

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146747.6 + phase: 0
(498 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
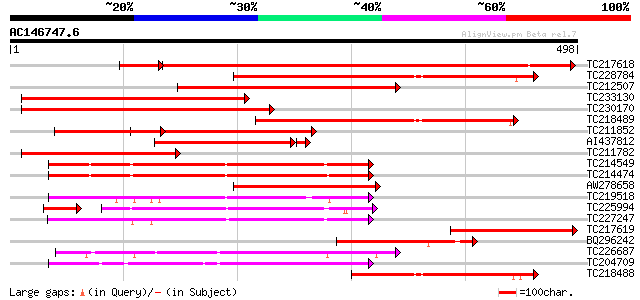
Score E
Sequences producing significant alignments: (bits) Value
TC217618 similar to GB|AAO64868.1|29028978|BT005933 At5g50860 {A... 597 0.0
TC228784 similar to UP|Q9LDC1 (Q9LDC1) CRK1 protein, partial (45%) 384 e-107
TC212507 similar to GB|AAO64868.1|29028978|BT005933 At5g50860 {A... 365 e-101
TC233130 similar to UP|Q9LNN0 (Q9LNN0) F8L10.9 protein, partial ... 332 2e-91
TC230170 similar to UP|Q9LDC1 (Q9LDC1) CRK1 protein, partial (39%) 320 7e-88
TC218489 similar to UP|Q9LDC1 (Q9LDC1) CRK1 protein, partial (35%) 308 5e-84
TC211852 homologue to UP|Q9LDC1 (Q9LDC1) CRK1 protein, partial (... 244 5e-65
AI437812 221 3e-59
TC211782 similar to UP|Q9LDC1 (Q9LDC1) CRK1 protein, partial (25%) 222 3e-58
TC214549 homologue to UP|O82135 (O82135) Cdc2, complete 220 1e-57
TC214474 homologue to UP|CDC2_VIGAC (Q41639) Cell division contr... 219 3e-57
AW278658 202 3e-52
TC219518 similar to UP|Q9FYT8 (Q9FYT8) Cyclin-dependent kinase B... 202 4e-52
TC225994 similar to UP|Q7XUF4 (Q7XUF4) OJ991113_30.14 protein, p... 184 3e-51
TC227247 UP|Q6T2Z8 (Q6T2Z8) Cyclin-dependent kinases CDKB, complete 198 4e-51
TC217619 similar to GB|AAO64868.1|29028978|BT005933 At5g50860 {A... 198 5e-51
BQ296242 similar to GP|29028978|gb| At5g50860 {Arabidopsis thali... 190 1e-48
TC226687 homologue to UP|MMK2_MEDSA (Q40353) Mitogen-activated p... 179 3e-45
TC204709 similar to UP|Q8H0X4 (Q8H0X4) Serine/threonine-protein ... 176 2e-44
TC218488 similar to UP|Q9LDC1 (Q9LDC1) CRK1 protein, partial (21%) 175 5e-44
>TC217618 similar to GB|AAO64868.1|29028978|BT005933 At5g50860 {Arabidopsis
thaliana;} , partial (60%)
Length = 1325
Score = 597 bits (1540), Expect(2) = 0.0
Identities = 292/363 (80%), Positives = 320/363 (87%)
Frame = +1
Query: 135 QVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQSMTS 194
QVKC+MKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATF++P KK MTS
Sbjct: 133 QVKCYMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFFDPKKKHPMTS 312
Query: 195 RVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGS 254
RVVTLWYRPPELLLG+T YGVG+DLWSAGCILAELLAGKP MPGRTEVEQLHKIFKLCGS
Sbjct: 313 RVVTLWYRPPELLLGSTSYGVGVDLWSAGCILAELLAGKPTMPGRTEVEQLHKIFKLCGS 492
Query: 255 PAEEYWRKHKLPNATIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTASAAL 314
P++EYW+K++LPNAT++KPQQPYKR I ETFKDFP SSLPLI++LLAIDPD RGT SAAL
Sbjct: 493 PSDEYWKKYRLPNATLYKPQQPYKRNILETFKDFPSSSLPLIETLLAIDPDDRGTTSAAL 672
Query: 315 NHEFFTTEPYACEPSSLPKYPPSKELDVKMRDEEARRQKALNGKANAVDGAKRVRARERG 374
N EFFTTEPYACEPS+LPKYPP+KELD+K+RDEEARRQKAL+GK NAVDGA+RVR R RG
Sbjct: 673 NSEFFTTEPYACEPSNLPKYPPTKELDIKLRDEEARRQKALSGKTNAVDGARRVRVR*RG 852
Query: 375 RAIPAPEANAEIQTNLDRWRVVTHANAKSKSEKFPPPHQDGAVGYPQDESSKGPVSFGAS 434
AIP PEAN EIQ N+DRWRVVTHANAKSKSEKFPPPHQDGAVGYP D+S+K VSFGA+
Sbjct: 853 LAIPGPEANVEIQNNVDRWRVVTHANAKSKSEKFPPPHQDGAVGYPLDDSNKRAVSFGAT 1032
Query: 435 DTSFSSGTFNVKPSGPTRSHDGTGLHKGTKTKKEESQMASSWKFMRPFKPSTIGLSMDLL 494
+TS +S F+ K G SHD KG KT K +SQMASSWKFMR FK S IG S DLL
Sbjct: 1033ETSSASTIFDSKSWGSVTSHD-AARDKGRKTSKADSQMASSWKFMRSFKLSAIGHSFDLL 1209
Query: 495 FRS 497
FRS
Sbjct: 1210FRS 1218
Score = 69.7 bits (169), Expect(2) = 0.0
Identities = 34/39 (87%), Positives = 36/39 (92%)
Frame = +2
Query: 97 LEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGVKFTEPQ 135
LEGLVTSR+S SLYLVFEYMEHDLAGLSA GVKF+EPQ
Sbjct: 2 LEGLVTSRISSSLYLVFEYMEHDLAGLSAAVGVKFSEPQ 118
>TC228784 similar to UP|Q9LDC1 (Q9LDC1) CRK1 protein, partial (45%)
Length = 1095
Score = 384 bits (985), Expect = e-107
Identities = 195/275 (70%), Positives = 222/275 (79%), Gaps = 7/275 (2%)
Frame = +1
Query: 197 VTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSPA 256
VTLWYRPPELLLGAT Y VG+DLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSP+
Sbjct: 1 VTLWYRPPELLLGATEYSVGVDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSPS 180
Query: 257 EEYWRKHKLPNATIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTASAALNH 316
+EYW+K KLP+ATIFKPQQ YKRCI+ET+KDFPPSSLPL+D+LLAI+PD R TA+AAL+
Sbjct: 181 DEYWKKSKLPHATIFKPQQSYKRCIAETYKDFPPSSLPLMDTLLAINPDERLTATAALHS 360
Query: 317 EFFTTEPYACEPSSLPKYPPSKELDVKMRDEEARRQKALNGKANAVDGAKRVRARER-GR 375
EFFTT+PYACEPSSLPKYPPSKE+D K+RDEEARR +A GKANA DG K+ R RER GR
Sbjct: 361 EFFTTKPYACEPSSLPKYPPSKEMDAKLRDEEARRLRAA-GKANA-DGVKKSRPRERVGR 534
Query: 376 AIPAPEANAEIQTNLDRWRVVTHANAKSKSEKFPPPHQDGAVGYPQDESSKGPVSFGASD 435
I PEANAE+Q N+DR R++TH+NAKSKSEKFPPPHQDGA+GYP S F D
Sbjct: 535 GIAVPEANAELQANIDRRRLITHSNAKSKSEKFPPPHQDGALGYPLGSSHHMDPVFDPPD 714
Query: 436 TSFSSGTF-----NVKP-SGPTRSHDGTGLHKGTK 464
FSS F N++ SGP G G+ + K
Sbjct: 715 VPFSSTNFSQPKANIQTWSGPLVDPSGVGVPRRKK 819
>TC212507 similar to GB|AAO64868.1|29028978|BT005933 At5g50860 {Arabidopsis
thaliana;} , partial (34%)
Length = 590
Score = 365 bits (938), Expect = e-101
Identities = 176/196 (89%), Positives = 185/196 (93%)
Frame = +2
Query: 148 EHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQSMTSRVVTLWYRPPELL 207
EHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFY+P KQ+MTSRVVTLWYRPPELL
Sbjct: 2 EHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYDPKIKQAMTSRVVTLWYRPPELL 181
Query: 208 LGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSPAEEYWRKHKLPN 267
LGAT YGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSP+EEYWRK++LPN
Sbjct: 182 LGATVYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSPSEEYWRKYRLPN 361
Query: 268 ATIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTASAALNHEFFTTEPYACE 327
ATIFKPQQPYKRCI ET+KDFPPSSLPLI++LLAI PD R TAS+ALN EFFTT PYACE
Sbjct: 362 ATIFKPQQPYKRCILETYKDFPPSSLPLIETLLAIYPDDRCTASSALNSEFFTTYPYACE 541
Query: 328 PSSLPKYPPSKELDVK 343
PSSLPKYPP K LDVK
Sbjct: 542 PSSLPKYPPIK*LDVK 589
>TC233130 similar to UP|Q9LNN0 (Q9LNN0) F8L10.9 protein, partial (30%)
Length = 767
Score = 332 bits (851), Expect = 2e-91
Identities = 160/200 (80%), Positives = 180/200 (90%)
Frame = +1
Query: 11 GWPSWLMAVAGEAIGDWTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDN 70
GWPSWL AVAGEAI W PRRA+SFEKL KIGQGTYSNVY+A+DL K+VALKKVRFDN
Sbjct: 166 GWPSWLAAVAGEAIKGWLPRRADSFEKLDKIGQGTYSNVYRARDLEQRKVVALKKVRFDN 345
Query: 71 LEPESVKFMAREILVLRKLDHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGVK 130
LEPESV+FMAREI +LR+LDHPNV+KLEGLVTSRMSCSLYLVFEYMEHDLAGL++ +K
Sbjct: 346 LEPESVRFMAREIHILRRLDHPNVIKLEGLVTSRMSCSLYLVFEYMEHDLAGLASHPKLK 525
Query: 131 FTEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQ 190
FTE QVKC+M+QLL GL+HCH+ GVLHRDI GSNLLIDN GILKIADFGLA+ ++PN+ Q
Sbjct: 526 FTEAQVKCYMQQLLQGLDHCHNCGVLHRDINGSNLLIDNNGILKIADFGLASVFDPNQTQ 705
Query: 191 SMTSRVVTLWYRPPELLLGA 210
+TSRVVTLWYRPPELLLGA
Sbjct: 706 PLTSRVVTLWYRPPELLLGA 765
>TC230170 similar to UP|Q9LDC1 (Q9LDC1) CRK1 protein, partial (39%)
Length = 826
Score = 320 bits (821), Expect = 7e-88
Identities = 162/223 (72%), Positives = 180/223 (80%), Gaps = 1/223 (0%)
Frame = +1
Query: 11 GWPSWLMAVAGEAIGDWTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDN 70
GWPSWL VAGEAI TPRRA++FEK+ KIGQGTYSNVYKA+D +TGKIVALKKVRFDN
Sbjct: 124 GWPSWLSKVAGEAINGLTPRRADTFEKIDKIGQGTYSNVYKARDTLTGKIVALKKVRFDN 303
Query: 71 LEPESVKFMAREILVLRKLDHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGVK 130
LEPESVKFMAREIL+LR+LDHPNV+KLEGLVTSRMSCSLYLVFEYM HDLAGL+ +K
Sbjct: 304 LEPESVKFMAREILILRRLDHPNVIKLEGLVTSRMSCSLYLVFEYMVHDLAGLATNPAIK 483
Query: 131 FTEPQVKCFMKQLLSGLEHCHSRGVLHRDI-KGSNLLIDNEGILKIADFGLATFYNPNKK 189
FTE QVKC+M QL SGL I +G NLLIDN+GIL I DFGL +F++PN K
Sbjct: 484 FTESQVKCYMHQLFSGLGTLSQPSCFCIXI*RGPNLLIDNDGILXIGDFGLXSFFDPNHK 663
Query: 190 QSMTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAG 232
MTS VTLWY P LLLGAT Y V +DLW+AGCILAELLAG
Sbjct: 664 HPMTSXXVTLWYXXPXLLLGATEYXVRMDLWNAGCILAELLAG 792
>TC218489 similar to UP|Q9LDC1 (Q9LDC1) CRK1 protein, partial (35%)
Length = 763
Score = 308 bits (788), Expect = 5e-84
Identities = 155/234 (66%), Positives = 188/234 (80%), Gaps = 3/234 (1%)
Frame = +2
Query: 217 IDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSPAEEYWRKHKLPNATIFKPQQP 276
IDLWS GCIL ELLAGKPI+PG TEVEQLHKI+KL GSP++EY +K K+ NAT+FKP+ P
Sbjct: 8 IDLWSVGCILGELLAGKPILPGPTEVEQLHKIYKL*GSPSDEYRKKSKMHNATLFKPRHP 187
Query: 277 YKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTASAALNHEFFTTEPYACEPSSLPKYPP 336
YKRCI+ETFKDFPPS+LPLID+LLAIDP R +A+ AL EFFTTEPYAC+PSSLPKYPP
Sbjct: 188 YKRCITETFKDFPPSALPLIDTLLAIDPAERKSATDALRSEFFTTEPYACDPSSLPKYPP 367
Query: 337 SKELDVKMRDEEARRQKALNGKANAVDGAKRVRARER-GRAIPAPEANAEIQTNLDRWRV 395
+KE+D K RD+EARR +A GKA+ VDGAK+ R R+R +A PAPE NAE+Q+N+DR R
Sbjct: 368 TKEMDAKRRDDEARRSRAA-GKAH-VDGAKKHRTRDRAAKAAPAPEGNAELQSNIDRRR* 541
Query: 396 VTHANAKSKSEKFPPPHQDGAVGYPQDESSKGPVSFGASDTSF--SSGTFNVKP 447
+THANAKSKSEK PPPH+DG +G+P S+ SD SF +S TF+ +P
Sbjct: 542 ITHANAKSKSEKLPPPHEDGQLGFPLGSSNHIDPDIDPSDVSFGSTSYTFSKEP 703
>TC211852 homologue to UP|Q9LDC1 (Q9LDC1) CRK1 protein, partial (39%)
Length = 706
Score = 244 bits (624), Expect = 5e-65
Identities = 119/163 (73%), Positives = 135/163 (82%)
Frame = +2
Query: 107 CSLYLVFEYMEHDLAGLSAGQGVKFTEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLL 166
CS+ +++ A S Q ++ QVKC+M QLLSGLEHCH+R VLHRDIKGSNLL
Sbjct: 221 CSIIWSMIWLDLLQAQESGSQSLRLD--QVKCYMHQLLSGLEHCHNRHVLHRDIKGSNLL 394
Query: 167 IDNEGILKIADFGLATFYNPNKKQSMTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCIL 226
IDNEG LKIADFGLA+ ++PN K MTSRVVTLWYRPPELLLGAT Y VG+DLWSAGCIL
Sbjct: 395 IDNEGTLKIADFGLASIFDPNHKHPMTSRVVTLWYRPPELLLGATDYSVGVDLWSAGCIL 574
Query: 227 AELLAGKPIMPGRTEVEQLHKIFKLCGSPAEEYWRKHKLPNAT 269
ELLAGKPIMPGRTEVEQLHKI+KLCGSP++EYW+K KLPNAT
Sbjct: 575 GELLAGKPIMPGRTEVEQLHKIYKLCGSPSDEYWKKSKLPNAT 703
Score = 178 bits (451), Expect = 6e-45
Identities = 85/98 (86%), Positives = 95/98 (96%)
Frame = +1
Query: 40 KIGQGTYSNVYKAKDLVTGKIVALKKVRFDNLEPESVKFMAREILVLRKLDHPNVVKLEG 99
KIGQGTYSNVYKAKD++TGKIVALKKVRFDN EPESVKFMAREIL+LR+LDHPNVVKL+G
Sbjct: 4 KIGQGTYSNVYKAKDMMTGKIVALKKVRFDNWEPESVKFMAREILILRRLDHPNVVKLQG 183
Query: 100 LVTSRMSCSLYLVFEYMEHDLAGLSAGQGVKFTEPQVK 137
LVTSRMSCSLYLVF+YMEHDLAGL+A G++FTEPQV+
Sbjct: 184 LVTSRMSCSLYLVFDYMEHDLAGLAASPGIRFTEPQVR 297
>AI437812
Length = 421
Score = 221 bits (562), Expect(2) = 3e-59
Identities = 103/124 (83%), Positives = 114/124 (91%)
Frame = +1
Query: 128 GVKFTEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPN 187
G+KFTE QVKC+M+QLL GL+HCHS GVLHRDIKGSNLLIDN GILKIADFGLA+F++PN
Sbjct: 10 GLKFTEAQVKCYMQQLLRGLDHCHSCGVLHRDIKGSNLLIDNSGILKIADFGLASFFDPN 189
Query: 188 KKQSMTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHK 247
+ Q +TSRVVTLWYRPPELLLGAT+YG +DLWS GCILAEL AGKPIMPGRTEVEQLHK
Sbjct: 190 QAQPLTSRVVTLWYRPPELLLGATYYGTAVDLWSTGCILAELYAGKPIMPGRTEVEQLHK 369
Query: 248 IFKL 251
IFKL
Sbjct: 370 IFKL 381
Score = 26.6 bits (57), Expect(2) = 3e-59
Identities = 9/12 (75%), Positives = 11/12 (91%)
Frame = +3
Query: 253 GSPAEEYWRKHK 264
GSP+E+YWRK K
Sbjct: 384 GSPSEDYWRKSK 419
>TC211782 similar to UP|Q9LDC1 (Q9LDC1) CRK1 protein, partial (25%)
Length = 482
Score = 222 bits (566), Expect = 3e-58
Identities = 102/140 (72%), Positives = 123/140 (87%)
Frame = +3
Query: 11 GWPSWLMAVAGEAIGDWTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDN 70
GWP+WL +VAGEAI W PR AN+FE+L KIGQGTYS VYKA+D++ K VALKKVRFDN
Sbjct: 63 GWPAWLSSVAGEAIKGWIPRSANTFERLHKIGQGTYSTVYKARDVINQKFVALKKVRFDN 242
Query: 71 LEPESVKFMAREILVLRKLDHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGVK 130
L+PESVKFM REI VLR+LDHPN++KLEGL+TS+MS SLYLVFEYMEHDL GL++ +K
Sbjct: 243 LDPESVKFMTREIHVLRRLDHPNIIKLEGLITSQMSRSLYLVFEYMEHDLTGLASNPDIK 422
Query: 131 FTEPQVKCFMKQLLSGLEHC 150
F+EPQ+KC+M+QLLSGL+HC
Sbjct: 423 FSEPQLKCYMQQLLSGLDHC 482
>TC214549 homologue to UP|O82135 (O82135) Cdc2, complete
Length = 1216
Score = 220 bits (561), Expect = 1e-57
Identities = 126/290 (43%), Positives = 182/290 (62%), Gaps = 5/290 (1%)
Frame = +3
Query: 35 FEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDNLEPESVKFMA-REILVLRKLDHPN 93
+EK+ KIG+GTY VYK +D VT + +ALKK+R + E E V A REI +L+++ H N
Sbjct: 165 YEKVEKIGEGTYGVVYKGRDRVTNETIALKKIRLEQ-EDEGVPSTAIREISLLKEMQHRN 341
Query: 94 VVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGVKFTEP-QVKCFMKQLLSGLEHCHS 152
+V+L+ +V S LYLVFEY++ DL +P QVK F+ Q+L G+ +CHS
Sbjct: 342 IVRLQDVVHDEKS--LYLVFEYLDLDLKKHMDSSPEFAKDPRQVKMFLYQILCGIAYCHS 515
Query: 153 RGVLHRDIKGSNLLIDNE-GILKIADFGLATFYNPNKKQSMTSRVVTLWYRPPELLLGAT 211
VLHRD+K NLLID LK+ADFGLA + + + T VVTLWYR PE+LLG+
Sbjct: 516 HRVLHRDLKPQNLLIDRSTNALKLADFGLARAFGIPVR-TFTHEVVTLWYRAPEILLGSR 692
Query: 212 FYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSPAEEYW-RKHKLPN-AT 269
Y +D+WS GCI AE++ +P+ PG +E+++L KIF++ G+P E+ W LP+ +
Sbjct: 693 QYSTPVDIWSVGCIFAEMVNQRPLFPGDSEIDELFKIFRIMGTPNEDTWPGVTSLPDFKS 872
Query: 270 IFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTASAALNHEFF 319
F QP + + + P+ L L+ S+L +DP +R TA +AL HE+F
Sbjct: 873 AFPKWQP--KDLKNVVPNLEPAGLDLLSSMLYLDPSKRITARSALEHEYF 1016
>TC214474 homologue to UP|CDC2_VIGAC (Q41639) Cell division control protein 2
homolog (p34cdc2) , complete
Length = 1204
Score = 219 bits (557), Expect = 3e-57
Identities = 124/289 (42%), Positives = 181/289 (61%), Gaps = 4/289 (1%)
Frame = +2
Query: 35 FEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDNLEPESVKFMA-REILVLRKLDHPN 93
+EK+ KIG+GTY VYKA+D T + +ALKK+R + E E V A REI +L+++ H N
Sbjct: 89 YEKVEKIGEGTYGVVYKARDRATNETIALKKIRLEQ-EDEGVPSTAIREISLLKEMQHRN 265
Query: 94 VVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGVKFTEP-QVKCFMKQLLSGLEHCHS 152
+V+L+ +V S LYLVFEY++ DL +P QVK F+ Q+L G+ +CHS
Sbjct: 266 IVRLQDVVHSEKR--LYLVFEYLDLDLKKHMDSSPEFVKDPRQVKMFLYQILCGIAYCHS 439
Query: 153 RGVLHRDIKGSNLLIDNE-GILKIADFGLATFYNPNKKQSMTSRVVTLWYRPPELLLGAT 211
VLHRD+K NLLID LK+ADFGLA + + + T VVTLWYR PE+LLG+
Sbjct: 440 HRVLHRDLKPQNLLIDRRTNSLKLADFGLARAFGIPVR-TFTHEVVTLWYRAPEILLGSR 616
Query: 212 FYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSPAEEYW-RKHKLPNATI 270
Y +D+WS GCI AE++ +P+ PG +E+++L KIF++ G+P E+ W LP+
Sbjct: 617 HYSTPVDVWSVGCIFAEMVNRRPLFPGDSEIDELFKIFRILGTPNEDTWPGVTSLPDFKS 796
Query: 271 FKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTASAALNHEFF 319
P+ P K ++ + + L L+ S+L +DP +R TA +A+ HE+F
Sbjct: 797 TFPKWPSKD-LANVVPNLDAAGLNLLSSMLCLDPSKRITARSAVEHEYF 940
>AW278658
Length = 395
Score = 202 bits (514), Expect = 3e-52
Identities = 90/129 (69%), Positives = 110/129 (84%)
Frame = +3
Query: 197 VTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSPA 256
VTLWYRPPELLLG+T YG +DLWS GC+ AELL GKPI+ GRTEVEQLHKIFKLCGSP
Sbjct: 9 VTLWYRPPELLLGSTAYGASVDLWSVGCVFAELLIGKPILQGRTEVEQLHKIFKLCGSPP 188
Query: 257 EEYWRKHKLPNATIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTASAALNH 316
EEYW+K +LP+AT+FKPQQPY C+ ETFKDF SS+ L+ +LL+++P +RGTAS+AL+
Sbjct: 189 EEYWKKTRLPHATLFKPQQPYDSCLRETFKDFHASSVNLLQTLLSVEPSKRGTASSALSL 368
Query: 317 EFFTTEPYA 325
E+F T+PYA
Sbjct: 369 EYFKTKPYA 395
>TC219518 similar to UP|Q9FYT8 (Q9FYT8) Cyclin-dependent kinase B1-2,
complete
Length = 1333
Score = 202 bits (513), Expect = 4e-52
Identities = 121/309 (39%), Positives = 173/309 (55%), Gaps = 24/309 (7%)
Frame = +2
Query: 35 FEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDNLEPESVKFMAREILVLRKLDHP-- 92
+EKL K+G+GTY VYKA++ +G +VALKK R + E RE+ +L+ L
Sbjct: 14 YEKLEKVGEGTYGKVYKAREKASGSLVALKKTRLEMDEEGVPPTALREVSLLQLLSQSIY 193
Query: 93 -----NVVKLEGLVTSRMSCS-------LYLVFEYMEHDLAGL--SAGQGVK---FTEPQ 135
+V ++ + S+ S S LYLVFEY++ DL S +G P
Sbjct: 194 IVRLLSVEHVDKVPKSQKSSSNPLTKPILYLVFEYLDTDLKKFIDSHRKGPNPRPLPPPL 373
Query: 136 VKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLID-NEGILKIADFGLATFYNPNKKQSMTS 194
++ F+ QL G+ HCHS GVLHRD+K NLL+D ++GILKIAD GL + K S T
Sbjct: 374 IQSFLFQLCKGVAHCHSHGVLHRDLKPQNLLLDQHKGILKIADLGLGRAFTVPLK-SYTH 550
Query: 195 RVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGS 254
+VTLWYR PE+LLG+T Y G+D+WS GCI AE++ + + PG +E +QL IFK+ G+
Sbjct: 551 EIVTLWYRAPEVLLGSTHYSTGVDIWSVGCIFAEMVRRQALFPGDSEFQQLIHIFKMLGT 730
Query: 255 PAEEYWRKHKLPNATIFKPQQPYKR----CISETFKDFPPSSLPLIDSLLAIDPDRRGTA 310
P EE W P T + Y R +++ P + L+ +L +P R +A
Sbjct: 731 PTEENW-----PGVTSLRDWHVYPRWEPQSLAKNVPSLGPDGVDLLSKMLKYNPSERISA 895
Query: 311 SAALNHEFF 319
AAL+H +F
Sbjct: 896 KAALDHPYF 922
>TC225994 similar to UP|Q7XUF4 (Q7XUF4) OJ991113_30.14 protein, partial (43%)
Length = 2770
Score = 184 bits (466), Expect(2) = 3e-51
Identities = 107/255 (41%), Positives = 154/255 (59%), Gaps = 12/255 (4%)
Frame = +2
Query: 81 REILVLRKLDHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGVKFTEPQVKCFM 140
REI +L +HP++V ++ +V ++V E+ME+DL GL + F+ ++K +
Sbjct: 788 REINILLSFNHPSIVNVKEVVVDDFD-GTFMVMEHMEYDLKGLMEVKKQPFSMSEIKSLV 964
Query: 141 KQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQSMTSRVVTLW 200
+QLL G+++ H V+HRD+K SN+L++++G LKI DFGL+ Y K T VVTLW
Sbjct: 965 RQLLEGVKYLHDNWVIHRDLKSSNILLNHDGELKICDFGLSRQYGSPLKP-YTPVVVTLW 1141
Query: 201 YRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSPAEEYW 260
YR PELLLGA Y ID+WS GCI+AEL+A +P+ G++E+EQL KIF+ G+P E+ W
Sbjct: 1142 YRAPELLLGAKEYSTSIDMWSVGCIMAELIAKEPLFRGKSELEQLDKIFRTLGTPDEKIW 1321
Query: 261 -RKHKLPNATIFKPQQPYKRCISETFKDFPPSS---LP--------LIDSLLAIDPDRRG 308
KLP A +QP I+ K FP +S LP L+ LL DP++R
Sbjct: 1322 PGLSKLPGAKANFVKQP----INTLRKKFPAASFTGLPVLSELGFDLLKRLLTYDPEKRI 1489
Query: 309 TASAALNHEFFTTEP 323
TA AL H++F P
Sbjct: 1490 TAEDALLHDWFHEAP 1534
Score = 37.0 bits (84), Expect(2) = 3e-51
Identities = 17/34 (50%), Positives = 23/34 (67%)
Frame = +3
Query: 30 RRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVAL 63
R F+ + KI +GTY VYKA+D TG++VAL
Sbjct: 630 RSVCEFKMIKKINEGTYGVVYKARDKKTGELVAL 731
>TC227247 UP|Q6T2Z8 (Q6T2Z8) Cyclin-dependent kinases CDKB, complete
Length = 1235
Score = 198 bits (504), Expect = 4e-51
Identities = 124/296 (41%), Positives = 170/296 (56%), Gaps = 10/296 (3%)
Frame = +2
Query: 34 SFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDNLEPESVKFMAREILVLRKLDH-P 92
+FEKL K+G+GTY VY+A++ TGKIVALKK R E RE+ +LR L P
Sbjct: 74 AFEKLEKVGEGTYGKVYRAREKATGKIVALKKTRLHEDEEGVPPTTLREVSILRMLSRDP 253
Query: 93 NVVKLEGLVTSRMS---CSLYLVFEYMEHDLAGL--SAGQGVKFTEPQ-VKCFMKQLLSG 146
+VV+L + + LYLVFEYM+ DL S Q + PQ +K M QL G
Sbjct: 254 HVVRLMDVKQGQNKEGKTVLYLVFEYMDTDLKKFIRSFRQTGQTVPPQTIKSLMYQLCKG 433
Query: 147 LEHCHSRGVLHRDIKGSNLLIDNEGI-LKIADFGLA-TFYNPNKKQSMTSRVVTLWYRPP 204
+ CH G+LHRD+K NLL+D + + LKIAD GLA F P KK T ++TLWYR P
Sbjct: 434 VAFCHGHGILHRDLKPHNLLMDPKTMMLKIADLGLARAFTVPIKKY--THEILTLWYRAP 607
Query: 205 ELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSPAEEYW-RKH 263
E+LLGAT Y + +D+WS GCI AEL+ + + PG +E++QL IF+L G+P E+ W
Sbjct: 608 EVLLGATHYSMAVDMWSVGCIFAELVTKQALFPGDSELQQLLHIFRLLGTPNEDVWPGVS 787
Query: 264 KLPNATIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTASAALNHEFF 319
KL N + P + +S L L+ +L +P +R +A A+ H +F
Sbjct: 788 KLMNWHEYPQWNP--QSLSTAVPSLDELGLDLLSQMLKYEPSKRISAKKAMEHVYF 949
>TC217619 similar to GB|AAO64868.1|29028978|BT005933 At5g50860 {Arabidopsis
thaliana;} , partial (7%)
Length = 883
Score = 198 bits (503), Expect = 5e-51
Identities = 93/111 (83%), Positives = 98/111 (87%)
Frame = +1
Query: 388 TNLDRWRVVTHANAKSKSEKFPPPHQDGAVGYPQDESSKGPVSFGASDTSFSSGTFNVKP 447
T RWRVVTHANAKSKSEKFPPPHQDGAVGYPQD S+KGPVSFGA DTSFSSG FN KP
Sbjct: 268 TEFQRWRVVTHANAKSKSEKFPPPHQDGAVGYPQDASNKGPVSFGAPDTSFSSGIFNSKP 447
Query: 448 SGPTRSHDGTGLHKGTKTKKEESQMASSWKFMRPFKPSTIGLSMDLLFRSK 498
SG R+H GLH+G KTKK+ESQMASSWKFMRPFKPST+GLSMDLLFRSK
Sbjct: 448 SGTVRNHGAAGLHRGRKTKKDESQMASSWKFMRPFKPSTVGLSMDLLFRSK 600
>BQ296242 similar to GP|29028978|gb| At5g50860 {Arabidopsis thaliana},
partial (16%)
Length = 426
Score = 190 bits (482), Expect = 1e-48
Identities = 99/126 (78%), Positives = 107/126 (84%), Gaps = 2/126 (1%)
Frame = +2
Query: 288 FPPSSLPLIDSLLAIDPDRRGTASAALNHEFFTTEPYACEPSSLPKYPPSKELDVKMRDE 347
FPPSSLPLI++LLAIDP+ RGTASA LN EFFTTEPYACEPSSLPKYPPSKELDVK+RDE
Sbjct: 2 FPPSSLPLIETLLAIDPEDRGTASATLNSEFFTTEPYACEPSSLPKYPPSKELDVKLRDE 181
Query: 348 EARRQKALNGKANAVDGAK--RVRARERGRAIPAPEANAEIQTNLDRWRVVTHANAKSKS 405
EARRQKALNGKA+AVDGAK RVR RERGRA+PAPEANAEIQTNLD V+H +
Sbjct: 182 EARRQKALNGKASAVDGAKKVRVRERERGRAVPAPEANAEIQTNLD----VSHTRSCLCL 349
Query: 406 EKFPPP 411
F PP
Sbjct: 350 SSFLPP 367
>TC226687 homologue to UP|MMK2_MEDSA (Q40353) Mitogen-activated protein
kinase homolog MMK2 , partial (97%)
Length = 1783
Score = 179 bits (454), Expect = 3e-45
Identities = 119/317 (37%), Positives = 170/317 (53%), Gaps = 14/317 (4%)
Frame = +3
Query: 41 IGQGTYSNVYKAKDLVTGKIVALKKV--RFDNLEPESVKFMAREILVLRKLDHPNVVKLE 98
+G+G Y V A + TG+ VA+KK+ FDN K REI +LR +DH N++ ++
Sbjct: 360 VGRGAYGIVCAAVNAETGEEVAIKKIGNAFDNRI--DAKRTLREIKLLRHMDHANIMSIK 533
Query: 99 GLVTSRMSCS---LYLVFEYMEHDLAGLSAGQGVKFTEPQVKCFMKQLLSGLEHCHSRGV 155
++ + +YLV E M+ DL + + T+ + F+ QLL GL++ HS V
Sbjct: 534 DIIRPPQKENFNDVYLVSELMDTDLHQIIRSNQ-QLTDDHCRYFLYQLLRGLKYVHSANV 710
Query: 156 LHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQSMTSRVVTLWYRPPELLLGATFYGV 215
LHRD+K SNLL++ LKIADFGLA ++ MT VVT WYR PELLL + Y
Sbjct: 711 LHRDLKPSNLLLNANCDLKIADFGLAR--TTSETDFMTEYVVTRWYRAPELLLNCSEYTA 884
Query: 216 GIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSPAEEYWRKHKLPNATIFKPQQ 275
ID+WS GCIL E++ +P+ PG+ V QL I +L GSP + + NA + Q
Sbjct: 885 AIDIWSVGCILGEIITRQPLFPGKDYVHQLRLITELIGSPDDASLGFLRSDNARRYVKQL 1064
Query: 276 PY--KRCISETFKDFPPSSLPLIDSLLAIDPDRRGTASAALNHEFFT------TEPYACE 327
P K+ S F P ++ L++ +L DP+RR T AL+H + + EP
Sbjct: 1065PQYPKQNFSARFPTMSPGAVDLLEKMLIFDPNRRITVDEALSHPYMSPLHDINEEPVCTR 1244
Query: 328 PSSLPKYPPS-KELDVK 343
P S PS E D+K
Sbjct: 1245PFSFDFEQPSFTEEDIK 1295
>TC204709 similar to UP|Q8H0X4 (Q8H0X4) Serine/threonine-protein kinase Mak
(Male germ cell-associated kinase)-like protein, partial
(59%)
Length = 2140
Score = 176 bits (446), Expect = 2e-44
Identities = 106/286 (37%), Positives = 166/286 (57%), Gaps = 1/286 (0%)
Frame = +2
Query: 35 FEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDNLEPESVKFMAREILVLRKLDHPNV 94
++ + ++G GT+ +V++A + TG++VA+KK++ E + RE+ LRK++HPN+
Sbjct: 539 YKLIKEVGDGTFGSVWRAINKQTGEVVAIKKMKKKYYSWEECVNL-REVKSLRKMNHPNI 715
Query: 95 VKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGVKFTEPQVKCFMKQLLSGLEHCHSRG 154
VKL+ ++ R S LY VFEYME +L L + F+E +V+ + Q+ GL + H RG
Sbjct: 716 VKLKEVI--RESDILYFVFEYMECNLYQLMKDREKLFSEAEVRNWCFQVFQGLAYMHQRG 889
Query: 155 VLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQSMTSRVVTLWYRPPELLLGATFYG 214
HRD+K NLL+ + +KIADFGLA + + T V T WYR PE+LL + Y
Sbjct: 890 YFHRDLKPENLLVTKD-FIKIADFGLAR--EISSQPPYTEYVSTRWYRAPEVLLQSYMYT 1060
Query: 215 VGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSPAEEYWRKH-KLPNATIFKP 273
+D+W+ G I+AEL + +P+ PG +E ++++KI + G+P E W KL ++
Sbjct: 1061SKVDMWAMGAIMAELFSLRPLFPGASEADEIYKICGVIGNPTFESWADGLKLARDINYQF 1240
Query: 274 QQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTASAALNHEFF 319
Q +S ++ LI SL + DP +R TAS AL H FF
Sbjct: 1241PQLAGVHLSALIPSASDDAISLITSLCSWDPCKRPTASEALQHPFF 1378
>TC218488 similar to UP|Q9LDC1 (Q9LDC1) CRK1 protein, partial (21%)
Length = 581
Score = 175 bits (443), Expect = 5e-44
Identities = 99/171 (57%), Positives = 118/171 (68%), Gaps = 7/171 (4%)
Frame = +2
Query: 301 AIDPDRRGTASAALNHEFFTTEPYACEPSSLPKYPPSKELDVKMRDEEARRQKALNGKAN 360
AIDP R TAS AL EFFTTEPYAC+PSSLPKYPPSKE+D K RD+E RR +A GKA
Sbjct: 2 AIDPVERKTASDALRSEFFTTEPYACDPSSLPKYPPSKEMDAKQRDDEMRRLRAA-GKAQ 178
Query: 361 AVDGAKRVRARERG-RAIPAPEANAEIQTNLDRWRVVTHANAKSKSEKFPPPHQDGAVGY 419
A DG K+ R R R +A PAPEANAE+Q+N+DR R++THANAKSKSEKFPPPHQDG VG+
Sbjct: 179 A-DGPKKHRTRNRAAKAFPAPEANAELQSNIDRRRLITHANAKSKSEKFPPPHQDGQVGF 355
Query: 420 PQDESSKGPVSFGASDTSFSSG--TFNVKP----SGPTRSHDGTGLHKGTK 464
P S +D SF+S T++ +P SGP + G+ K K
Sbjct: 356 PLGSSHHIDPDTVPTDVSFTSTSYTYSKEPFQAWSGPIGNAADIGVPKRKK 508
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.317 0.134 0.403
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 23,000,367
Number of Sequences: 63676
Number of extensions: 332609
Number of successful extensions: 2764
Number of sequences better than 10.0: 746
Number of HSP's better than 10.0 without gapping: 2428
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2467
length of query: 498
length of database: 12,639,632
effective HSP length: 101
effective length of query: 397
effective length of database: 6,208,356
effective search space: 2464717332
effective search space used: 2464717332
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 61 (28.1 bits)
Medicago: description of AC146747.6


