

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146747.7 - phase: 0 /pseudo
(305 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
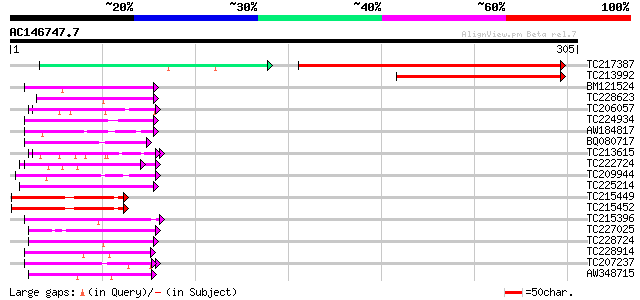
Score E
Sequences producing significant alignments: (bits) Value
TC217387 similar to GB|AAO24564.1|27808568|BT003132 At3g06060 {A... 234 5e-64
TC213992 120 1e-27
BM121524 weakly similar to SP|Q9FPQ6|GP1_C Vegetative cell wall ... 49 3e-06
TC228623 similar to UP|Q9SA21 (Q9SA21) F3O9.2 protein, partial (... 46 2e-05
TC206057 homologue to UP|Q9ZRX1 (Q9ZRX1) Cytosolic chaperonin, d... 46 2e-05
TC224934 similar to UP|Q9SII5 (Q9SII5) Expressed protein (At2g17... 45 3e-05
AW184817 45 4e-05
BQ080717 45 5e-05
TC213615 similar to UP|Q9SUL7 (Q9SUL7) SNF1 like protein kinase ... 44 7e-05
TC222724 weakly similar to UP|Q6VBJ4 (Q6VBJ4) Epa5p, partial (5%) 44 9e-05
TC209944 weakly similar to UP|Q9NP08 (Q9NP08) H6 homeodomain pro... 44 9e-05
TC225214 homologue to PIR|T04582|T04582 ribosomal protein L8, cy... 44 9e-05
TC215449 similar to UP|P93488 (P93488) NAP1Ps, partial (97%) 44 1e-04
TC215452 similar to UP|P93488 (P93488) NAP1Ps, partial (29%) 44 1e-04
TC215396 weakly similar to UP|Q8H9F4 (Q8H9F4) 41 kD chloroplast ... 44 1e-04
TC227025 homologue to UP|Q9ZPA0 (Q9ZPA0) Inosine monophosphate d... 44 1e-04
TC228724 similar to UP|Q9FML2 (Q9FML2) Histone deacetylase, part... 43 2e-04
TC228914 similar to UP|DAH1_MOUSE (Q9QYB2) Dachshund homolog 1 (... 43 2e-04
TC207237 weakly similar to UP|EWS_HUMAN (Q01844) RNA-binding pro... 43 2e-04
AW348715 43 2e-04
>TC217387 similar to GB|AAO24564.1|27808568|BT003132 At3g06060 {Arabidopsis
thaliana;} , partial (92%)
Length = 1545
Score = 234 bits (596), Expect(2) = 5e-64
Identities = 116/144 (80%), Positives = 127/144 (87%)
Frame = +2
Query: 156 ITSWSAYSASKFGLRGLAEALQQEVIGDNIHVSLIFPPDTDTPGLVEENKRKPELTKIIA 215
I + AYSASKFGLRGLAE+LQQEVI DNIHVS+IFPPDTDTPGL EENKR+PELTKII
Sbjct: 716 IYGYVAYSASKFGLRGLAESLQQEVIEDNIHVSMIFPPDTDTPGLAEENKRRPELTKIIT 895
Query: 216 ASSGFMKADEVAQKAFDGIRSGSFIISCNLEGIALSLATSGLSPQRSFLMAFVEVIAAGI 275
ASSG MKADEVAQKA DGI+ G FI+SCN EGIALSLAT+GLSPQRSFLMAF+EV+AAGI
Sbjct: 896 ASSGSMKADEVAQKALDGIKCGDFIVSCNFEGIALSLATAGLSPQRSFLMAFLEVVAAGI 1075
Query: 276 MRIAALCFQWNWYGSIEKWHKQRK 299
+RI AL QW WY SIEK+H QRK
Sbjct: 1076LRIVALGMQWTWYRSIEKYHSQRK 1147
Score = 28.5 bits (62), Expect(2) = 5e-64
Identities = 43/137 (31%), Positives = 52/137 (37%), Gaps = 12/137 (8%)
Frame = +3
Query: 17 STSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPNSKKLRTPS 76
STS S P RS+ T T SS A A S T S S + ++ T S
Sbjct: 243 STSWSVRVP*RSRLRTGTFSSRADPAVSGWRWRTARRRRVRASRSSPGHRTS*RRRATRS 422
Query: 77 NTPPALK*-------GYLMR*RKWLMKLDPLMCCC*IMEC-----FMRWN*VMSSLRLML 124
+P + G R* P CCC* ++RWN*V SS
Sbjct: 423 ASPQGWRWRHSRQTCGTSRR*SARSTMRAPSTCCC*TTASSWRWSWIRWN*VRSSSPWT* 602
Query: 125 I*WVV*I*LKLLCLI*R 141
I W *I + CL *R
Sbjct: 603 ISWER*ISSRPRCLQ*R 653
>TC213992
Length = 804
Score = 120 bits (300), Expect = 1e-27
Identities = 62/91 (68%), Positives = 67/91 (73%)
Frame = +3
Query: 209 ELTKIIAASSGFMKADEVAQKAFDGIRSGSFIISCNLEGIALSLATSGLSPQRSFLMAFV 268
ELTKI ASSG M DEVAQ A DG G FI EGIALSLAT+GL QRSFLMAF+
Sbjct: 3 ELTKIXXASSGSMXXDEVAQXALDGXXCGDFIXXXXFEGIALSLATAGLXXQRSFLMAFL 182
Query: 269 EVIAAGIMRIAALCFQWNWYGSIEKWHKQRK 299
EV+AAGI+RI AL QW WY SIEK+H QRK
Sbjct: 183 EVVAAGILRIVALGMQWTWYRSIEKYHSQRK 275
>BM121524 weakly similar to SP|Q9FPQ6|GP1_C Vegetative cell wall protein gp1
precursor (Hydroxyproline-rich glycoprotein 1)., partial
(10%)
Length = 875
Score = 48.9 bits (115), Expect = 3e-06
Identities = 31/75 (41%), Positives = 42/75 (55%), Gaps = 3/75 (4%)
Frame = +3
Query: 9 SLSSSSPSSTSSSAPAPSR--SQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHD-P 65
S S S PSS+++S+P+P S SP+ T SSP + + A+ P PP S S P
Sbjct: 288 SRSHSPPSSSAASSPSPPAPSSSSPSPTPSSPPS*PSPAASPPLTPPTHFTASSSTRALP 467
Query: 66 SPNSKKLRTPSNTPP 80
SPNS+ TP + PP
Sbjct: 468 SPNSRASVTPPSPPP 512
Score = 32.3 bits (72), Expect = 0.27
Identities = 24/63 (38%), Positives = 33/63 (52%), Gaps = 3/63 (4%)
Frame = +3
Query: 11 SSSSPSSTSSSAPA-PSRSQSPTVTSSS--PAAQAASASPSPTVPPLMAHVSPSWHDPSP 67
SSSSPS T SS P+ PS + SP +T + A+ + A PSP + SP +P
Sbjct: 348 SSSSPSPTPSSPPS*PSPAASPPLTPPTHFTASSSTRALPSPNSRASVTPPSPPPPEPEF 527
Query: 68 NSK 70
+ K
Sbjct: 528 SRK 536
>TC228623 similar to UP|Q9SA21 (Q9SA21) F3O9.2 protein, partial (85%)
Length = 1082
Score = 46.2 bits (108), Expect = 2e-05
Identities = 26/71 (36%), Positives = 35/71 (48%), Gaps = 5/71 (7%)
Frame = -3
Query: 15 PSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSP-----TVPPLMAHVSPSWHDPSPNS 69
PS++S+ P+PS S V+ S P + + S +PSP PPL A + P P P S
Sbjct: 441 PSASSAPPPSPSAPGSTPVSPSCPPSSSTSPAPSPPAPPTVAPPLSARLPPPSFRPLPPS 262
Query: 70 KKLRTPSNTPP 80
R N PP
Sbjct: 261 SSRRVTLNPPP 229
>TC206057 homologue to UP|Q9ZRX1 (Q9ZRX1) Cytosolic chaperonin,
delta-subunit, complete
Length = 1930
Score = 45.8 bits (107), Expect = 2e-05
Identities = 34/88 (38%), Positives = 46/88 (51%), Gaps = 17/88 (19%)
Frame = +3
Query: 11 SSSSPSSTSSSAPAP------------SRSQSP---TVTSSSPAAQAASASPSPT--VPP 53
SSS P SSS+P+P S S +P + +SSSP A +SP+P+ +P
Sbjct: 309 SSSPPPRCSSSSPSPRTPPPATAPLPSSSSLAPSSSSASSSSPTASTPPSSPTPSTRLP* 488
Query: 54 LMAHVSPSWHDPSPNSKKLRTPSNTPPA 81
+ SP W PSP+S TPS +PPA
Sbjct: 489 RPSTFSPPW--PSPSSSPTATPS*SPPA 566
Score = 42.7 bits (99), Expect = 2e-04
Identities = 26/72 (36%), Positives = 36/72 (49%), Gaps = 3/72 (4%)
Frame = +3
Query: 13 SSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPNSKKL 72
S P T SS+P + S SSSP + +S+SPSP PP PS +P+S
Sbjct: 243 SPPPPTRSSSPMTAPPSSTRCRSSSPPPRCSSSSPSPRTPPPATAPLPSSSSLAPSSSSA 422
Query: 73 RTPS---NTPPA 81
+ S +TPP+
Sbjct: 423 SSSSPTASTPPS 458
Score = 38.5 bits (88), Expect = 0.004
Identities = 34/100 (34%), Positives = 46/100 (46%), Gaps = 21/100 (21%)
Frame = +3
Query: 11 SSSSPSS------TSSSAPAPSRSQSPTVTSS-SPAAQAASA-----SPSPTVPPLMAHV 58
SS +PS+ ++ S P PS S SPT T S SP A ++A +P ++P
Sbjct: 456 SSPTPSTRLP*RPSTFSPPWPSPSSSPTATPS*SPPAHPSTARSSANTPRSSLPSPSTPF 635
Query: 59 SPSWHDPSP---------NSKKLRTPSNTPPALK*GYLMR 89
SPSW PSP + + PS TP + K L R
Sbjct: 636 SPSWMPPSPTWSISAMSRS*RNSAAPSTTPSS*KVSSLTR 755
>TC224934 similar to UP|Q9SII5 (Q9SII5) Expressed protein
(At2g17230/T23A1.9), partial (70%)
Length = 953
Score = 45.4 bits (106), Expect = 3e-05
Identities = 26/72 (36%), Positives = 41/72 (56%)
Frame = +3
Query: 9 SLSSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPN 68
+LS S+P+ + ++PAPS S + T TS++P A + ASP T P PS PSP+
Sbjct: 384 ALSPSTPTRPAPTSPAPSPSPASTPTSATPTAHTSPASPCRTSSP-----PPSRPSPSPS 548
Query: 69 SKKLRTPSNTPP 80
+ + ++PP
Sbjct: 549 TTATGST*SSPP 584
Score = 30.0 bits (66), Expect = 1.4
Identities = 20/73 (27%), Positives = 33/73 (44%), Gaps = 6/73 (8%)
Frame = +3
Query: 14 SPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSW------HDPSP 67
S S++S +A P+ S + TSS+ + +A P P +SPS P+P
Sbjct: 255 STSTSSGTASGPNPRSSSSRTSSTQSPTTTTAPPPLPPSPTGGALSPSTPTRPAPTSPAP 434
Query: 68 NSKKLRTPSNTPP 80
+ TP++ P
Sbjct: 435 SPSPASTPTSATP 473
>AW184817
Length = 455
Score = 45.1 bits (105), Expect = 4e-05
Identities = 32/75 (42%), Positives = 44/75 (58%), Gaps = 3/75 (4%)
Frame = +1
Query: 9 SLSSSSPS---STSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDP 65
S+++SSP+ +TS ++PA RS T TSSSP + + S+ PSPT PP SPS
Sbjct: 19 SVAASSPAPTTTTSPTSPATPRSSPSTATSSSPPS-SPSSPPSPTTPP---PTSPSSSTS 186
Query: 66 SPNSKKLRTPSNTPP 80
SP K+ TP + P
Sbjct: 187SPG--KISTP*SPSP 225
Score = 34.3 bits (77), Expect = 0.072
Identities = 22/60 (36%), Positives = 33/60 (54%), Gaps = 2/60 (3%)
Frame = +1
Query: 11 SSSSPSSTSSSAPAPSRSQ--SPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPN 68
+SSSP S+ SS P+P+ SP+ ++SSP + SPSPT S + P+P+
Sbjct: 103 TSSSPPSSPSSPPSPTTPPPTSPSSSTSSPGKISTP*SPSPTTTTSTR**SSTIASPAPH 282
>BQ080717
Length = 426
Score = 44.7 bits (104), Expect = 5e-05
Identities = 29/69 (42%), Positives = 39/69 (56%), Gaps = 1/69 (1%)
Frame = +2
Query: 9 SLSSSSPSSTSSS-APAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSP 67
S SSSS SS SSS +P P+ S SP + S+P+ + S SP+ PP SPS+ P
Sbjct: 149 SPSSSSVSSASSSPSPCPAMSTSPPSSPSAPSTPSPSGSPT---PPTSTSPSPSFRCSKP 319
Query: 68 NSKKLRTPS 76
+ + TPS
Sbjct: 320 SCQSPSTPS 346
Score = 30.8 bits (68), Expect = 0.80
Identities = 19/52 (36%), Positives = 30/52 (57%), Gaps = 2/52 (3%)
Frame = +2
Query: 12 SSSPSSTSSSAPAPS-RSQSPTVTS-SSPAAQAASASPSPTVPPLMAHVSPS 61
S SP+ +S++P+PS R P+ S S+P+A + + + T P L SPS
Sbjct: 257 SGSPTPPTSTSPSPSFRCSKPSCQSPSTPSASCSVKNHTKTTPCLTCSPSPS 412
>TC213615 similar to UP|Q9SUL7 (Q9SUL7) SNF1 like protein kinase
(CBL-interacting protein kinase 4), partial (45%)
Length = 674
Score = 44.3 bits (103), Expect = 7e-05
Identities = 37/89 (41%), Positives = 45/89 (49%), Gaps = 16/89 (17%)
Frame = +3
Query: 11 SSSSP----SSTSSSAPAPSRSQSPTVT--SSSPAAQAASASPSPT----------VPPL 54
SS+SP SS SS A A SRS SP T SSSP + +A+A SPT PP
Sbjct: 327 SSTSPVAASSSPSSPAAAASRSPSPAATSRSSSPLSASATAMASPTETSSHRTSSSTPPA 506
Query: 55 MAHVSPSWHDPSPNSKKLRTPSNTPPALK 83
+ S PS N+ T S+TPPA +
Sbjct: 507 TSRSPTSASPPSRNTS--TTVSSTPPAAR 587
Score = 41.6 bits (96), Expect = 5e-04
Identities = 35/79 (44%), Positives = 41/79 (51%), Gaps = 10/79 (12%)
Frame = +3
Query: 13 SSPSSTSSSAPAP---SRSQSPTVTSSSPA--AQAASASPSPTV-----PPLMAHVSPSW 62
+S ST SS P P S S SP SSSP+ A AAS SPSP PL A + +
Sbjct: 276 TSSKSTRSSPPRPRSTSSSTSPVAASSSPSSPAAAASRSPSPAATSRSSSPLSASAT-AM 452
Query: 63 HDPSPNSKKLRTPSNTPPA 81
P+ S RT S+TPPA
Sbjct: 453 ASPTETSSH-RTSSSTPPA 506
Score = 32.7 bits (73), Expect = 0.21
Identities = 19/37 (51%), Positives = 26/37 (69%), Gaps = 3/37 (8%)
Frame = +3
Query: 11 SSSSPSSTS---SSAPAPSRSQSPTVTSSSPAAQAAS 44
SSS+P +TS +SA PSR+ S TV+S+ PAA+ S
Sbjct: 486 SSSTPPATSRSPTSASPPSRNTSTTVSSTPPAARRPS 596
Score = 32.0 bits (71), Expect = 0.36
Identities = 24/74 (32%), Positives = 36/74 (48%), Gaps = 7/74 (9%)
Frame = +3
Query: 12 SSSPSSTSSSAPA-PSRSQSPTVTSSSPAAQAASASPSPTVPP------LMAHVSPSWHD 64
SSSP S S++A A P+ + S +SS+P A S SP+ PP ++ P+
Sbjct: 417 SSSPLSASATAMASPTETSSHRTSSSTP--PATSRSPTSASPPSRNTSTTVSSTPPAARR 590
Query: 65 PSPNSKKLRTPSNT 78
PSP + + T
Sbjct: 591 PSPRQRSSAASATT 632
Score = 32.0 bits (71), Expect = 0.36
Identities = 20/63 (31%), Positives = 32/63 (50%)
Frame = +3
Query: 14 SPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPNSKKLR 73
SP+S S P + + +PT + S+ ++ S S + P+ A SPS SP + R
Sbjct: 222 SPASCVRSTPCAASTTTPTSSKSTRSSPPRPRSTSSSTSPVAASSSPS----SPAAAASR 389
Query: 74 TPS 76
+PS
Sbjct: 390 SPS 398
>TC222724 weakly similar to UP|Q6VBJ4 (Q6VBJ4) Epa5p, partial (5%)
Length = 714
Score = 43.9 bits (102), Expect = 9e-05
Identities = 31/83 (37%), Positives = 44/83 (52%), Gaps = 7/83 (8%)
Frame = +2
Query: 6 FIFSLSSSSPSSTS--SSAPAPSRSQSPTVTS-----SSPAAQAASASPSPTVPPLMAHV 58
F FS +S+SPSS+S SS +PSRS+SP +S S PA ++S SP+P+ PP
Sbjct: 98 FAFS-NSASPSSSSRGSSRASPSRSRSPPSSSAASSPSPPAPSSSSPSPTPSSPPSSLSP 274
Query: 59 SPSWHDPSPNSKKLRTPSNTPPA 81
+ S P + PP+
Sbjct: 275 AASRPPTPPTHSTASSSKRAPPS 343
Score = 42.7 bits (99), Expect = 2e-04
Identities = 27/68 (39%), Positives = 38/68 (55%), Gaps = 3/68 (4%)
Frame = +2
Query: 9 SLSSSSPSSTSSSAPAPSR--SQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWH-DP 65
S S S PSS+++S+P+P S SP+ T SSP + + A+ P PP + S S P
Sbjct: 161 SRSRSPPSSSAASSPSPPAPSSSSPSPTPSSPPSSLSPAASRPPTPPTHSTASSSKRAPP 340
Query: 66 SPNSKKLR 73
SPN + R
Sbjct: 341 SPNLTRRR 364
>TC209944 weakly similar to UP|Q9NP08 (Q9NP08) H6 homeodomain protein,
partial (5%)
Length = 589
Score = 43.9 bits (102), Expect = 9e-05
Identities = 28/86 (32%), Positives = 44/86 (50%), Gaps = 8/86 (9%)
Frame = +3
Query: 4 AFFIFSLSSSSPSST--------SSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLM 55
A F+FS + SP+S+ ++ P+PS S SP + SP++ S+ PSP P +
Sbjct: 90 AMFVFSAVAQSPASSPEFAAAPPKTATPSPSSSMSPPAAAPSPSSSVVSSPPSP--PTSV 263
Query: 56 AHVSPSWHDPSPNSKKLRTPSNTPPA 81
SPS PS + + +P + PA
Sbjct: 264 PASSPS---PSISPSSISSPPSDAPA 332
>TC225214 homologue to PIR|T04582|T04582 ribosomal protein L8, cytosolic -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(63%)
Length = 527
Score = 43.9 bits (102), Expect = 9e-05
Identities = 22/75 (29%), Positives = 40/75 (53%)
Frame = +1
Query: 6 FIFSLSSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDP 65
F S+ S+ + A R S TS+S A + ++SP+ + P+ A +SP WH
Sbjct: 52 FALSVRVLGRCSSLTLTTARVRRGSAASTSASATATSRASSPTSSTTPVAAPLSPRWHSA 231
Query: 66 SPNSKKLRTPSNTPP 80
+P++ + + S++PP
Sbjct: 232 TPSATRSKRSSSSPP 276
>TC215449 similar to UP|P93488 (P93488) NAP1Ps, partial (97%)
Length = 1495
Score = 43.5 bits (101), Expect = 1e-04
Identities = 31/63 (49%), Positives = 40/63 (63%)
Frame = -2
Query: 2 ADAFFIFSLSSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPS 61
AD FF+ LSSSS SS+SSS+ + S S S SSS ++ ++S+S SP P MA SP
Sbjct: 1164 ADDFFLVLLSSSSSSSSSSSSSSSSSSSS----SSSISSSSSSSSRSPNSSP*MA--SPV 1003
Query: 62 WHD 64
HD
Sbjct: 1002 NHD 994
>TC215452 similar to UP|P93488 (P93488) NAP1Ps, partial (29%)
Length = 609
Score = 43.5 bits (101), Expect = 1e-04
Identities = 31/63 (49%), Positives = 40/63 (63%)
Frame = -3
Query: 2 ADAFFIFSLSSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPS 61
AD FF+ LSSSS SS+SSS+ + S S S SSS ++ ++S+S SP P MA SP
Sbjct: 259 ADDFFLVLLSSSSSSSSSSSSSSSSSSSS----SSSISSSSSSSSRSPNSSP*MA--SPV 98
Query: 62 WHD 64
HD
Sbjct: 97 NHD 89
>TC215396 weakly similar to UP|Q8H9F4 (Q8H9F4) 41 kD chloroplast nucleoid DNA
binding protein (CND41), partial (69%)
Length = 1808
Score = 43.5 bits (101), Expect = 1e-04
Identities = 31/89 (34%), Positives = 42/89 (46%), Gaps = 14/89 (15%)
Frame = +1
Query: 9 SLSSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASAS--------------PSPTVPPL 54
S S+++PS +S+ P SPT + PA AAS S PSP PP
Sbjct: 841 SASAATPSPSSNKPPPYIVKSSPTASPPPPAPPAASPSAPPPLPTSSTPPSPPSPAAPPS 1020
Query: 55 MAHVSPSWHDPSPNSKKLRTPSNTPPALK 83
A SP+ +PNS+ PS PPA++
Sbjct: 1021TASTSPASPWAAPNSQSPPPPS--PPAVQ 1101
Score = 40.0 bits (92), Expect = 0.001
Identities = 29/79 (36%), Positives = 39/79 (48%), Gaps = 4/79 (5%)
Frame = +1
Query: 9 SLSSSSPSSTSSSAPAPSRSQSPTVTSS----SPAAQAASASPSPTVPPLMAHVSPSWHD 64
S S+ P TSS+ P+P +P T+S SP A S SP P PP + +P+
Sbjct: 946 SPSAPPPLPTSSTPPSPPSPAAPPSTASTSPASPWAAPNSQSPPPPSPPAVQSSTPA--P 1119
Query: 65 PSPNSKKLRTPSNTPPALK 83
SP S TP + PP+ K
Sbjct: 1120 *SPASPPPPTPPSAPPSGK 1176
Score = 38.1 bits (87), Expect = 0.005
Identities = 26/77 (33%), Positives = 37/77 (47%), Gaps = 5/77 (6%)
Frame = +1
Query: 9 SLSSSSP-----SSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWH 63
S +S SP SST+SS+ A +++ V + +A AA+ SPS PP S
Sbjct: 736 SAASGSP*PPPTSSTTSSSVAAKTTRASLVAPPASSASAATPSPSSNKPPPYIVKSSPTA 915
Query: 64 DPSPNSKKLRTPSNTPP 80
P P + +PS PP
Sbjct: 916 SPPPPAPPAASPSAPPP 966
Score = 32.0 bits (71), Expect = 0.36
Identities = 23/69 (33%), Positives = 34/69 (48%), Gaps = 2/69 (2%)
Frame = +1
Query: 14 SPSSTSSSAPAPSRSQSPTVTSSSPAAQAASAS--PSPTVPPLMAHVSPSWHDPSPNSKK 71
SP +TS+++ +P + + TSSS AA+ AS P A SPS + P P K
Sbjct: 721 SPLATSAASGSP*PPPTSSTTSSSVAAKTTRASLVAPPASSASAATPSPSSNKPPPYIVK 900
Query: 72 LRTPSNTPP 80
++ PP
Sbjct: 901 SSPTASPPP 927
Score = 28.9 bits (63), Expect = 3.0
Identities = 18/62 (29%), Positives = 28/62 (45%)
Frame = +1
Query: 13 SSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPNSKKL 72
++P+S S P+P P V SS+PA + ++ P PT P + P P +
Sbjct: 1051 AAPNSQSPPPPSP-----PAVQSSTPAP*SPASPPPPTPPSAPPSGKACPNTPPPGNSPY 1215
Query: 73 RT 74
T
Sbjct: 1216 ST 1221
>TC227025 homologue to UP|Q9ZPA0 (Q9ZPA0) Inosine monophosphate
dehydrogenase, complete
Length = 2097
Score = 43.5 bits (101), Expect = 1e-04
Identities = 30/72 (41%), Positives = 41/72 (56%), Gaps = 1/72 (1%)
Frame = +2
Query: 11 SSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASP-SPTVPPLMAHVSPSWHDPSPNS 69
S++SPSST +S P PSR P+ SPAA +S +P SP +PP + ++PS PS +S
Sbjct: 380 SAASPSSTPTSPP-PSRR--PSSAKRSPAASPSSPTPPSPHLPPSWSTMTPSGPPPSFSS 550
Query: 70 KKLRTPSNTPPA 81
L P A
Sbjct: 551 PTLAPPPGNSSA 586
Score = 28.1 bits (61), Expect = 5.2
Identities = 22/69 (31%), Positives = 30/69 (42%), Gaps = 4/69 (5%)
Frame = -3
Query: 15 PSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPT----VPPLMAHVSPSWHDPSPNSK 70
P+ S+ P+P+ SP SSP Q +P P PP + PS PSP ++
Sbjct: 934 PTPQHSTQPSPNAPTSPYRPPSSPP-QPPP*TPRPPPL*PAPPALGIP*PSPPPPSPPNQ 758
Query: 71 KLRTPSNTP 79
PS P
Sbjct: 757 PPHHPSPKP 731
>TC228724 similar to UP|Q9FML2 (Q9FML2) Histone deacetylase, partial (82%)
Length = 1677
Score = 43.1 bits (100), Expect = 2e-04
Identities = 25/74 (33%), Positives = 38/74 (50%), Gaps = 4/74 (5%)
Frame = +2
Query: 11 SSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSP----TVPPLMAHVSPSWHDPS 66
SS++PS+ + + APSR PT +S+P + S+ PSP +PP + S S +
Sbjct: 221 SSTTPSTAAWRSTAPSRPPPPTSAASTPTTTSTSSPPSPPRPSPIPPSLVTSSASTSART 400
Query: 67 PNSKKLRTPSNTPP 80
S +PS PP
Sbjct: 401 APSSTASSPSARPP 442
Score = 32.7 bits (73), Expect = 0.21
Identities = 23/80 (28%), Positives = 38/80 (46%), Gaps = 10/80 (12%)
Frame = +2
Query: 11 SSSSPSSTSSSAPA---------PSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPS 61
++S+P++TS+S+P PS S TS+ A + ++SPS PP PS
Sbjct: 290 AASTPTTTSTSSPPSPPRPSPIPPSLVTSSASTSARTAPSSTASSPSARPPPAAPSAPPS 469
Query: 62 WHD-PSPNSKKLRTPSNTPP 80
P+P S + ++ P
Sbjct: 470 NSTAPTPTSPSIGPAASIMP 529
>TC228914 similar to UP|DAH1_MOUSE (Q9QYB2) Dachshund homolog 1 (Dach1),
partial (3%)
Length = 784
Score = 43.1 bits (100), Expect = 2e-04
Identities = 28/82 (34%), Positives = 42/82 (51%), Gaps = 12/82 (14%)
Frame = +3
Query: 9 SLSSSSPSSTSSSAPAPSRSQSPTVTSSSP-----AAQAASASPSPTVP-------PLMA 56
SL +S S S PAP+ S S T +++SP AA ASA+ +P++P P A
Sbjct: 354 SLGTSMSSPASPPPPAPNASTSSTTSAASPALASSAANTASATTAPSLPSIPSRTTPTTA 533
Query: 57 HVSPSWHDPSPNSKKLRTPSNT 78
+PS +P+ ++ TP T
Sbjct: 534 RSTPSSSNPTSSTSPTATPKRT 599
Score = 37.7 bits (86), Expect = 0.007
Identities = 22/75 (29%), Positives = 41/75 (54%)
Frame = +3
Query: 9 SLSSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPN 68
S ++++ ++T+ S P+ +PT S+P++ ++S SPT P VS P+P+
Sbjct: 456 SAANTASATTAPSLPSIPSRTTPTTARSTPSSSNPTSSTSPTATPKRTRVS----SPTPS 623
Query: 69 SKKLRTPSNTPPALK 83
S T ++PP+ K
Sbjct: 624 SSS--TSRSSPPSPK 662
Score = 35.4 bits (80), Expect = 0.032
Identities = 28/84 (33%), Positives = 35/84 (41%), Gaps = 14/84 (16%)
Frame = +3
Query: 12 SSSPSSTSSSAPAPSRSQSP---------TVTSSSPAAQAASAS-----PSPTVPPLMAH 57
S SP AP P RSQ+P T TS+ P A S+S PSP+ P +
Sbjct: 24 SLSPRWRKPRAPPPHRSQTPTRTTTATRRTTTSTLPPASPRSSSPPSSPPSPSTTPTSSA 203
Query: 58 VSPSWHDPSPNSKKLRTPSNTPPA 81
+ H SK+ TPS A
Sbjct: 204 PGNAPHSSPSASKRRPTPSGPSSA 275
Score = 34.3 bits (77), Expect = 0.072
Identities = 23/71 (32%), Positives = 32/71 (44%)
Frame = +3
Query: 11 SSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPNSK 70
SSS PSS S + P+ S SSP+ A+ P+P+ P A SP S +
Sbjct: 147 SSSPPSSPPSPSTTPTSSAPGNAPHSSPS--ASKRRPTPSGPSSAASTSPKPTSTSSRAA 320
Query: 71 KLRTPSNTPPA 81
++P P A
Sbjct: 321 PSKSPFTWPSA 353
Score = 33.5 bits (75), Expect = 0.12
Identities = 26/76 (34%), Positives = 37/76 (48%), Gaps = 2/76 (2%)
Frame = +3
Query: 9 SLSSSSPSSTS-SSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSP 67
S +S+SP TS SS APS+S ++S + ++ ASP P P + S P+
Sbjct: 270 SAASTSPKPTSTSSRAAPSKSPFTWPSASLGTSMSSPASPPPPAPNASTSSTTSAASPAL 449
Query: 68 NSKKLRTPS-NTPPAL 82
S T S T P+L
Sbjct: 450 ASSAANTASATTAPSL 497
Score = 33.5 bits (75), Expect = 0.12
Identities = 28/79 (35%), Positives = 35/79 (43%), Gaps = 8/79 (10%)
Frame = +3
Query: 11 SSSSPSST-SSSAPAPSRSQSPTVTSSSPA----AQAASASPSPTVPPLMAHVSPS---W 62
S SPS+T +SSAP + SP+ + P + AAS SP PT A S S W
Sbjct: 165 SPPSPSTTPTSSAPGNAPHSSPSASKRRPTPSGPSSAASTSPKPTSTSSRAAPSKSPFTW 344
Query: 63 HDPSPNSKKLRTPSNTPPA 81
S + S PPA
Sbjct: 345 PSASLGTSMSSPASPPPPA 401
Score = 32.3 bits (72), Expect = 0.27
Identities = 17/49 (34%), Positives = 29/49 (58%), Gaps = 2/49 (4%)
Frame = +3
Query: 11 SSSSPSSTSSSAPAPSRSQ--SPTVTSSSPAAQAASASPSPTVPPLMAH 57
SSS+P+S++S P R++ SPT +SSS + + + PT ++H
Sbjct: 546 SSSNPTSSTSPTATPKRTRVSSPTPSSSSTSRSSPPSPKEPTATVTVSH 692
>TC207237 weakly similar to UP|EWS_HUMAN (Q01844) RNA-binding protein EWS
(EWS oncogene) (Ewing sarcoma breakpoint region 1
protein), partial (5%)
Length = 812
Score = 42.7 bits (99), Expect = 2e-04
Identities = 28/76 (36%), Positives = 41/76 (53%), Gaps = 3/76 (3%)
Frame = +3
Query: 9 SLSSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPN 68
S SS++ SS S S+P+PS + +P+ TS+S AS +PSP W PS +
Sbjct: 420 SSSSATSSSRSPSSPSPSSASTPSPTSASLGPTWASRTPSPWTRATSPASPTPW--PSSS 593
Query: 69 SKKLRT---PSNTPPA 81
S +L + PS PP+
Sbjct: 594 SPRLTSPWAPSTAPPS 641
Score = 42.0 bits (97), Expect = 3e-04
Identities = 25/73 (34%), Positives = 41/73 (55%), Gaps = 2/73 (2%)
Frame = +3
Query: 9 SLSSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSW--HDPS 66
S SS PS +++ P P+ S S T +S SP++ + S++ +P+ P A + P+W PS
Sbjct: 366 SPSSPPPSWPTTTLPTPTSSSSATSSSRSPSSPSPSSASTPS--PTSASLGPTWASRTPS 539
Query: 67 PNSKKLRTPSNTP 79
P ++ S TP
Sbjct: 540 PWTRATSPASPTP 578
Score = 36.6 bits (83), Expect = 0.015
Identities = 26/74 (35%), Positives = 37/74 (49%)
Frame = +3
Query: 9 SLSSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPN 68
S SSS+ S+++S + SRS + SS P + + P+PT S S PSP+
Sbjct: 294 SSSSSTRRSSTTSRSSASRSSATNSPSSPPPSWPTTTLPTPTSSSSATSSSRSPSSPSPS 473
Query: 69 SKKLRTPSNTPPAL 82
S TPS T +L
Sbjct: 474 SAS--TPSPTSASL 509
Score = 32.0 bits (71), Expect = 0.36
Identities = 17/37 (45%), Positives = 23/37 (61%)
Frame = +3
Query: 11 SSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASP 47
SSSSP TS AP+ + SP +SSS + ++S SP
Sbjct: 585 SSSSPRLTSPWAPSTAPPSSPATSSSS*PSSSSSGSP 695
>AW348715
Length = 713
Score = 42.7 bits (99), Expect = 2e-04
Identities = 28/75 (37%), Positives = 38/75 (50%), Gaps = 6/75 (8%)
Frame = +2
Query: 11 SSSSPSSTSSSAPAPSRSQSPTVTS--SSPAAQAASASPSPTVPP---LMAHVSPSWHD- 64
SS P+ T+SS PAP R+ P S + P SA+P T P L +SP+ D
Sbjct: 233 SSQPPTCTASSPPAPPRTPVPRAASPPTHPLCSCTSAAPGQTPPKSTLLSPPISPTSEDP 412
Query: 65 PSPNSKKLRTPSNTP 79
P P++ + TPS P
Sbjct: 413 PQPHAPEKCTPSTEP 457
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.337 0.143 0.471
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,219,851
Number of Sequences: 63676
Number of extensions: 305563
Number of successful extensions: 14012
Number of sequences better than 10.0: 1432
Number of HSP's better than 10.0 without gapping: 7702
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 11316
length of query: 305
length of database: 12,639,632
effective HSP length: 97
effective length of query: 208
effective length of database: 6,463,060
effective search space: 1344316480
effective search space used: 1344316480
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.8 bits)
S2: 59 (27.3 bits)
Medicago: description of AC146747.7


