

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146751.11 - phase: 0
(399 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
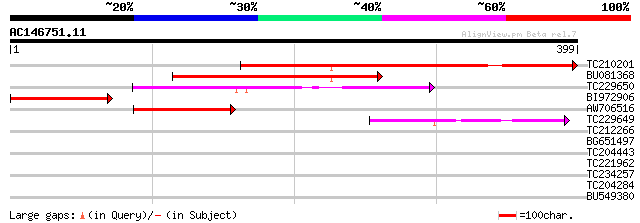
Score E
Sequences producing significant alignments: (bits) Value
TC210201 similar to UP|Q39216 (Q39216) RNA polymerase subunit (I... 336 1e-92
BU081368 206 1e-53
TC229650 similar to UP|RP3A_ARATH (Q39211) DNA-directed RNA poly... 112 2e-25
BI972906 weakly similar to PIR|H96633|H966 RNA polymerase subuni... 82 3e-16
AW706516 similar to SP|Q39211|RP3A DNA-directed RNA polymerase I... 70 1e-12
TC229649 similar to UP|Q8LAJ1 (Q8LAJ1) DNA-directed RNA polymera... 70 2e-12
TC212266 similar to UP|Q8LAJ1 (Q8LAJ1) DNA-directed RNA polymera... 37 0.021
BG651497 similar to GP|28854439|gb tail tape meausure protein {P... 30 1.9
TC204443 homologue to UP|Q8SBC6 (Q8SBC6) Transcription factor LI... 30 1.9
TC221962 similar to GB|BAA98195.1|8978342|AP002030 short chain a... 30 2.5
TC234257 29 3.3
TC204284 similar to UP|Q9ZTK7 (Q9ZTK7) CONSTANS-like protein 2, ... 28 7.3
BU549380 similar to GP|16323057|gb| AT5g57660/MRI1_1 {Arabidopsi... 28 7.3
>TC210201 similar to UP|Q39216 (Q39216) RNA polymerase subunit (Isoform B),
partial (55%)
Length = 863
Score = 336 bits (861), Expect = 1e-92
Identities = 171/240 (71%), Positives = 193/240 (80%), Gaps = 3/240 (1%)
Frame = +1
Query: 163 DDAGGENNEKNTIVFKQHVRCEKGQPRLTVKSDTLKWLPNGSELIAEGTKSAADPAPKTF 222
++AG + NEKNTIVFK HVRC+ GQPR+TVKSD LKWLPNGSEL E K A PKTF
Sbjct: 55 ENAGDDKNEKNTIVFKLHVRCQVGQPRITVKSDKLKWLPNGSELPCEDVKPNAGSKPKTF 234
Query: 223 TTF--NQKSLPKFSKDPAPYNL-DVIVAKLGPGQEIELEAHAVKGIGKTHAKWSPVATAW 279
T+F +Q SLP+FS + D+I+AKLGPGQEIELEAHAVKG GKTHAKWSPVAT W
Sbjct: 235 TSFTCSQDSLPEFSNNSIGLTYSDIILAKLGPGQEIELEAHAVKGTGKTHAKWSPVATTW 414
Query: 280 YRMLPEVVLTKDVKDELAEELVSKCPAKVFDIEDIGRGRKKAVVKNARACTLCRECIREA 339
YRMLPEVVL KDV+DELAEEL +KCP VFDIEDIG+G+++A V R CTLCRECIR
Sbjct: 415 YRMLPEVVLLKDVEDELAEELKNKCPVNVFDIEDIGKGKRRAKVARPRDCTLCRECIR-- 588
Query: 340 DEGEEGGEGEKWTDRVSLRRVKDHFIFTIESTGAIPPDVLFTEAVKILEDKCERVITELS 399
G++W DRVSLRRVK+HFIFTIESTGA+PP+VLFTEAVKILEDKCERVITELS
Sbjct: 589 -------GGKEWEDRVSLRRVKNHFIFTIESTGALPPEVLFTEAVKILEDKCERVITELS 747
>BU081368
Length = 456
Score = 206 bits (525), Expect = 1e-53
Identities = 109/151 (72%), Positives = 122/151 (80%), Gaps = 3/151 (1%)
Frame = +3
Query: 115 RILIAEVPTMAIERVYIANNTSLVQDEVLSHRLGLIPIDADPKLFEYPDDAGGENNEKNT 174
RILIAEVPTMAIERVYIANNTS+VQDEVLSHRLGLIPI ADP+LFEYP++AG + NEKNT
Sbjct: 3 RILIAEVPTMAIERVYIANNTSVVQDEVLSHRLGLIPIRADPRLFEYPENAGDDKNEKNT 182
Query: 175 IVFKQHVRCEKGQPRLTVKSDTLKWLPNGSELIAEGTKSAADPAPKTFTTF--NQKSLPK 232
IVFK RC+ GQPR+TVKS LKWLPNGSEL E K A PKTFT+F +Q SLP+
Sbjct: 183 IVFKLLFRCQVGQPRITVKSHKLKWLPNGSELPCEDVKPNAGSKPKTFTSFTCSQDSLPE 362
Query: 233 FSKDPAPYNL-DVIVAKLGPGQEIELEAHAV 262
FS + D+I+AKLGPGQEIELEAHAV
Sbjct: 363 FSNNSIGLTYSDIILAKLGPGQEIELEAHAV 455
>TC229650 similar to UP|RP3A_ARATH (Q39211) DNA-directed RNA polymerase II 36
kDa polypeptide A (RNA polymerase II subunit 3) ,
partial (60%)
Length = 726
Score = 112 bits (281), Expect = 2e-25
Identities = 73/218 (33%), Positives = 114/218 (51%), Gaps = 5/218 (2%)
Frame = +1
Query: 87 KVEVKRITDEEMEFDMIGIDPAVANAFRRILIAEVPTMAIERVYIANNTSLVQDEVLSHR 146
+V+++ + D+ +F++ D ++ANA RR++IAEVPT+AI+ V I N+S++ DE ++HR
Sbjct: 124 RVKIRELKDDYAKFELRDTDASIANALRRVMIAEVPTVAIDLVEIEVNSSVLNDEFIAHR 303
Query: 147 LGLIPIDADPKL---FEYPDDA--GGENNEKNTIVFKQHVRCEKGQPRLTVKSDTLKWLP 201
LGLIP+ ++ + F DA G E ++ F V+C Q D P
Sbjct: 304 LGLIPLTSERAMSMRFSRDCDACDGDGQCEFCSVEFHLRVKCMTDQTLDVTSKDLYSSDP 483
Query: 202 NGSELIAEGTKSAADPAPKTFTTFNQKSLPKFSKDPAPYNLDVIVAKLGPGQEIELEAHA 261
S + +DP+ + D N +I+ KL GQE++L A A
Sbjct: 484 TVSPV------DFSDPS---------------ATDSPDNNRGIIIVKLRRGQELKLRAIA 600
Query: 262 VKGIGKTHAKWSPVATAWYRMLPEVVLTKDVKDELAEE 299
KGIGK HAKWSP AT + PE+ + +D+ + L E
Sbjct: 601 RKGIGKDHAKWSPAATVTFMYEPEIHINEDLMETLTLE 714
>BI972906 weakly similar to PIR|H96633|H966 RNA polymerase subunit
10595-12672 [imported] - Arabidopsis thaliana, partial
(9%)
Length = 286
Score = 82.4 bits (202), Expect = 3e-16
Identities = 40/72 (55%), Positives = 50/72 (68%)
Frame = +2
Query: 1 MSSSGDESMASEIDEQEQNGGRGIPDYIMNLEDVPSTLPTHLELQKTRVFCNLDAPQHTD 60
MSSS S +SE + + D+I++L++VPS LP HLEL KTRV CN DAP HTD
Sbjct: 71 MSSSSSSSSSSEDESMSEQSNELGLDFILSLDNVPSKLPPHLELFKTRVLCNNDAPVHTD 250
Query: 61 TIQYSGAYAALG 72
T+QYSGAYA +G
Sbjct: 251 TVQYSGAYAIMG 286
>AW706516 similar to SP|Q39211|RP3A DNA-directed RNA polymerase II 36 kDa
polypeptide A (EC 2.7.7.6) (RNA polymerase II subunit
3)., partial (39%)
Length = 463
Score = 70.5 bits (171), Expect = 1e-12
Identities = 31/72 (43%), Positives = 53/72 (73%)
Frame = +2
Query: 88 VEVKRITDEEMEFDMIGIDPAVANAFRRILIAEVPTMAIERVYIANNTSLVQDEVLSHRL 147
V+++ + D+ +F++ D ++ANA RR++IAEVPT+AI+ V I N+S++ DE + HRL
Sbjct: 35 VKIRELKDDYAKFELRDTDASIANALRRVMIAEVPTVAIDLVEIEVNSSVLNDEFICHRL 214
Query: 148 GLIPIDADPKLF 159
GLIP+ ++ +F
Sbjct: 215 GLIPLTSERAMF 250
>TC229649 similar to UP|Q8LAJ1 (Q8LAJ1) DNA-directed RNA polymerase II, third
largest subunit, partial (50%)
Length = 659
Score = 70.1 bits (170), Expect = 2e-12
Identities = 46/145 (31%), Positives = 75/145 (51%), Gaps = 4/145 (2%)
Frame = +1
Query: 254 EIELEAHAVKGIGKTHAKWSPVATAWYRMLPEVVLTKDVKDELA----EELVSKCPAKVF 309
E++ A KGI HAKWS AT + PE+ + +D+ + L E V P +VF
Sbjct: 1 ELKXRAIXXKGIXXDHAKWSXAATVTFMYEPEIHINEDLMETLTLEEKREWVDSSPTRVF 180
Query: 310 DIEDIGRGRKKAVVKNARACTLCRECIREADEGEEGGEGEKWTDRVSLRRVKDHFIFTIE 369
+I+ + ++ +V +A A T E +++A+ + G V + +D FIFT+E
Sbjct: 181 EIDPV---TQQVMVVDAEAYTYDDEVLKKAEAMGKPG-------LVEIIARQDSFIFTVE 330
Query: 370 STGAIPPDVLFTEAVKILEDKCERV 394
STGA+ L A++IL+ K + V
Sbjct: 331 STGAVKASQLVLNAIEILKQKLDAV 405
>TC212266 similar to UP|Q8LAJ1 (Q8LAJ1) DNA-directed RNA polymerase II, third
largest subunit, partial (28%)
Length = 1061
Score = 36.6 bits (83), Expect = 0.021
Identities = 24/73 (32%), Positives = 39/73 (52%)
Frame = +2
Query: 322 VVKNARACTLCRECIREADEGEEGGEGEKWTDRVSLRRVKDHFIFTIESTGAIPPDVLFT 381
+V +A A T E +++A+ + G V + +D FIFT+ESTGA+ L
Sbjct: 551 MVVDAEAYTYDDEVLKKAEAMGKPG-------LVEIIARQDSFIFTVESTGAVKASQLVL 709
Query: 382 EAVKILEDKCERV 394
A++IL+ K + V
Sbjct: 710 NAIEILKQKLDAV 748
>BG651497 similar to GP|28854439|gb tail tape meausure protein {Pseudomonas
syringae pv. tomato str. DC3000}, partial (2%)
Length = 386
Score = 30.0 bits (66), Expect = 1.9
Identities = 18/50 (36%), Positives = 26/50 (52%), Gaps = 2/50 (4%)
Frame = +1
Query: 297 AEELVSKCPA--KVFDIEDIGRGRKKAVVKNARACTLCRECIREADEGEE 344
AEELV +C + + +D++D G CREC +EA+E EE
Sbjct: 106 AEELVMRCLSCEEEYDVDDSGT---------------CRECYQEANEAEE 210
>TC204443 homologue to UP|Q8SBC6 (Q8SBC6) Transcription factor LIM, partial
(91%)
Length = 1080
Score = 30.0 bits (66), Expect = 1.9
Identities = 19/67 (28%), Positives = 36/67 (53%)
Frame = -2
Query: 81 NFYQNFKVEVKRITDEEMEFDMIGIDPAVANAFRRILIAEVPTMAIERVYIANNTSLVQD 140
NF+ F+ VK + E ++ + PAV N+F+R+++AE + +A+ SL+
Sbjct: 395 NFWCPFEAFVKTTSSFEQ---LVKVWPAVKNSFKRVIVAE---LECSFAVVASEASLMVH 234
Query: 141 EVLSHRL 147
V+ +L
Sbjct: 233 SVICGQL 213
>TC221962 similar to GB|BAA98195.1|8978342|AP002030 short chain alcohol
dehydrogenase-like {Arabidopsis thaliana;} , partial
(91%)
Length = 1245
Score = 29.6 bits (65), Expect = 2.5
Identities = 15/40 (37%), Positives = 22/40 (54%)
Frame = +2
Query: 190 LTVKSDTLKWLPNGSELIAEGTKSAADPAPKTFTTFNQKS 229
LTV S WLP+ +L+ AA P P +F+T +K+
Sbjct: 119 LTVISTKCIWLPS*IQLLIFHYSQAAQPPPLSFSTIEEKN 238
>TC234257
Length = 771
Score = 29.3 bits (64), Expect = 3.3
Identities = 8/19 (42%), Positives = 13/19 (68%)
Frame = +1
Query: 264 GIGKTHAKWSPVATAWYRM 282
G+GK H +W+P W+R+
Sbjct: 346 GLGKEHRRWTPEPVTWFRL 402
>TC204284 similar to UP|Q9ZTK7 (Q9ZTK7) CONSTANS-like protein 2, partial
(60%)
Length = 1202
Score = 28.1 bits (61), Expect = 7.3
Identities = 20/61 (32%), Positives = 26/61 (41%), Gaps = 7/61 (11%)
Frame = +1
Query: 315 GRGRKKAVVKNARACTLCRECIREADEGEEGGEGEK-------WTDRVSLRRVKDHFIFT 367
G GR+ + R RE R G+E EG + W D S RRV D+ I+
Sbjct: 850 GTGRRGRTESSRRPFVTHREK-RMPKRGQESKEGSRNALMRTLWLDTASFRRVDDYHIYH 1026
Query: 368 I 368
I
Sbjct: 1027I 1029
>BU549380 similar to GP|16323057|gb| AT5g57660/MRI1_1 {Arabidopsis thaliana},
partial (18%)
Length = 657
Score = 28.1 bits (61), Expect = 7.3
Identities = 20/61 (32%), Positives = 26/61 (41%), Gaps = 7/61 (11%)
Frame = -3
Query: 315 GRGRKKAVVKNARACTLCRECIREADEGEEGGEGEK-------WTDRVSLRRVKDHFIFT 367
G GR+ + R RE R G+E EG + W D S RRV D+ I+
Sbjct: 247 GTGRRGRTESSRRPFVTHREK-RMPKRGQESKEGSRNALMRTLWLDTASFRRVDDYHIYH 71
Query: 368 I 368
I
Sbjct: 70 I 68
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.315 0.134 0.389
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,403,385
Number of Sequences: 63676
Number of extensions: 201511
Number of successful extensions: 1008
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 994
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1003
length of query: 399
length of database: 12,639,632
effective HSP length: 99
effective length of query: 300
effective length of database: 6,335,708
effective search space: 1900712400
effective search space used: 1900712400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.5 bits)
S2: 60 (27.7 bits)
Medicago: description of AC146751.11


