

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146751.3 + phase: 1 /pseudo
(439 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
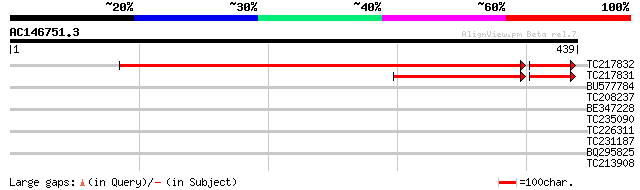
Sequences producing significant alignments: (bits) Value
TC217832 373 e-117
TC217831 128 1e-43
BU577784 30 2.8
TC208237 similar to UP|Q711Q9 (Q711Q9) Allene oxide cyclase prec... 29 3.7
BE347228 29 4.8
TC235090 weakly similar to UP|Q91S26 (Q91S26) Neurofilament trip... 29 4.8
TC226311 similar to UP|Q84TR6 (Q84TR6) Subtilase, partial (18%) 28 6.3
TC231187 similar to UP|Q9LV37 (Q9LV37) MAP kinase, partial (6%) 28 6.3
BQ295825 28 8.2
TC213908 similar to GB|CAA54498.1|602891|SCIILDNA YBL0422 {Sacch... 28 8.2
>TC217832
Length = 1381
Score = 373 bits (957), Expect(2) = e-117
Identities = 192/314 (61%), Positives = 227/314 (72%)
Frame = +3
Query: 86 GRE*CCSNG**LHH*VL*YCF*TN*WVGCAI*HD*NCDWD*AFCDINSC*LFNTFKSFGP 145
GRE*CC +G *LH+ +* C *++ WVG A+*HD* CDW *AFCDI S *LF TF+ F
Sbjct: 3 GRE*CCWDGG*LHYEDI*DCV*SDRWVGSAV*HD*ICDWH*AFCDIQSG*LFGTFEGFDD 182
Query: 146 TFTIHRKVFEFEKSGQEP*MESNL*GTRFTFPRACDHA*RSG*EGKRVCF*RHRCSSTYF 205
+ + R+V E E G EP*ME +L F RACDH+ G*EGKRVC R+ C S F
Sbjct: 183 SLAVCREVLELENPG*EP*MEPDLCRG*LDFARACDHSRGFG*EGKRVCLQRNGCVSADF 362
Query: 206 R**YNCAWKQYKIYVIHC*FHSLWCCRSFQFCVSNNGVYLGFVYSHYIRVWWSN*TSDAY 265
* NCAWKQYK+YV+H FH+LW R FQFCV +G+YLGFV H+IR WWS+ TSD +
Sbjct: 363 HQ**NCAWKQYKVYVLHREFHNLWSRRGFQFCVPVDGIYLGFVLPHHIREWWSDRTSDVH 542
Query: 266 ASNFNLH*S*MC*CFG*SDFRCSFGYR*DCIFSRMSYMAFV*VMQNSFLVYVNCTCIH*S 325
ASNF LH* MC* FG*S FR FG+ *DCIFSR+SY+AFV*V QN+FLV+VNC CIH S
Sbjct: 543 ASNFKLH*EQMC*GFG*SYFRSPFGHC*DCIFSRVSYLAFV*VEQNTFLVHVNCACIHKS 722
Query: 326 FAANISILACDDSGSFATSDGRQIHTGRFLGCYSPFPYGLWCVRDSGRCSWK*CLSYWTQ 385
ANISILAC++ S ATS GRQIH G + YSPF +GLWC+ DS R + +*C+SYW +
Sbjct: 723 SPANISILACNNPSSLATSAGRQIHNGHCVVYYSPFSHGLWCIGDSRRRAGE*CISYWVE 902
Query: 386 HHWGNDIVPISS*G 399
HHWGNDIV I S G
Sbjct: 903 HHWGNDIVSICSGG 944
Score = 66.6 bits (161), Expect(2) = e-117
Identities = 31/36 (86%), Positives = 36/36 (99%)
Frame = +1
Query: 403 LQGAIMGPLITTVMIALKDLYAEFVLEEPKDRAKKK 438
L+GAIMGPLITTVMIALKDLYAEFVL+EPKD++K+K
Sbjct: 937 LEGAIMGPLITTVMIALKDLYAEFVLQEPKDKSKQK 1044
>TC217831
Length = 971
Score = 128 bits (322), Expect(2) = 1e-43
Identities = 64/102 (62%), Positives = 78/102 (75%)
Frame = +1
Query: 298 SRMSYMAFV*VMQNSFLVYVNCTCIH*SFAANISILACDDSGSFATSDGRQIHTGRFLGC 357
SR+SY+AFV*V QN+FLV+VN CIH S ANISILAC++ S ATS GRQIH G +
Sbjct: 1 SRVSYLAFV*VEQNTFLVHVNRACIHKSSPANISILACNNPSSGATSVGRQIHNGHCVVY 180
Query: 358 YSPFPYGLWCVRDSGRCSWK*CLSYWTQHHWGNDIVPISS*G 399
SPF +GL C+RDS R + +*C+S+W +HHWGNDIVPI S G
Sbjct: 181 CSPFSHGLRCIRDSRRRAGE*CISHWVEHHWGNDIVPICSGG 306
Score = 66.6 bits (161), Expect(2) = 1e-43
Identities = 31/36 (86%), Positives = 36/36 (99%)
Frame = +2
Query: 403 LQGAIMGPLITTVMIALKDLYAEFVLEEPKDRAKKK 438
L+GAIMGPLITTVMIALKDLYAEFVL+EPKD++K+K
Sbjct: 299 LEGAIMGPLITTVMIALKDLYAEFVLQEPKDKSKQK 406
>BU577784
Length = 352
Score = 29.6 bits (65), Expect = 2.8
Identities = 18/53 (33%), Positives = 29/53 (53%), Gaps = 1/53 (1%)
Frame = +2
Query: 374 CSWK*CLSYWTQHHWGNDIVPISS*GYVVLQGAIMGPLITT-VMIALKDLYAE 425
C W S + H WG+ ++ +SS VVLQ +GP+ + ++ KD+Y E
Sbjct: 197 CMWCLLCSATSFHSWGSGLLELSSTSLVVLQ*T-LGPIFSN*GIMREKDIYLE 352
>TC208237 similar to UP|Q711Q9 (Q711Q9) Allene oxide cyclase precursor ,
partial (49%)
Length = 631
Score = 29.3 bits (64), Expect = 3.7
Identities = 10/32 (31%), Positives = 19/32 (59%)
Frame = +1
Query: 285 FRCSFGYR*DCIFSRMSYMAFV*VMQNSFLVY 316
F C +GYR C+F+++ + + +MQ S +
Sbjct: 316 FSCCYGYRASCLFAKLHQLIMIMMMQLSLFCF 411
>BE347228
Length = 469
Score = 28.9 bits (63), Expect = 4.8
Identities = 11/22 (50%), Positives = 16/22 (72%)
Frame = -2
Query: 13 EKCN*CFFYKKGGEEVEYYCCN 34
E+C+*C F +GG EV ++ CN
Sbjct: 270 ERCS*CHFAIEGGVEVFFFFCN 205
>TC235090 weakly similar to UP|Q91S26 (Q91S26) Neurofilament triplet H1-like
protein, partial (11%)
Length = 448
Score = 28.9 bits (63), Expect = 4.8
Identities = 18/57 (31%), Positives = 21/57 (36%), Gaps = 9/57 (15%)
Frame = -1
Query: 34 NWAYCCYD-CRIFDWGDFLF--------V*DWC*RKRCRCFS*VTC*GK*LF*ENWC 81
+W+ C CR FDW F + DWC RCR F NWC
Sbjct: 325 SWSGCTLSRCRTFDWCTFNWCGTFNWCRTFDWCTFSRCRTF-------------NWC 194
>TC226311 similar to UP|Q84TR6 (Q84TR6) Subtilase, partial (18%)
Length = 943
Score = 28.5 bits (62), Expect = 6.3
Identities = 10/25 (40%), Positives = 14/25 (56%)
Frame = +1
Query: 213 WKQYKIYVIHC*FHSLWCCRSFQFC 237
W Q+ ++ IH WCC SF+ C
Sbjct: 394 WLQFAVWNIHGMPSCFWCCCSFESC 468
>TC231187 similar to UP|Q9LV37 (Q9LV37) MAP kinase, partial (6%)
Length = 717
Score = 28.5 bits (62), Expect = 6.3
Identities = 7/16 (43%), Positives = 11/16 (68%)
Frame = +2
Query: 30 YYCCNWAYCCYDCRIF 45
YYCCN+ +C C ++
Sbjct: 395 YYCCNYLFCVRKCNVY 442
>BQ295825
Length = 348
Score = 28.1 bits (61), Expect = 8.2
Identities = 10/28 (35%), Positives = 19/28 (67%)
Frame = -1
Query: 381 SYWTQHHWGNDIVPISS*GYVVLQGAIM 408
+Y Q+HWG D+V + + ++ +GAI+
Sbjct: 279 NYKVQYHWGEDLVAVKTEFHIWKRGAII 196
>TC213908 similar to GB|CAA54498.1|602891|SCIILDNA YBL0422 {Saccharomyces
cerevisiae;} , partial (5%)
Length = 514
Score = 28.1 bits (61), Expect = 8.2
Identities = 8/15 (53%), Positives = 10/15 (66%)
Frame = -3
Query: 26 EEVEYYCCNWAYCCY 40
E YYCC + +CCY
Sbjct: 221 EHSYYYCCYYCFCCY 177
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.369 0.166 0.734
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 25,521,952
Number of Sequences: 63676
Number of extensions: 471695
Number of successful extensions: 7725
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 4322
Number of HSP's successfully gapped in prelim test: 331
Number of HSP's that attempted gapping in prelim test: 3115
Number of HSP's gapped (non-prelim): 5028
length of query: 439
length of database: 12,639,632
effective HSP length: 100
effective length of query: 339
effective length of database: 6,272,032
effective search space: 2126218848
effective search space used: 2126218848
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.8 bits)
S2: 60 (27.7 bits)
Medicago: description of AC146751.3


