

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146759.10 + phase: 1 /partial
(178 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
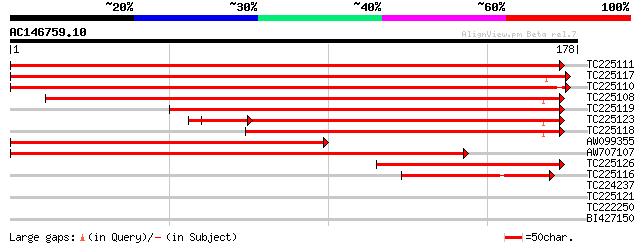
Sequences producing significant alignments: (bits) Value
TC225111 similar to UP|Q9FJA6 (Q9FJA6) 40S ribosomal protein S3 ... 340 3e-94
TC225117 homologue to UP|Q9FJA6 (Q9FJA6) 40S ribosomal protein S... 338 8e-94
TC225110 homologue to UP|Q9M339 (Q9M339) Ribosomal protein S3a-l... 336 3e-93
TC225108 similar to UP|Q9M339 (Q9M339) Ribosomal protein S3a-lik... 311 1e-85
TC225119 similar to UP|Q9FJA6 (Q9FJA6) 40S ribosomal protein S3 ... 242 6e-65
TC225123 similar to UP|Q9FJA6 (Q9FJA6) 40S ribosomal protein S3 ... 189 5e-49
TC225118 similar to UP|Q9FJA6 (Q9FJA6) 40S ribosomal protein S3 ... 189 8e-49
AW099355 138 1e-33
AW707107 134 2e-32
TC225126 similar to UP|Q9FJA6 (Q9FJA6) 40S ribosomal protein S3 ... 117 4e-27
TC225116 similar to UP|Q7X9L6 (Q7X9L6) 40S ribosomal protein (Fr... 80 4e-16
TC224237 similar to UP|Q8HDK9 (Q8HDK9) Ribosomal protein S3, par... 30 0.79
TC225121 UP|G3PA_PEA (P12858) Glyceraldehyde-3-phosphate dehydro... 30 0.79
TC222250 similar to UP|Q7SH02 (Q7SH02) Probable AMP deaminase, p... 27 5.1
BI427150 27 6.7
>TC225111 similar to UP|Q9FJA6 (Q9FJA6) 40S ribosomal protein S3
(AT5g35530/MOK9_14), partial (91%)
Length = 1035
Score = 340 bits (871), Expect = 3e-94
Identities = 170/174 (97%), Positives = 172/174 (98%)
Frame = +3
Query: 1 EKGRRIRELTSVVQKRFKFPENSVELYAEKVNNRGLCAIAQAESLRYKLLGGLAVRRACY 60
EKGRRIRELTSVVQKRFKFPENSVELYAEKVNNRGLCAIAQAESLRYKLLGGLAVRRACY
Sbjct: 297 EKGRRIRELTSVVQKRFKFPENSVELYAEKVNNRGLCAIAQAESLRYKLLGGLAVRRACY 476
Query: 61 GVLRFVMESGAKGCEVIVSGKLRAQRAKSMKFKDGYMISSGQPVKDYIDSAVRHVLLRQG 120
GVLRFVMESGAKGCEVIVSGKLRAQRAKSMKFKDGYMISSGQPVKDYIDSAVRHVLLRQG
Sbjct: 477 GVLRFVMESGAKGCEVIVSGKLRAQRAKSMKFKDGYMISSGQPVKDYIDSAVRHVLLRQG 656
Query: 121 VLGIKVKIMLDWDPKGKQGPKTPLPDIVTIHTPKEEEEYIRPAAVVANDIEVPV 174
VLGIKVKIMLDWDPKGKQGPKTPLPD+VTIHTPKEEEEY RPAAV+A DIEVPV
Sbjct: 657 VLGIKVKIMLDWDPKGKQGPKTPLPDLVTIHTPKEEEEYARPAAVLATDIEVPV 818
>TC225117 homologue to UP|Q9FJA6 (Q9FJA6) 40S ribosomal protein S3
(AT5g35530/MOK9_14), partial (91%)
Length = 1116
Score = 338 bits (867), Expect = 8e-94
Identities = 171/178 (96%), Positives = 175/178 (98%), Gaps = 2/178 (1%)
Frame = +3
Query: 1 EKGRRIRELTSVVQKRFKFPENSVELYAEKVNNRGLCAIAQAESLRYKLLGGLAVRRACY 60
EKGRRIRELTSVVQKRFKFPENSVELYAEKVNNRGLCAIAQAESLRYKLLGGLAVRRACY
Sbjct: 288 EKGRRIRELTSVVQKRFKFPENSVELYAEKVNNRGLCAIAQAESLRYKLLGGLAVRRACY 467
Query: 61 GVLRFVMESGAKGCEVIVSGKLRAQRAKSMKFKDGYMISSGQPVKDYIDSAVRHVLLRQG 120
GVLRFVMESGAKGCEVIVSGKLRAQRAKSMKFKDGYMISSGQPVKDYIDSAVRHVLLRQG
Sbjct: 468 GVLRFVMESGAKGCEVIVSGKLRAQRAKSMKFKDGYMISSGQPVKDYIDSAVRHVLLRQG 647
Query: 121 VLGIKVKIMLDWDPKGKQGPKTPLPDIVTIHTPKEEEEYIRPAAVVA--NDIEVPVPV 176
VLGIKVKIMLDWDPKGKQGPKTPLPD+VTIH+PKEEEEYI+PA V+A NDIEVPVPV
Sbjct: 648 VLGIKVKIMLDWDPKGKQGPKTPLPDLVTIHSPKEEEEYIQPAPVLAANNDIEVPVPV 821
>TC225110 homologue to UP|Q9M339 (Q9M339) Ribosomal protein S3a-like protein
(AT3g53870/F5K20_170), partial (87%)
Length = 965
Score = 336 bits (862), Expect = 3e-93
Identities = 171/176 (97%), Positives = 173/176 (98%)
Frame = +2
Query: 1 EKGRRIRELTSVVQKRFKFPENSVELYAEKVNNRGLCAIAQAESLRYKLLGGLAVRRACY 60
EKGRRIRELTSVVQKRFKFPENSVELYAEKVNNRGLCAIAQAESLRYKLLGGLAVRRACY
Sbjct: 275 EKGRRIRELTSVVQKRFKFPENSVELYAEKVNNRGLCAIAQAESLRYKLLGGLAVRRACY 454
Query: 61 GVLRFVMESGAKGCEVIVSGKLRAQRAKSMKFKDGYMISSGQPVKDYIDSAVRHVLLRQG 120
GVLRFVMESGAKGCEVIVSGKLRAQRAKSMKFKDGYMISSGQPVKDYIDSAVRHVLLRQG
Sbjct: 455 GVLRFVMESGAKGCEVIVSGKLRAQRAKSMKFKDGYMISSGQPVKDYIDSAVRHVLLRQG 634
Query: 121 VLGIKVKIMLDWDPKGKQGPKTPLPDIVTIHTPKEEEEYIRPAAVVANDIEVPVPV 176
VLGIKVKIMLDWDPKGKQGPKTPLPD+VTIHTPKEEEEY RP AV+ANDIEV VPV
Sbjct: 635 VLGIKVKIMLDWDPKGKQGPKTPLPDLVTIHTPKEEEEYNRPPAVLANDIEV-VPV 799
>TC225108 similar to UP|Q9M339 (Q9M339) Ribosomal protein S3a-like protein
(AT3g53870/F5K20_170), partial (62%)
Length = 708
Score = 311 bits (797), Expect = 1e-85
Identities = 157/164 (95%), Positives = 161/164 (97%), Gaps = 1/164 (0%)
Frame = +2
Query: 12 VVQKRFKFPENSVELYAEKVNNRGLCAIAQAESLRYKLLGGLAVRRACYGVLRFVMESGA 71
VVQKRFKFPE SVELYAEKVNNRGLCAIAQAESLRYKLLGGLAVRRACYGVLRFVMESGA
Sbjct: 2 VVQKRFKFPEXSVELYAEKVNNRGLCAIAQAESLRYKLLGGLAVRRACYGVLRFVMESGA 181
Query: 72 KGCEVIVSGKLRAQRAKSMKFKDGYMISSGQPVKDYIDSAVRHVLLRQGVLGIKVKIMLD 131
KGCEVIVSGKLRAQRAKSMKFKDGYMISSGQPVKDYIDSAVRHVLLRQGVLGIKVKIMLD
Sbjct: 182 KGCEVIVSGKLRAQRAKSMKFKDGYMISSGQPVKDYIDSAVRHVLLRQGVLGIKVKIMLD 361
Query: 132 WDPKGKQGPKTPLPDIVTIHTPKEEEEYIRPAAVV-ANDIEVPV 174
WDPKGKQGPKTPLPD+VTIH+PKEEEEYI+PA V+ ANDIEVPV
Sbjct: 362 WDPKGKQGPKTPLPDLVTIHSPKEEEEYIQPAPVLAANDIEVPV 493
>TC225119 similar to UP|Q9FJA6 (Q9FJA6) 40S ribosomal protein S3
(AT5g35530/MOK9_14), partial (46%)
Length = 749
Score = 242 bits (618), Expect = 6e-65
Identities = 119/124 (95%), Positives = 122/124 (97%)
Frame = +1
Query: 51 GGLAVRRACYGVLRFVMESGAKGCEVIVSGKLRAQRAKSMKFKDGYMISSGQPVKDYIDS 110
GGLAVRRACYGVLRFVMESGAKGCEVIVSGKLRAQRAKSMKFKDGYMISSGQPVKDYIDS
Sbjct: 10 GGLAVRRACYGVLRFVMESGAKGCEVIVSGKLRAQRAKSMKFKDGYMISSGQPVKDYIDS 189
Query: 111 AVRHVLLRQGVLGIKVKIMLDWDPKGKQGPKTPLPDIVTIHTPKEEEEYIRPAAVVANDI 170
AVRHVLLRQGVLGIKVKIMLDWDPKGKQGPKTPLPD+VTIHTPKEEEEY RPAAV+A +I
Sbjct: 190 AVRHVLLRQGVLGIKVKIMLDWDPKGKQGPKTPLPDLVTIHTPKEEEEYARPAAVLATNI 369
Query: 171 EVPV 174
EVPV
Sbjct: 370 EVPV 381
>TC225123 similar to UP|Q9FJA6 (Q9FJA6) 40S ribosomal protein S3
(AT5g35530/MOK9_14), partial (44%)
Length = 604
Score = 189 bits (481), Expect = 5e-49
Identities = 99/115 (86%), Positives = 104/115 (90%), Gaps = 1/115 (0%)
Frame = +2
Query: 61 GVLRFVMESGAKGCEVIVSGKLRAQRAKSMKFKDGYMISSGQPVKDYIDSAVRHVLLRQG 120
GVL VM +VIVSGKLRAQRAKSMKFKDGYMISSGQPVKDYIDSAVRHVLLRQG
Sbjct: 104 GVLLCVMVLVVWFEQVIVSGKLRAQRAKSMKFKDGYMISSGQPVKDYIDSAVRHVLLRQG 283
Query: 121 VLGIKVKIMLDWDPKGKQGPKTPLPDIVTIHTPKEEEEYIRPAAVV-ANDIEVPV 174
VLGIKVKIMLDWDPKGKQGPKTPLPD+VTIH+PKEEEEYI+PA V+ ANDIEVPV
Sbjct: 284 VLGIKVKIMLDWDPKGKQGPKTPLPDLVTIHSPKEEEEYIQPAPVLAANDIEVPV 448
Score = 44.3 bits (103), Expect = 3e-05
Identities = 19/20 (95%), Positives = 19/20 (95%)
Frame = +1
Query: 57 RACYGVLRFVMESGAKGCEV 76
R CYGVLRFVMESGAKGCEV
Sbjct: 4 RGCYGVLRFVMESGAKGCEV 63
>TC225118 similar to UP|Q9FJA6 (Q9FJA6) 40S ribosomal protein S3
(AT5g35530/MOK9_14), partial (38%)
Length = 556
Score = 189 bits (479), Expect = 8e-49
Identities = 94/101 (93%), Positives = 99/101 (97%), Gaps = 1/101 (0%)
Frame = +3
Query: 75 EVIVSGKLRAQRAKSMKFKDGYMISSGQPVKDYIDSAVRHVLLRQGVLGIKVKIMLDWDP 134
+VIVSGKLRAQRAKSMKFKDGYMISSGQPVKDYIDSAVRHVLLRQGVLGIKVKIMLDWDP
Sbjct: 99 QVIVSGKLRAQRAKSMKFKDGYMISSGQPVKDYIDSAVRHVLLRQGVLGIKVKIMLDWDP 278
Query: 135 KGKQGPKTPLPDIVTIHTPKEEEEYIRPAAVV-ANDIEVPV 174
KGKQGPKTPLPD+VTIH+PKEEEEYI+PA V+ ANDIEVPV
Sbjct: 279 KGKQGPKTPLPDLVTIHSPKEEEEYIQPAPVLAANDIEVPV 401
>AW099355
Length = 484
Score = 138 bits (348), Expect = 1e-33
Identities = 73/100 (73%), Positives = 77/100 (77%)
Frame = +2
Query: 1 EKGRRIRELTSVVQKRFKFPENSVELYAEKVNNRGLCAIAQAESLRYKLLGGLAVRRACY 60
E +R ELTSVV RFK PE SVELYAE VNNR +CAI QAE+LRYKL GLAVR ACY
Sbjct: 179 EDVKRXMELTSVVHNRFKVPETSVELYAELVNNRRVCAIVQAETLRYKLFDGLAVRMACY 358
Query: 61 GVLRFVMESGAKGCEVIVSGKLRAQRAKSMKFKDGYMISS 100
LRFVMESGA GCEVI SG LRAQRA S +F D YMISS
Sbjct: 359 SALRFVMESGANGCEVIASG*LRAQRATSHEFNDRYMISS 478
>AW707107
Length = 583
Score = 134 bits (338), Expect = 2e-32
Identities = 69/144 (47%), Positives = 99/144 (67%)
Frame = +2
Query: 1 EKGRRIRELTSVVQKRFKFPENSVELYAEKVNNRGLCAIAQAESLRYKLLGGLAVRRACY 60
EK R I +LTS++ +FKFPEN++ LY ++NN LC I QA+SL Y LL L++ Y
Sbjct: 149 EK*RTITKLTSLIHNKFKFPENTI*LYTNRINNTYLCTITQAKSLLYNLLNNLSILTTYY 328
Query: 61 GVLRFVMESGAKGCEVIVSGKLRAQRAKSMKFKDGYMISSGQPVKDYIDSAVRHVLLRQG 120
+LRF++ + + E++ + KLR+ R+KS+ FK Y+ISS QP+KDYI SA RHVLLRQ
Sbjct: 329 NILRFII*TKTQKYEIMFNIKLRSHRSKSIIFKHIYIISSRQPIKDYIYSAXRHVLLRQT 508
Query: 121 VLGIKVKIMLDWDPKGKQGPKTPL 144
+L +++ IML + P K +TPL
Sbjct: 509 ILIMQINIMLYYYPNKKHSSQTPL 580
>TC225126 similar to UP|Q9FJA6 (Q9FJA6) 40S ribosomal protein S3
(AT5g35530/MOK9_14), partial (20%)
Length = 1251
Score = 117 bits (292), Expect = 4e-27
Identities = 55/59 (93%), Positives = 57/59 (96%)
Frame = +2
Query: 116 LLRQGVLGIKVKIMLDWDPKGKQGPKTPLPDIVTIHTPKEEEEYIRPAAVVANDIEVPV 174
LLRQGVLGIKVKIMLDWDPKGKQGPKTPLPD+VTIHTPKEEEEY RPAAV+A DIEVPV
Sbjct: 878 LLRQGVLGIKVKIMLDWDPKGKQGPKTPLPDLVTIHTPKEEEEYARPAAVLATDIEVPV 1054
>TC225116 similar to UP|Q7X9L6 (Q7X9L6) 40S ribosomal protein (Fragment),
partial (51%)
Length = 900
Score = 80.5 bits (197), Expect = 4e-16
Identities = 39/48 (81%), Positives = 41/48 (85%)
Frame = +1
Query: 124 IKVKIMLDWDPKGKQGPKTPLPDIVTIHTPKEEEEYIRPAAVVANDIE 171
IKVKIMLDWDPKG Q PKTP PD+VTIHTPK EEEY P AV+ANDIE
Sbjct: 1 IKVKIMLDWDPKGNQSPKTPFPDLVTIHTPK-EEEYK*PPAVLANDIE 141
>TC224237 similar to UP|Q8HDK9 (Q8HDK9) Ribosomal protein S3, partial (51%)
Length = 800
Score = 29.6 bits (65), Expect = 0.79
Identities = 23/64 (35%), Positives = 33/64 (50%), Gaps = 1/64 (1%)
Frame = +3
Query: 66 VMESGAKGCEVIVSGKLR-AQRAKSMKFKDGYMISSGQPVKDYIDSAVRHVLLRQGVLGI 124
VM+ G +G + SG+L A+ A++ K G +S ID A V R G+LG+
Sbjct: 606 VMKKGVEGIRICCSGRLEGAEIARTECGKYGK--TSCNVFNQKIDYASAEVSTRYGILGV 779
Query: 125 KVKI 128
KV I
Sbjct: 780 KVWI 791
>TC225121 UP|G3PA_PEA (P12858) Glyceraldehyde-3-phosphate dehydrogenase A,
chloroplast precursor (NADP-dependent
glyceraldehydephosphate dehydrogenase subunit A) ,
partial (8%)
Length = 372
Score = 29.6 bits (65), Expect = 0.79
Identities = 15/19 (78%), Positives = 17/19 (88%), Gaps = 1/19 (5%)
Frame = +2
Query: 157 EEYIRPAAVVA-NDIEVPV 174
EEYI+PA V+A NDIEVPV
Sbjct: 95 EEYIQPAPVLAANDIEVPV 151
>TC222250 similar to UP|Q7SH02 (Q7SH02) Probable AMP deaminase, partial (10%)
Length = 1059
Score = 26.9 bits (58), Expect = 5.1
Identities = 18/70 (25%), Positives = 31/70 (43%), Gaps = 6/70 (8%)
Frame = +2
Query: 8 ELTSVVQKRFKFPENSVELYAEKVNNRGLCAIAQAE----SLRY--KLLGGLAVRRACYG 61
E F+ + + +YA K + L +A + + Y K++ VR +CY
Sbjct: 512 EPVEATSHHFRMEDGVIHVYASKTDTEELFPVASSTRFFTDMHYILKVMSIGNVRTSCYH 691
Query: 62 VLRFVMESGA 71
LRF+ E G+
Sbjct: 692 RLRFLEEIGS 721
>BI427150
Length = 422
Score = 26.6 bits (57), Expect = 6.7
Identities = 10/22 (45%), Positives = 14/22 (63%)
Frame = +3
Query: 119 QGVLGIKVKIMLDWDPKGKQGP 140
Q +L +K+ LDWDP+ Q P
Sbjct: 105 QSLLRLKISSFLDWDPETLQTP 170
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.319 0.138 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,251,976
Number of Sequences: 63676
Number of extensions: 71611
Number of successful extensions: 359
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 352
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 354
length of query: 178
length of database: 12,639,632
effective HSP length: 91
effective length of query: 87
effective length of database: 6,845,116
effective search space: 595525092
effective search space used: 595525092
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 56 (26.2 bits)
Medicago: description of AC146759.10


