

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146723.12 + phase: 0
(172 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
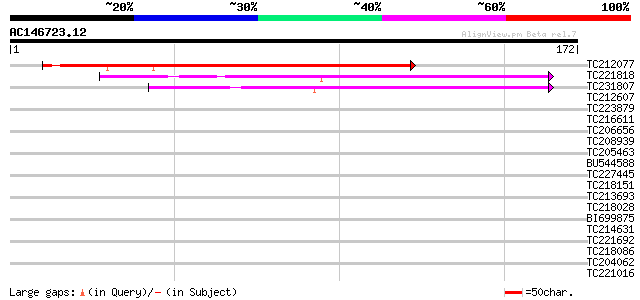
Sequences producing significant alignments: (bits) Value
TC212077 weakly similar to UP|Q9Z2Z7 (Q9Z2Z7) Phosphatidylglycer... 167 2e-42
TC221818 122 6e-29
TC231807 similar to UP|Q7XJ59 (Q7XJ59) At1g10530, partial (9%) 95 1e-20
TC212607 similar to UP|Q9ASS9 (Q9ASS9) AT4g24840/F6I7_50, partia... 36 0.008
TC223879 30 0.56
TC216611 30 0.73
TC206656 30 0.73
TC208939 UP|Q39830 (Q39830) BiP isoform A, complete 29 0.96
TC205463 UP|Q39804 (Q39804) BiP isoform B, complete 29 0.96
BU544588 29 0.96
TC227445 homologue to UP|O22639 (O22639) Endoplasmic reticulum H... 29 1.3
TC218151 homologue to UP|O22639 (O22639) Endoplasmic reticulum H... 29 1.3
TC213693 similar to UP|Q9NJY8 (Q9NJY8) Fruitless protein type B ... 28 2.8
TC218028 28 2.8
BI699875 similar to PIR|E96668|E96 protein F1N19.3 [imported] - ... 28 2.8
TC214631 weakly similar to GB|AAL31248.1|16974489|AY061921 At2g2... 27 3.6
TC221692 UP|Q9FEC5 (Q9FEC5) Glycinin subunit G7, complete 27 3.6
TC218086 27 3.6
TC204062 homologue to UP|Q09083 (Q09083) Hydroxyproline-rich gly... 27 3.6
TC221016 similar to GB|AAB72116.2|13699777|ATU69533 AtKAP alpha ... 27 6.2
>TC212077 weakly similar to UP|Q9Z2Z7 (Q9Z2Z7) Phosphatidylglycerophosphate
synthase, partial (3%)
Length = 414
Score = 167 bits (423), Expect = 2e-42
Identities = 86/116 (74%), Positives = 98/116 (84%), Gaps = 3/116 (2%)
Frame = +2
Query: 11 FLIPSSSSSIPSLCFSQT-TTKCIQRFSFPSIN--RKVSTLPFSSIKRSITRAAEYKFPD 67
FLI SSSS PSLCF+QT + C QRF FPS+ RK STLPFSS KR +TRAAEYK PD
Sbjct: 71 FLI--SSSSFPSLCFAQTHSITCKQRFPFPSVKGRRKFSTLPFSSTKRLVTRAAEYKCPD 244
Query: 68 PIPEFADSETEKFQNHLLNKLSKKDVFEESVEEVVGVCTEIFSTFLHSEYGGPGTL 123
PIPEFAD+ET+KF++HLL KLSKKD++ ES+EEVVG+CTEI STFLHSEYGGPGTL
Sbjct: 245 PIPEFADAETQKFKSHLLKKLSKKDMYGESIEEVVGICTEIISTFLHSEYGGPGTL 412
>TC221818
Length = 680
Score = 122 bits (307), Expect = 6e-29
Identities = 67/140 (47%), Positives = 84/140 (59%), Gaps = 2/140 (1%)
Frame = +2
Query: 28 TTTKCIQRFSFPSINRKVSTLPFSSIKRSITRAAEYKFPDPIPEFADSETEKFQNHLLNK 87
T C FS I+R T ++ S T Y PDP+PEFA+ ET KF+ L K
Sbjct: 182 TLHHCFPPFSAKPISRNSFTC---ALTHSPTHTHVY--PDPVPEFAEHETHKFKVELFRK 346
Query: 88 LSKKDV--FEESVEEVVGVCTEIFSTFLHSEYGGPGTLLVDPFIDMADIVNERGLPGGPQ 145
LS+ DV F + ++ VV VC +IFS FLH EYGGPGTLLV+PF DM + ++ LPG
Sbjct: 347 LSEDDVDEFGDDLDAVVAVCAQIFSEFLHKEYGGPGTLLVEPFTDMMVALKKKKLPGAAL 526
Query: 146 AARAAINWAQAHVDKDWNEW 165
AARA++ WA VDKDW W
Sbjct: 527 AARASLLWAPNFVDKDWEVW 586
>TC231807 similar to UP|Q7XJ59 (Q7XJ59) At1g10530, partial (9%)
Length = 971
Score = 95.1 bits (235), Expect = 1e-20
Identities = 53/125 (42%), Positives = 72/125 (57%), Gaps = 2/125 (1%)
Frame = +1
Query: 43 RKVSTLPFSSIKRSITRAAEYKFPDPIPEFADSETEKFQNHLLNKLSKK--DVFEESVEE 100
R+ +L S I + +FP+ + ET+KF+ L KLS++ D F + +
Sbjct: 13 RETPSLALSHIHPLTSTCTPTQFPN---SQST*ETQKFKFELFWKLSEEGVDEFGDDLHS 183
Query: 101 VVGVCTEIFSTFLHSEYGGPGTLLVDPFIDMADIVNERGLPGGPQAARAAINWAQAHVDK 160
VV VC +IFS FLH EY GPGTLLV+PF DM + ++ LPG AAR ++ W Q VDK
Sbjct: 184 VVDVCAQIFSEFLHKEYRGPGTLLVEPFTDMMVALKKKKLPGAALAARESLLWLQNFVDK 363
Query: 161 DWNEW 165
DW W
Sbjct: 364 DWEVW 378
>TC212607 similar to UP|Q9ASS9 (Q9ASS9) AT4g24840/F6I7_50, partial (19%)
Length = 653
Score = 36.2 bits (82), Expect = 0.008
Identities = 22/59 (37%), Positives = 26/59 (43%)
Frame = +2
Query: 26 SQTTTKCIQRFSFPSINRKVSTLPFSSIKRSITRAAEYKFPDPIPEFADSETEKFQNHL 84
S+ T C F FPS N K + SS K+ A K DPIP S T F + L
Sbjct: 47 SERTRSCSY*FQFPSPNHKQTDKSISSTKKHTPNRANQKMADPIPAPPRSATFLFSDPL 223
>TC223879
Length = 424
Score = 30.0 bits (66), Expect = 0.56
Identities = 18/48 (37%), Positives = 25/48 (51%), Gaps = 2/48 (4%)
Frame = +3
Query: 46 STLPFSSIKRSITRAAEYKFPDPIPEFADSET--EKFQNHLLNKLSKK 91
ST+P S I+ S RA+ + P PIP F D +K + NK +K
Sbjct: 189 STIPMSPIRYSPIRASGTEIPIPIPIFPDQSLWWKKLKKRKYNKWLRK 332
>TC216611
Length = 634
Score = 29.6 bits (65), Expect = 0.73
Identities = 20/48 (41%), Positives = 26/48 (53%), Gaps = 6/48 (12%)
Frame = -2
Query: 6 WC--LVPFLIPSSSSSIPSL----CFSQTTTKCIQRFSFPSINRKVST 47
WC PF +PS+S+ +P L CFS T C Q F SI KV++
Sbjct: 363 WC*PWTPFSLPSTSTQLPPLTPACCFSFT---C*QMFPMGSIFLKVAS 229
>TC206656
Length = 486
Score = 29.6 bits (65), Expect = 0.73
Identities = 46/169 (27%), Positives = 63/169 (37%), Gaps = 17/169 (10%)
Frame = +3
Query: 16 SSSSIPSLCFSQTTTKCIQRFSFPSINRKVSTL--------------PFSSIKRSITRAA 61
SSSS PS F RF+F R S L PFSS S A
Sbjct: 12 SSSSPPSFTFLTRFESATARFNFSPSTRSHSHLKRLSSICCQHANPSPFSS---STPFLA 182
Query: 62 EYKFPDPIPEF-ADSETEKFQNHLLNKLSKKDVFEESVEEVVGVCTEIFSTFLHSEYGGP 120
P P F + S+ ++H LN++ K F + +V+ T + + L
Sbjct: 183 SSNHPINFPSFSSSSDPSTLESHSLNQI-KTGAFPK--PKVITARTLVILSAL------- 332
Query: 121 GTLLVDPFIDMADIV--NERGLPGGPQAARAAINWAQAHVDKDWNEWTG 167
+L+ P I A GGP AA AA+ H + + WTG
Sbjct: 333 AMILIQPTIAPAAFATFQTAAKTGGP-AAAAAVGRKLIHTELLSSAWTG 476
>TC208939 UP|Q39830 (Q39830) BiP isoform A, complete
Length = 2452
Score = 29.3 bits (64), Expect = 0.96
Identities = 13/36 (36%), Positives = 19/36 (52%)
Frame = +3
Query: 86 NKLSKKDVFEESVEEVVGVCTEIFSTFLHSEYGGPG 121
N+ +K+ +EE ++EV VC I S G PG
Sbjct: 1899 NQSVEKEEYEEKLKEVEAVCNPIISAVYQRSGGAPG 2006
>TC205463 UP|Q39804 (Q39804) BiP isoform B, complete
Length = 2383
Score = 29.3 bits (64), Expect = 0.96
Identities = 13/36 (36%), Positives = 19/36 (52%)
Frame = +1
Query: 86 NKLSKKDVFEESVEEVVGVCTEIFSTFLHSEYGGPG 121
N+ +K+ +EE ++EV VC I S G PG
Sbjct: 1933 NQSMEKEDYEEKLKEVEAVCNPIISAVYQRSGGAPG 2040
>BU544588
Length = 414
Score = 29.3 bits (64), Expect = 0.96
Identities = 13/36 (36%), Positives = 19/36 (52%)
Frame = -2
Query: 86 NKLSKKDVFEESVEEVVGVCTEIFSTFLHSEYGGPG 121
N+ +K+ +EE ++EV VC I S G PG
Sbjct: 191 NQSVEKEEYEEKLKEVEAVCNPIISAVYQRSGGAPG 84
>TC227445 homologue to UP|O22639 (O22639) Endoplasmic reticulum HSC70-cognate
binding protein precursor, complete
Length = 2254
Score = 28.9 bits (63), Expect = 1.3
Identities = 13/36 (36%), Positives = 19/36 (52%)
Frame = +2
Query: 86 NKLSKKDVFEESVEEVVGVCTEIFSTFLHSEYGGPG 121
N+ +K+ +EE ++EV VC I S G PG
Sbjct: 1952 NQSVEKEDYEEKLKEVEAVCNPIISAVYQRSGGAPG 2059
>TC218151 homologue to UP|O22639 (O22639) Endoplasmic reticulum HSC70-cognate
binding protein precursor, partial (35%)
Length = 922
Score = 28.9 bits (63), Expect = 1.3
Identities = 13/36 (36%), Positives = 19/36 (52%)
Frame = +1
Query: 86 NKLSKKDVFEESVEEVVGVCTEIFSTFLHSEYGGPG 121
N+ +K+ +EE ++EV VC I S G PG
Sbjct: 634 NQSVEKEDYEEKLKEVEAVCNPIISAVYQRSGGAPG 741
>TC213693 similar to UP|Q9NJY8 (Q9NJY8) Fruitless protein type B (Fragment),
partial (10%)
Length = 375
Score = 27.7 bits (60), Expect = 2.8
Identities = 14/34 (41%), Positives = 17/34 (49%)
Frame = +1
Query: 8 LVPFLIPSSSSSIPSLCFSQTTTKCIQRFSFPSI 41
L+P IP+ S P L F TT RFS P +
Sbjct: 175 LIPQQIPNPSKHFPFLSFRATTLSTSSRFSPPPL 276
>TC218028
Length = 1655
Score = 27.7 bits (60), Expect = 2.8
Identities = 17/56 (30%), Positives = 24/56 (42%)
Frame = -3
Query: 13 IPSSSSSIPSLCFSQTTTKCIQRFSFPSINRKVSTLPFSSIKRSITRAAEYKFPDP 68
+P S+IPSL K ++ S + LPFS++ IT A F P
Sbjct: 1386 LPVVHSAIPSLVIRGACKKLLRLSS*KYFLASIKILPFSNLASPITLACSMNFIPP 1219
>BI699875 similar to PIR|E96668|E96 protein F1N19.3 [imported] - Arabidopsis
thaliana, partial (37%)
Length = 421
Score = 27.7 bits (60), Expect = 2.8
Identities = 22/61 (36%), Positives = 28/61 (45%), Gaps = 11/61 (18%)
Frame = +2
Query: 16 SSSSIPSLCFSQTTTKCI---------QRFSFPSINRKVSTLPFSSI--KRSITRAAEYK 64
SSSS SLC S+ T K + R + PS +R ST P ++ RS T A
Sbjct: 50 SSSSFTSLCSSRRTPKSL*APSRSLPSPRLTAPSTSRSHSTPPLGTLTEARSRTTTARSS 229
Query: 65 F 65
F
Sbjct: 230 F 232
>TC214631 weakly similar to GB|AAL31248.1|16974489|AY061921
At2g27080/T20P8.13 {Arabidopsis thaliana;} , partial
(22%)
Length = 1068
Score = 27.3 bits (59), Expect = 3.6
Identities = 16/58 (27%), Positives = 28/58 (47%), Gaps = 3/58 (5%)
Frame = -1
Query: 2 ASSTWCLVPFLIPSSSSSIP---SLCFSQTTTKCIQRFSFPSINRKVSTLPFSSIKRS 56
A S W P + SS+ L SQT ++C+ P+ R ++ P+ +++RS
Sbjct: 495 APSVWSAPPATLFGPRSSVRFRHPLTPSQTPSRCLTAGKTPAERRYTASRPYKTLRRS 322
>TC221692 UP|Q9FEC5 (Q9FEC5) Glycinin subunit G7, complete
Length = 1674
Score = 27.3 bits (59), Expect = 3.6
Identities = 17/49 (34%), Positives = 26/49 (52%), Gaps = 4/49 (8%)
Frame = -1
Query: 3 SSTWCL---VPFLIPSSSSSIPSLCF-SQTTTKCIQRFSFPSINRKVST 47
S WCL + F +SSS PSLCF S ++C + F S + +++
Sbjct: 912 SPMWCLRFLISFSCTTSSSPSPSLCFLSFWHSQCSRVLGFSSTSASLTS 766
>TC218086
Length = 1385
Score = 27.3 bits (59), Expect = 3.6
Identities = 18/50 (36%), Positives = 27/50 (54%), Gaps = 1/50 (2%)
Frame = +3
Query: 3 SSTWCLVPFLIPSSSSSIPSLCFSQT-TTKCIQRFSFPSINRKVSTLPFS 51
SS+ C+V FL +P CF +KCI FSF S R+ +++ F+
Sbjct: 573 SSSSCMVLFL------PLPIQCFILVLASKCIISFSFSSAFRECTSITFT 704
>TC204062 homologue to UP|Q09083 (Q09083) Hydroxyproline-rich glycoprotein
precursor, partial (34%)
Length = 722
Score = 27.3 bits (59), Expect = 3.6
Identities = 21/54 (38%), Positives = 30/54 (54%)
Frame = +3
Query: 13 IPSSSSSIPSLCFSQTTTKCIQRFSFPSINRKVSTLPFSSIKRSITRAAEYKFP 66
+P SSSS + FS TT+ IQ S S+ +VST SSI+ I+ + + P
Sbjct: 360 VPISSSSSIQVPFSSTTSL*IQISSSTSLQIQVST---SSIQVPISTTSTLQVP 512
>TC221016 similar to GB|AAB72116.2|13699777|ATU69533 AtKAP alpha {Arabidopsis
thaliana;} , partial (27%)
Length = 851
Score = 26.6 bits (57), Expect = 6.2
Identities = 13/32 (40%), Positives = 20/32 (61%), Gaps = 1/32 (3%)
Frame = +2
Query: 75 SETEKFQNHLLNKLSKKDVF-EESVEEVVGVC 105
SET F +HL++ S +D+F + + VGVC
Sbjct: 422 SETMSFPSHLVDSTSAEDLFLRRDLLDEVGVC 517
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.316 0.132 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,035,801
Number of Sequences: 63676
Number of extensions: 133930
Number of successful extensions: 942
Number of sequences better than 10.0: 55
Number of HSP's better than 10.0 without gapping: 931
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 938
length of query: 172
length of database: 12,639,632
effective HSP length: 91
effective length of query: 81
effective length of database: 6,845,116
effective search space: 554454396
effective search space used: 554454396
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 55 (25.8 bits)
Medicago: description of AC146723.12


