {Oryza sativa ... 65 5e-11
BM143422 64 2e-10
AW395660 62 3e-10
TC211805 similar to UP|Q9K6E7 (Q9K6E7) BH3782 protein, partial (... 60 1e-09
TC223618 60 1e-09
TC216702 similar to UP|Q9LDP5 (Q9LDP5) FLORICAULA/LEAFY-like pro... 60 2e-09
TC213957 similar to GB|AAP37713.1|30725382|BT008354 At3g23880 {A... 59 3e-09
BI787361 59 5e-09
TC219822 homologue to UP|CENB_HUMAN (P07199) Major centromere au... 57 1e-08
TC233972 weakly similar to GB|AAP37713.1|30725382|BT008354 At3g2... 55 4e-08
TC234332 54 2e-07
TC216704 similar to UP|Q7PIN1 (Q7PIN1) ENSANGP00000023661 (Fragm... 50 2e-06
TC227645 similar to UP|Q9FZK1 (Q9FZK1) F17L21.13, partial (61%) 44 1e-04
TC235149 43 3e-04
>TC231109 weakly similar to UP|Q9LUP7 (Q9LUP7) Gb|AAD25583.1, partial (8%)
Length = 1061
Score = 117 bits (294), Expect = 7e-27
Identities = 104/353 (29%), Positives = 170/353 (47%), Gaps = 30/353 (8%)
Frame = +3
Query: 34 ADMPEEIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQFAKKHVSISTAYPQLVSVFVSI 93
A++P E++ EIL RLPV+S+++ R CK W+++I F H++ S S +
Sbjct: 87 ANLPVEVVTEILSRLPVKSVIRLRSTCKWWRSIIDSRHFVLFHLNKSH------SSLILR 248
Query: 94 AKCNLVSYPLKPLLDNPSAHRVEPADFEMIHTTSMTIIGSCNGLLCLSDFY-QFTLWNPS 152
+ +L S LK +P + VE + M ++ S+ ++GS NGLLC+S+ LWNP
Sbjct: 249 HRSHLYSLDLK----SPEQNPVELSHPLMCYSNSIKVLGSSNGLLCISNVADDIALWNPF 416
Query: 153 IKLKSKPSPTIIAFDSF---DSKRFLYR--GFGYDQVNDRYKVLAVVQNCYNLD------ 201
++ I+ D F S F R GFG+ ++ YK+L++ Y +D
Sbjct: 417 LR-----KHRILPADRFHRPQSSLFAARVYGFGHHSPSNDYKLLSIT---YFVDLQKRTF 572
Query: 202 ETKTLIYTFGGKDWTTIQKFPCDPSRCDLGRLGVGKFVSGNLNWIVSKK-------VIVF 254
+++ +YT W + P + C +GV FVSG+L+W+V++K +IV
Sbjct: 573 DSQVQLYTLKSDSWKNLPSMPY--ALCCARTMGV--FVSGSLHWLVTRKLQPHEPDLIVS 740
Query: 255 FDIEKETYGEMSLPQDY-GDKNTVLYVSSNRIYVSFDHSNKTHWVVWMMKEYGVVESWTK 313
FD+ +ET+ E+ LP GD + + + + V +H T + VW+M+ YG SW K
Sbjct: 741 FDLTRETFHEVPLPVTVNGDFDMQVALLGGCLCV-VEHRG-TGFDVWVMRVYGSRNSWEK 914
Query: 314 LMIIPQD----------KLTSPGPYCLSDALFISEHGVLLMRPQHSKLSVYNL 356
L + ++ KL P L + EH + KL YNL
Sbjct: 915 LFTLLENNDHHEMMGSGKLKYVRPLALDGDRVLFEHNRI-------KLCWYNL 1052
>TC232958
Length = 758
Score = 97.4 bits (241), Expect = 9e-21
Identities = 77/234 (32%), Positives = 111/234 (46%), Gaps = 12/234 (5%)
Frame = +1
Query: 36 MPEEIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQFAKKHVSISTAYPQLVSVFVSIAK 95
+P+++I EILLRLPV+SL++F+ VCK W LISDP+FAK H + A + +F++ +
Sbjct: 61 LPQDLITEILLRLPVKSLVRFKSVCKSWLFLISDPRFAKSHFDLFAALADRI-LFIASSA 237
Query: 96 CNLVSYPLKPLLDNPSAHRVEPADF--EMIHTTSMTIIGSCNGLLCLSDFYQFTLWNPSI 153
L S L + SA D + + IIGSC G + L +WNP+
Sbjct: 238 PELRSIDFNASLHDDSASVAVTVDLPAPKPYFHFVEIIGSCRGFILLHCLSHLCVWNPTT 417
Query: 154 KLKSKPSPTIIAFDSFDSKRF-LYRGFGYDQVNDRYKVLAVVQNCYNLDETKTL--IYTF 210
+ K P F + D+ F L GFGYD D + VV CYN I++
Sbjct: 418 GV-HKVVPLSPIFFNKDAVFFTLLCGFGYDPSTDDF---LVVHACYNPKHQANCAEIFSL 585
Query: 211 GGKDWTTIQ--KFPCDPSRCDLGRLGVGKFVSGNLNWI-----VSKKVIVFFDI 257
W I+ FP R G F++G ++W+ S VIV FD+
Sbjct: 586 RANAWKGIEGIHFPYTHFRYTNRYNQFGSFLNGAIHWLAFRINASINVIVAFDL 747
>BM121480
Length = 917
Score = 76.6 bits (187), Expect = 2e-14
Identities = 85/280 (30%), Positives = 123/280 (43%), Gaps = 9/280 (3%)
Frame = +3
Query: 7 NRSKTRRRLRQHSQNVTVFTQSVSEITADMPEEIIVEILLRLPVRSLLQFRCVCKLWKTL 66
+ K R+R + QN ++ T +P+E+I EILLRLPV+SLL+F+CV L+KT
Sbjct: 6 HEKKERKRSMKKKQNQSLTT---------LPQELIREILLRLPVKSLLRFKCV--LFKTC 152
Query: 67 ISDPQFAKKHVSISTAYPQLVSVFVSIAKCNLVSYPLKPLLDNPSAHRVEPADFEMIHTT 126
D +V +PL PL PS + DF +
Sbjct: 153 SRD----------------------------VVYFPL-PL---PSIPCLRLDDFGI---- 224
Query: 127 SMTIIGSCNGLLCL--SDFYQFTLWNPSIKLKSKPSPTIIAFDSFDSKRFLYRGFGYDQV 184
I+GSC GL+ L D LWNPS+ + K P D F Y GFGYD+
Sbjct: 225 RPKILGSCRGLVLLYYDDSANLILWNPSLG-RHKXLPNY----RDDITSFPY-GFGYDES 386
Query: 185 NDRYKVLAVVQNCYNLDETKTLIYTFGGKDW--TTIQKFPCDPSRCDLGRLGVGKFVSGN 242
D Y ++ + + ET IY+F + W TI P + G ++G
Sbjct: 387 KDEYLLILIGLPKFG-PETGADIYSFKTESWKTDTIVYDPLXRYXAEDXIARAGSLLNGA 563
Query: 243 LNWIVSKK-----VIVFFDIEKETYGEMSLPQDYGDKNTV 277
L+W V + VI+ FD+ + T + LP D++TV
Sbjct: 564 LHWFVFSESKXDHVIIAFDLVERTLSXIPLP--LADRSTV 677
>AI443156
Length = 414
Score = 73.6 bits (179), Expect = 1e-13
Identities = 49/146 (33%), Positives = 74/146 (50%), Gaps = 6/146 (4%)
Frame = +3
Query: 12 RRRLRQHSQNVTVFTQSVSEITADMPEEIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQ 71
RRR H+ +TV A++P E++ EIL RLPV+S+++ R CK W+++I
Sbjct: 36 RRRNWMHANPITVSNM------ANLPVEVVTEILSRLPVKSVIRLRSTCKWWRSIIDSRH 197
Query: 72 FAKKHVSISTAYPQLVSVFVSIAKCNLVSYPLKPLLDNPSAHRVEPADFEMIH-----TT 126
F H++ S + + + L S LK LLD P FE+ H +
Sbjct: 198 FILFHLNKSH------TSLILRHRSQLYSLDLKSLLD--------PNPFELSHPLMCYSN 335
Query: 127 SMTIIGSCNGLLCLSDFY-QFTLWNP 151
S+ ++GS NGLLC+S+ LWNP
Sbjct: 336 SIKVLGSSNGLLCISNVXDDIALWNP 413
>TC233794 similar to GB|AAP37713.1|30725382|BT008354 At3g23880 {Arabidopsis
thaliana;} , partial (9%)
Length = 449
Score = 69.7 bits (169), Expect = 2e-12
Identities = 54/146 (36%), Positives = 81/146 (54%), Gaps = 15/146 (10%)
Frame = +1
Query: 10 KTRRRLRQHSQNVTVFTQSVSEITADMPEEIIVEILLRLPVRSLLQFRCVCKLWKTLISD 69
K R+R + QN ++ T +P+E+I EILLRLPV+SLL+F+CVCK + +LISD
Sbjct: 58 KERKRSMKKKQNQSLTT---------LPQELIREILLRLPVKSLLRFKCVCKSFLSLISD 210
Query: 70 PQFAKKHVSISTA------------YPQLV---SVFVSIAKCNLVSYPLKPLLDNPSAHR 114
PQF H +++ + Y Q + SVF + ++ ++V +PL PL PS
Sbjct: 211 PQFVISHYALAASPTHRLILRSHDFYAQSIATESVFKTCSR-DVVYFPL-PL---PSIPC 375
Query: 115 VEPADFEMIHTTSMTIIGSCNGLLCL 140
+ DF + I+GSC GL+ L
Sbjct: 376 LRLDDFGI----RPKILGSCRGLVLL 441
>TC218059 weakly similar to UP|Q84KQ9 (Q84KQ9) F-box, partial (7%)
Length = 1185
Score = 68.2 bits (165), Expect = 6e-12
Identities = 77/344 (22%), Positives = 135/344 (38%), Gaps = 45/344 (13%)
Frame = +3
Query: 15 LRQHSQNVTVFTQSVSEITADMPEEIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQFAK 74
L+ N VF + +P E++ +L RLP + LL +CVCK W LI+DP F
Sbjct: 3 LQTSHSNFQVFQRR*VMSMEHLPGELVSNVLSRLPSKVLLLCKCVCKSWFDLITDPHFVS 182
Query: 75 KHVSISTAYPQLVSVFVSIAK---CNLVSY--PLKPLLDNPSAHRVE-----PADFEMIH 124
+ + + + I + L +Y L ++P H P ++ H
Sbjct: 183 NYYVVYNSLQSQEEHLLVIRRPFFSGLKTYISVLSWNTNDPKKHVSSDVLNPPYEYNSDH 362
Query: 125 TTSMTIIGSCNGLLCLSDFYQFTLWNPSIKLKSKPSPTIIAFDSFDSKRFL--------- 175
I+G CNG+ L NP++ + +P++ F + F
Sbjct: 363 KYWTEILGPCNGIYFLEG-------NPNVLM----NPSLGEFKALPKSHFTSPHGTYTFT 509
Query: 176 -YRGFGYDQVNDRYKVLAVVQNCYNLDETKTL------IYTFGGKDWTTIQKFPCDPSRC 228
Y GFG+D + YKV+ + + + + +Y+ W + DPS
Sbjct: 510 DYAGFGFDPKTNDYKVVVLKDLWLKETDEREIGYWSAELYSLNSNSWRKL-----DPSLL 674
Query: 229 DL-----GRLGVGKFVSGNLNW-------IVSKKVIVFFDIEKETYGEMSLP--QDYGDK 274
L G V + + +W ++ V++ FD+ KE++ ++ +P +D D+
Sbjct: 675 PLPIEIWGSSRVFTYANNCCHWWGFVEESDATQDVVLAFDMVKESFRKIRVPKIRDSSDE 854
Query: 275 NTVLYV-----SSNRIYVSFDHSNKTHWVVWMMKEYGVVESWTK 313
V +S V + + VW+MK+Y SW K
Sbjct: 855 KFGTLVPFEESASIGFLVYPVRGTEKRFDVWVMKDYWDEGSWVK 986
>BG404995 similar to GP|22831068|dbj OJ1131_E05.32 {Oryza sativa (japonica
cultivar-group)}, partial (8%)
Length = 394
Score = 65.1 bits (157), Expect = 5e-11
Identities = 41/107 (38%), Positives = 59/107 (54%)
Frame = +1
Query: 36 MPEEIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQFAKKHVSISTAYPQLVSVFVSIAK 95
+P E++ EIL RLPVRSLL+FR K WK+LI H++ S S+ + +
Sbjct: 40 LPREVLTEILSRLPVRSLLRFRSTSKSWKSLIDSQHLNWLHLTRSLTLASNTSLILRV-D 216
Query: 96 CNLVSYPLKPLLDNPSAHRVEPADFEMIHTTSMTIIGSCNGLLCLSD 142
+L P LD P V M ++ S+T++GSCNGLLC+S+
Sbjct: 217 SDLYQTNF-PTLDPP----VSLNHPLMCYSNSITLLGSCNGLLCISN 342
>BM143422
Length = 424
Score = 63.5 bits (153), Expect = 2e-10
Identities = 28/60 (46%), Positives = 42/60 (69%)
Frame = +1
Query: 25 FTQSVSEITADMPEEIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQFAKKHVSISTAYP 84
F++S+ T +P ++I ILL+LPV+S+ +F+CVCK W +LISDPQF H ++ A P
Sbjct: 196 FSRSMKNHTQTLPLDLIELILLKLPVKSVTRFKCVCKSWLSLISDPQFGFSHFDLALAVP 375
>AW395660
Length = 381
Score = 62.4 bits (150), Expect = 3e-10
Identities = 27/39 (69%), Positives = 33/39 (84%)
Frame = +3
Query: 36 MPEEIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQFAK 74
+P+E++VEIL RLPV+SLLQFRCVCK W +LI DP F K
Sbjct: 264 LPDELVVEILSRLPVKSLLQFRCVCKSWMSLIYDPYFMK 380
>TC211805 similar to UP|Q9K6E7 (Q9K6E7) BH3782 protein, partial (13%)
Length = 693
Score = 60.5 bits (145), Expect = 1e-09
Identities = 39/100 (39%), Positives = 58/100 (58%)
Frame = +1
Query: 287 VSFDHSNKTHWVVWMMKEYGVVESWTKLMIIPQDKLTSPGPYCLSDALFISEHGVLLMRP 346
+++D+ KTH+VVWMMK+YG ESW KL+ IP + +P + S +ISE+G +L+
Sbjct: 31 MNYDYK-KTHFVVWMMKDYGARESWVKLVSIPY--VPNPENFSYSGPYYISENGEVLLMF 201
Query: 347 QHSKLSVYNLNNDGGLDYCTTISGQFARYLHIYNESLVSP 386
+ L +YN D Y SG+ +Y E+LVSP
Sbjct: 202 EFD-LILYN-PRDNSFKYPKIESGKGWFDAEVYVETLVSP 315
>TC223618
Length = 444
Score = 60.5 bits (145), Expect = 1e-09
Identities = 49/154 (31%), Positives = 69/154 (43%), Gaps = 2/154 (1%)
Frame = +1
Query: 71 QFAKKHVSISTAYPQLVSVFVSIAKCNLVSYPLKPLLDNPSAHRVEPADFEMIHTTSMTI 130
+F K S+S+ + L +V C+ ++YP+K N H I
Sbjct: 22 EFHLKSCSLSSLFNNLSTV------CDELNYPVK----NKFRHD--------------GI 129
Query: 131 IGSCNGLLCLSDFYQFTL-WNPSIKLKSKPSPTIIAFDSFDSKRFLYRGFGYDQVNDRYK 189
+GSCNGLLC + L WNPSI++ K P +++ F G GYD VN+ YK
Sbjct: 130 VGSCNGLLCFAIKGDCVLLWNPSIRVSKKSPPL---GNNWRPGCFTAFGLGYDHVNEDYK 300
Query: 190 VLAV-VQNCYNLDETKTLIYTFGGKDWTTIQKFP 222
V+AV E K +Y+ W IQ FP
Sbjct: 301 VVAVFCDPSEYFIECKVKVYSMATNSWRKIQDFP 402
>TC216702 similar to UP|Q9LDP5 (Q9LDP5) FLORICAULA/LEAFY-like protein,
partial (5%)
Length = 1631
Score = 59.7 bits (143), Expect = 2e-09
Identities = 57/229 (24%), Positives = 103/229 (44%), Gaps = 21/229 (9%)
Frame = +3
Query: 149 WNPSIKL-KSKPSPTIIAFDSFDSKRFLYR--GFGYDQVNDRYKVLAVVQNCYNLD---- 201
WNPS++ + P + D+ F R GFG+D YK++ + Y +D
Sbjct: 39 WNPSLRQHRILPYLPVPRRRHPDTTLFAARVCGFGFDHKTRDYKLVRI---SYFVDLHDR 209
Query: 202 --ETKTLIYTFGGKDWTTIQKFPCDPSRCDLGRLGVGKFVSGNLNWIVSKKV-------I 252
+ + +YT W T+ P + C +GV FV +L+W+V++K+ I
Sbjct: 210 SFDAQVKLYTLRANAWKTLPSLPY--ALCCARTMGV--FVGNSLHWVVTRKLEPDQPDLI 377
Query: 253 VFFDIEKETYGEMSLPQDYG-----DKNTVLYVSSNRIYVSFDHSNKTHWVVWMMKEYGV 307
+ FD+ + + E+ LP G + + L S + V+F +KT VW+M+EY
Sbjct: 378 IAFDLTHDIFRELPLPDTGGVDGGFEIDLALLGGSLCMTVNF---HKTRIDVWVMREYNR 548
Query: 308 VESWTKLMIIPQDKLTSPGPYCLSDALFISEHGVLLMRPQHSKLSVYNL 356
+SW K+ + + + C+ + S+ +L+ +L Y+L
Sbjct: 549 RDSWCKVFTLEESR-EMRSLKCVRPLGYSSDGNKVLLEHDRKRLFWYDL 692
>TC213957 similar to GB|AAP37713.1|30725382|BT008354 At3g23880 {Arabidopsis
thaliana;} , partial (9%)
Length = 696
Score = 59.3 bits (142), Expect = 3e-09
Identities = 27/52 (51%), Positives = 38/52 (72%)
Frame = +3
Query: 32 ITADMPEEIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQFAKKHVSISTAY 83
++ +P E+I EILLR PVRS+L+F+CVCK W +LISDPQF ++ S +
Sbjct: 21 LSVTLPLELIREILLRSPVRSVLRFKCVCKSWLSLISDPQFTHFDLAASPTH 176
>BI787361
Length = 423
Score = 58.5 bits (140), Expect = 5e-09
Identities = 41/132 (31%), Positives = 65/132 (49%), Gaps = 20/132 (15%)
Frame = +3
Query: 52 SLLQFRCVCKLWKTLISDPQFAKKHVSIS-----------TAYPQLVSVFVSIAKCNLVS 100
+L++FR V + W +LI DP F K H+ S Y + V V +A C++
Sbjct: 3 ALMRFRYVSETWNSLIFDPTFVKLHLERSPKNTHVLLEFQAIYDRDVGQQVGVAPCSI-- 176
Query: 101 YPLKPLLDNPSAHRVEPADFEMIHTTSMTIIGSCNGLLCLSDFY---------QFTLWNP 151
+ L++NPS ++ HT S I GSCNGL+C++ + Q+ LWNP
Sbjct: 177 ---RRLVENPS-FTIDDCLTLFKHTNS--IFGSCNGLVCMTKCFDVREFEEECQYRLWNP 338
Query: 152 SIKLKSKPSPTI 163
+ + S+ SP +
Sbjct: 339 ATGIMSEYSPPL 374
>TC219822 homologue to UP|CENB_HUMAN (P07199) Major centromere autoantigen B
(Centromere protein B) (CENP-B), partial (5%)
Length = 710
Score = 57.0 bits (136), Expect = 1e-08
Identities = 24/45 (53%), Positives = 34/45 (75%)
Frame = +1
Query: 36 MPEEIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQFAKKHVSIS 80
+PE +++EIL+ + V + LQ RCVCK WK+L+ DPQF KKH+ S
Sbjct: 196 LPEGLMIEILVWIRVSNPLQLRCVCKRWKSLVVDPQFVKKHLHTS 330
>TC233972 weakly similar to GB|AAP37713.1|30725382|BT008354 At3g23880
{Arabidopsis thaliana;} , partial (10%)
Length = 631
Score = 55.5 bits (132), Expect = 4e-08
Identities = 27/50 (54%), Positives = 37/50 (74%)
Frame = -2
Query: 36 MPEEIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQFAKKHVSISTAYPQ 85
+P+EII+EILLRLPV+SL F+ VCK +LIS+P FAK H + A+ +
Sbjct: 540 LPQEIIIEILLRLPVKSLPSFKFVCKS*LSLISNPHFAKWHFERNAAHTE 391
>TC234332
Length = 644
Score = 53.5 bits (127), Expect = 2e-07
Identities = 45/177 (25%), Positives = 75/177 (41%), Gaps = 25/177 (14%)
Frame = +2
Query: 236 GKFVSGNLNWI---------------VSKKVIVFFDIEKETYGEMSLPQ---DYGDKNTV 277
G V G +NW+ + VI +D++ E+Y + +P +
Sbjct: 20 GASVRGTVNWLALPNSSSDYQWETVTIDDLVIFSYDLKNESYRYLLMPDGLLEVPHSPPE 199
Query: 278 LYVSSNRIYVSFDHSNKTHWVVWMMKEYGVVESWTKLMIIPQDKLTSPGPY-------CL 330
L V + +S H H+ W+MKE+GV +SWT+ + I D+L G + C+
Sbjct: 200 LVVLKGCLCLSHRHGGN-HFGFWLMKEFGVEKSWTRFLNISYDQLHIHGGFLDHPVILCM 376
Query: 331 SDALFISEHGVLLMRPQHSKLSVYNLNNDGGLDYCTTISGQFARYLHIYNESLVSPH 387
S+ + VLL H K +YN ++ Y G+F + Y +S V P+
Sbjct: 377 SE----DDGVVLLENGGHGKFILYNKRDNTIECYGELDKGRFQFLSYDYAQSFVMPY 535
>TC216704 similar to UP|Q7PIN1 (Q7PIN1) ENSANGP00000023661 (Fragment),
partial (5%)
Length = 1232
Score = 50.1 bits (118), Expect = 2e-06
Identities = 36/131 (27%), Positives = 61/131 (46%), Gaps = 12/131 (9%)
Frame = +2
Query: 238 FVSGNLNWIVSKKV-------IVFFDIEKETYGEMSLPQDYG-----DKNTVLYVSSNRI 285
FV +L+W+V++K+ IV FD+ E + E+ LP G + + L S +
Sbjct: 2 FVGNSLHWVVTRKLEPDQPDLIVAFDLTHEIFTELPLPDTGGVGGGFEIDVALLGDSLCM 181
Query: 286 YVSFDHSNKTHWVVWMMKEYGVVESWTKLMIIPQDKLTSPGPYCLSDALFISEHGVLLMR 345
V+F +S VW+M+EY +SW KL + + CL + S+ +L+
Sbjct: 182 TVNFHNSKMD---VWVMREYNRGDSWCKLFTLEESA*VEIVQXCLRPLGYSSDGNKVLLE 352
Query: 346 PQHSKLSVYNL 356
+L Y+L
Sbjct: 353 HDRKRLCWYDL 385
>TC227645 similar to UP|Q9FZK1 (Q9FZK1) F17L21.13, partial (61%)
Length = 1671
Score = 43.9 bits (102), Expect = 1e-04
Identities = 22/82 (26%), Positives = 44/82 (52%), Gaps = 2/82 (2%)
Frame = +3
Query: 10 KTRR-RLRQHSQNVTVFTQSVS-EITADMPEEIIVEILLRLPVRSLLQFRCVCKLWKTLI 67
K+RR R R S + T+ + EI D PE++ ++ RLP+ + +FR VC+ W +++
Sbjct: 279 KSRRDRSRGKSSGRSCTTEVMEQEIWKDFPEDLFEAVIARLPISTFFRFRSVCRQWNSML 458
Query: 68 SDPQFAKKHVSISTAYPQLVSV 89
+ F++ ++ P ++
Sbjct: 459 NSQSFSQHCTQVTQENPWFYTI 524
>TC235149
Length = 435
Score = 42.7 bits (99), Expect = 3e-04
Identities = 31/101 (30%), Positives = 50/101 (48%), Gaps = 10/101 (9%)
Frame = +2
Query: 297 WVVWMMKEYGVVESWTKLMIIPQDKLTSPG---------PYCLSDALFISEHGVLLMRPQ 347
+VVW+ +E+GV SWT+L+ + + + G P C+S+ +E +LL +
Sbjct: 2 FVVWLTREFGVERSWTRLLNVSYEHFRNHGCPPYYRFVTPLCMSE----NEDVLLLANDE 169
Query: 348 HSKLSVYNLNNDGGLDYCTTI-SGQFARYLHIYNESLVSPH 387
S+ YNL D +D S +F+ H Y SLV P+
Sbjct: 170 GSEFVFYNL-RDNRIDRIQDFDSYKFSFLSHDYVPSLVLPY 289
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.323 0.138 0.430
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,920,779
Number of Sequences: 63676
Number of extensions: 301195
Number of successful extensions: 1558
Number of sequences better than 10.0: 85
Number of HSP's better than 10.0 without gapping: 1535
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1542
length of query: 388
length of database: 12,639,632
effective HSP length: 99
effective length of query: 289
effective length of database: 6,335,708
effective search space: 1831019612
effective search space used: 1831019612
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 60 (27.7 bits)
Medicago: description of AC146776.6


