

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146722.4 - phase: 1
(88 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
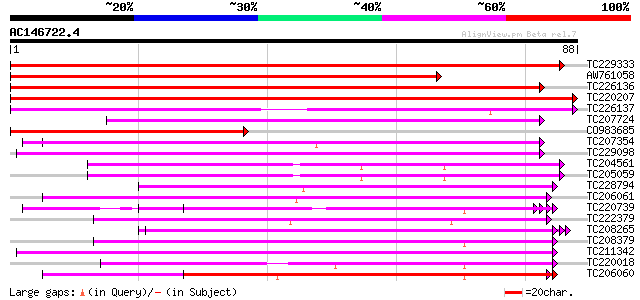
Score E
Sequences producing significant alignments: (bits) Value
TC229333 similar to UP|Q6H976 (Q6H976) STYLOSA protein, partial ... 160 1e-40
AW761058 124 1e-29
TC226136 similar to UP|O48847 (O48847) Expressed protein (At2g32... 120 1e-28
TC220207 similar to UP|Q6H975 (Q6H975) STY-L protein, partial (43%) 97 1e-21
TC226137 similar to UP|O48847 (O48847) Expressed protein (At2g32... 61 1e-10
TC207724 similar to PIR|A96839|A96839 F23A5.2(form2) [imported] ... 58 9e-10
CO983685 55 5e-09
TC207354 weakly similar to UP|WDR5_HUMAN (P61964) WD-repeat prot... 55 8e-09
TC229098 similar to PIR|T46032|T46032 WD-40 repeat regulatory pr... 50 2e-07
TC204561 homologue to UP|GBLP_SOYBN (Q39836) Guanine nucleotide-... 48 7e-07
TC205059 homologue to UP|GBLP_SOYBN (Q39836) Guanine nucleotide-... 48 7e-07
TC228794 similar to UP|Q6V8V0 (Q6V8V0) Transducin-like (Fragment... 48 1e-06
TC206061 weakly similar to UP|Q9FNN2 (Q9FNN2) WD-repeat protein-... 48 1e-06
TC220739 weakly similar to UP|PRP4_HUMAN (O43172) U4/U6 small nu... 47 2e-06
TC222379 homologue to UP|Q9HGV7 (Q9HGV7) Gbeta like protein, par... 47 2e-06
TC208265 similar to UP|Q6S7B0 (Q6S7B0) TAF5, partial (36%) 46 4e-06
TC208379 similar to UP|Q9FN19 (Q9FN19) Arabidopsis thaliana geno... 45 5e-06
TC211342 similar to UP|Q9LVF2 (Q9LVF2) Arabidopsis thaliana geno... 44 1e-05
TC220018 similar to UP|Q9FN19 (Q9FN19) Arabidopsis thaliana geno... 44 1e-05
TC206060 weakly similar to UP|Q9FNN2 (Q9FNN2) WD-repeat protein-... 43 2e-05
>TC229333 similar to UP|Q6H976 (Q6H976) STYLOSA protein, partial (32%)
Length = 982
Score = 160 bits (405), Expect = 1e-40
Identities = 77/86 (89%), Positives = 82/86 (94%)
Frame = +3
Query: 1 FTFSELNSVRASTNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITD 60
FTFS++NSVRAST+KV C HFSSDGKLLASGGHDK+VVLWYTDSLKQKATLEEHS LITD
Sbjct: 624 FTFSDVNSVRASTSKVACCHFSSDGKLLASGGHDKRVVLWYTDSLKQKATLEEHSSLITD 803
Query: 61 VRFSPSMPRLATSSYDKTVRVWDVDN 86
VRFSPSMPRLATSS+DKTV VWDVDN
Sbjct: 804 VRFSPSMPRLATSSFDKTVXVWDVDN 881
>AW761058
Length = 355
Score = 124 bits (310), Expect = 1e-29
Identities = 59/67 (88%), Positives = 64/67 (95%)
Frame = +3
Query: 1 FTFSELNSVRASTNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITD 60
FT+S++NSVRAST+KV C HFSSDGKLLASGGHDK+VVLWYTDSLKQKATLEEHS LITD
Sbjct: 153 FTYSDVNSVRASTSKVACCHFSSDGKLLASGGHDKRVVLWYTDSLKQKATLEEHSSLITD 332
Query: 61 VRFSPSM 67
VRFSPSM
Sbjct: 333 VRFSPSM 353
>TC226136 similar to UP|O48847 (O48847) Expressed protein
(At2g32700/F24L7.16), partial (42%)
Length = 1463
Score = 120 bits (301), Expect = 1e-28
Identities = 56/83 (67%), Positives = 70/83 (83%)
Frame = +3
Query: 1 FTFSELNSVRASTNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITD 60
F+FSE+ S+R S +KVVC HFSSDGKLLAS GHDKKVVLW ++L+ ++T EEHSL+ITD
Sbjct: 195 FSFSEVGSIRKSNSKVVCCHFSSDGKLLASAGHDKKVVLWNMETLQTESTPEEHSLIITD 374
Query: 61 VRFSPSMPRLATSSYDKTVRVWD 83
VRF P+ +LATSS+D TVR+WD
Sbjct: 375 VRFRPNSTQLATSSFDTTVRLWD 443
Score = 32.0 bits (71), Expect = 0.055
Identities = 20/65 (30%), Positives = 34/65 (51%)
Frame = +3
Query: 18 CSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSYDK 77
C S LL GG+ + + LW K T+ H +I+ + SP +A++S+DK
Sbjct: 873 CVFHPSYSTLLVIGGY-QSLELWNMAENKCM-TIPAHECVISALAQSPLTGMVASASHDK 1046
Query: 78 TVRVW 82
+V++W
Sbjct: 1047SVKIW 1061
>TC220207 similar to UP|Q6H975 (Q6H975) STY-L protein, partial (43%)
Length = 1244
Score = 97.4 bits (241), Expect = 1e-21
Identities = 47/88 (53%), Positives = 61/88 (68%)
Frame = +3
Query: 1 FTFSELNSVRASTNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITD 60
FTF+E+ R +KV C HFSSDGK LAS G D KV +W D+L+ ++T EH +ITD
Sbjct: 375 FTFAEVGCRRTRNSKVTCCHFSSDGKWLASAGDDMKVDIWNMDTLETESTPAEHKSVITD 554
Query: 61 VRFSPSMPRLATSSYDKTVRVWDVDNVS 88
VRF P+ +LAT+S DK+VR+WD N S
Sbjct: 555 VRFRPNSSQLATASTDKSVRLWDTTNPS 638
>TC226137 similar to UP|O48847 (O48847) Expressed protein
(At2g32700/F24L7.16), partial (15%)
Length = 417
Score = 60.8 bits (146), Expect = 1e-10
Identities = 34/89 (38%), Positives = 45/89 (50%), Gaps = 1/89 (1%)
Frame = +3
Query: 1 FTFSELNSVRASTNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITD 60
F+F E+ S+R S +KVVC HFSSDGKLL+S GHDKK T H+ +
Sbjct: 114 FSFDEVGSIRKSNSKVVCCHFSSDGKLLSSAGHDKKPTF-------PLHTYSGHTSHVVS 272
Query: 61 VRFSPSMPRLATS-SYDKTVRVWDVDNVS 88
+ F P L S + +R W + S
Sbjct: 273 LDFHPKKTELFCSCDNNNEIRFWSISQYS 359
>TC207724 similar to PIR|A96839|A96839 F23A5.2(form2) [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(98%)
Length = 1343
Score = 57.8 bits (138), Expect = 9e-10
Identities = 28/68 (41%), Positives = 40/68 (58%)
Frame = +2
Query: 16 VVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSY 75
V+CS + DG + SGG DK+V +W S Q T+ H + D+ + P M LAT S+
Sbjct: 287 VLCSAWKDDGTTVFSGGCDKQVKMWPLMSGGQPMTVAMHDAPVKDIAWIPEMNLLATGSW 466
Query: 76 DKTVRVWD 83
DKT++ WD
Sbjct: 467 DKTLKYWD 490
Score = 26.2 bits (56), Expect = 3.0
Identities = 10/27 (37%), Positives = 16/27 (59%)
Frame = +2
Query: 58 ITDVRFSPSMPRLATSSYDKTVRVWDV 84
I+ + FSP L +S+D VR W++
Sbjct: 146 ISSICFSPKANFLVATSWDNQVRCWEI 226
>CO983685
Length = 720
Score = 55.5 bits (132), Expect = 5e-09
Identities = 27/37 (72%), Positives = 30/37 (80%)
Frame = -3
Query: 1 FTFSELNSVRASTNKVVCSHFSSDGKLLASGGHDKKV 37
F+FSE+ S+ S NKVVC HFSSDGKLLAS G DKKV
Sbjct: 205 FSFSEVGSICKSNNKVVCCHFSSDGKLLASVGLDKKV 95
>TC207354 weakly similar to UP|WDR5_HUMAN (P61964) WD-repeat protein 5,
partial (57%)
Length = 1124
Score = 54.7 bits (130), Expect = 8e-09
Identities = 28/81 (34%), Positives = 46/81 (56%)
Frame = +1
Query: 3 FSELNSVRASTNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVR 62
+ L ++ N V C FS+DG LLAS DK +++W + +L L HS I+D+
Sbjct: 106 YRHLKTLTDHENAVSCVKFSNDGTLLASASLDKTLIIWSSATLTLCHRLVGHSEGISDLA 285
Query: 63 FSPSMPRLATSSYDKTVRVWD 83
+S + ++S D+T+R+WD
Sbjct: 286 WSSDSHYICSASDDRTLRIWD 348
Score = 42.4 bits (98), Expect = 4e-05
Identities = 18/79 (22%), Positives = 44/79 (54%), Gaps = 1/79 (1%)
Frame = +1
Query: 6 LNSVRASTNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLK-QKATLEEHSLLITDVRFS 64
+++++ T V H++ DG L+ S HD +W T++ K +E+ + ++ +FS
Sbjct: 496 VHTIKGHTMPVTSVHYNRDGNLIISASHDGSCKIWDTETGNLLKTLIEDKAPAVSFAKFS 675
Query: 65 PSMPRLATSSYDKTVRVWD 83
P+ + ++ + T+++W+
Sbjct: 676 PNGKLILAATLNDTLKLWN 732
>TC229098 similar to PIR|T46032|T46032 WD-40 repeat regulatory protein tup1
homolog - Arabidopsis thaliana {Arabidopsis thaliana;} ,
partial (59%)
Length = 580
Score = 50.1 bits (118), Expect = 2e-07
Identities = 27/82 (32%), Positives = 41/82 (49%)
Frame = +3
Query: 2 TFSELNSVRASTNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDV 61
T S + ++ TN V C +F+ ++ SG D+ V +W S K L HS +T V
Sbjct: 21 TGSLIKTLHGHTNYVFCVNFNPQSNIIVSGSFDETVRVWDVKSGKCLKVLPAHSDPVTAV 200
Query: 62 RFSPSMPRLATSSYDKTVRVWD 83
F+ + +SSYD R+WD
Sbjct: 201 DFNRDGSLIVSSSYDGLCRIWD 266
Score = 36.6 bits (83), Expect = 0.002
Identities = 19/79 (24%), Positives = 40/79 (50%), Gaps = 1/79 (1%)
Frame = +3
Query: 6 LNSVRASTNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLK-QKATLEEHSLLITDVRFS 64
L + A ++ V F+ DG L+ S +D +W + K +++ + ++ V+FS
Sbjct: 159 LKVLPAHSDPVTAVDFNRDGSLIVSSSYDGLCRIWDASTGHCMKTLIDDDNPPVSFVKFS 338
Query: 65 PSMPRLATSSYDKTVRVWD 83
P+ + + D T+R+W+
Sbjct: 339 PNAKFILVGTLDNTLRLWN 395
Score = 35.4 bits (80), Expect = 0.005
Identities = 17/46 (36%), Positives = 25/46 (53%)
Frame = +3
Query: 39 LWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSYDKTVRVWDV 84
LW + TL H+ + V F+P + + S+D+TVRVWDV
Sbjct: 6 LWDVPTGSLIKTLHGHTNYVFCVNFNPQSNIIVSGSFDETVRVWDV 143
>TC204561 homologue to UP|GBLP_SOYBN (Q39836) Guanine nucleotide-binding
protein beta subunit-like protein, complete
Length = 1419
Score = 48.1 bits (113), Expect = 7e-07
Identities = 28/79 (35%), Positives = 47/79 (59%), Gaps = 5/79 (6%)
Frame = +1
Query: 13 TNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEE---HSLLITDVRFSPS--M 67
T V+ FS D + + S D+ + LW T + K T+++ HS ++ VRFSPS
Sbjct: 457 TKDVLSVAFSIDNRQIVSASRDRTIKLWNTLG-ECKYTIQDGDAHSDWVSCVRFSPSTLQ 633
Query: 68 PRLATSSYDKTVRVWDVDN 86
P + ++S+D+TV+VW++ N
Sbjct: 634 PTIVSASWDRTVKVWNLTN 690
Score = 37.7 bits (86), Expect = 0.001
Identities = 22/67 (32%), Positives = 41/67 (60%)
Frame = +1
Query: 22 SSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSYDKTVRV 81
S DG L ASGG D ++LW K+ +L+ S +I + FSP+ L ++ ++++++
Sbjct: 748 SPDGSLCASGGKDGVILLWDLAEGKRLYSLDAGS-IIHALCFSPNRYWLCAAT-EQSIKI 921
Query: 82 WDVDNVS 88
WD+++ S
Sbjct: 922 WDLESKS 942
Score = 34.7 bits (78), Expect = 0.008
Identities = 20/71 (28%), Positives = 35/71 (49%), Gaps = 6/71 (8%)
Frame = +1
Query: 19 SHF------SSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLAT 72
SHF SSDG+ SG D ++ LW + H+ + V FS ++ +
Sbjct: 331 SHFVQDVVLSSDGQFALSGSWDGELRLWDLAAGTSARRFVGHTKDVLSVAFSIDNRQIVS 510
Query: 73 SSYDKTVRVWD 83
+S D+T+++W+
Sbjct: 511 ASRDRTIKLWN 543
>TC205059 homologue to UP|GBLP_SOYBN (Q39836) Guanine nucleotide-binding
protein beta subunit-like protein, complete
Length = 1492
Score = 48.1 bits (113), Expect = 7e-07
Identities = 28/79 (35%), Positives = 47/79 (59%), Gaps = 5/79 (6%)
Frame = +2
Query: 13 TNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEE---HSLLITDVRFSPS--M 67
T V+ FS D + + S D+ + LW T + K T+++ HS ++ VRFSPS
Sbjct: 533 TKDVLSVAFSIDNRQIVSASRDRTIKLWNTLG-ECKYTIQDGDAHSDWVSCVRFSPSTLQ 709
Query: 68 PRLATSSYDKTVRVWDVDN 86
P + ++S+D+TV+VW++ N
Sbjct: 710 PTIVSASWDRTVKVWNLTN 766
Score = 37.7 bits (86), Expect = 0.001
Identities = 22/67 (32%), Positives = 41/67 (60%)
Frame = +2
Query: 22 SSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSYDKTVRV 81
S DG L ASGG D ++LW K+ +L+ S +I + FSP+ L ++ ++++++
Sbjct: 824 SPDGSLCASGGKDGVILLWDLAEGKRLYSLDAGS-IIHALCFSPNRYWLCAAT-EQSIKI 997
Query: 82 WDVDNVS 88
WD+++ S
Sbjct: 998 WDLESKS 1018
Score = 34.7 bits (78), Expect = 0.008
Identities = 20/71 (28%), Positives = 35/71 (49%), Gaps = 6/71 (8%)
Frame = +2
Query: 19 SHF------SSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLAT 72
SHF SSDG+ SG D ++ LW + H+ + V FS ++ +
Sbjct: 407 SHFVQDVVLSSDGQFALSGSWDGELRLWDLAAGTSARRFVGHTKDVLSVAFSIDNRQIVS 586
Query: 73 SSYDKTVRVWD 83
+S D+T+++W+
Sbjct: 587 ASRDRTIKLWN 619
>TC228794 similar to UP|Q6V8V0 (Q6V8V0) Transducin-like (Fragment), complete
Length = 870
Score = 47.8 bits (112), Expect = 1e-06
Identities = 22/66 (33%), Positives = 38/66 (57%), Gaps = 1/66 (1%)
Frame = +3
Query: 21 FSSDGKLLASGGHDKKVVLWYTDS-LKQKATLEEHSLLITDVRFSPSMPRLATSSYDKTV 79
F+ G ++ASG HD+++ LW K L+ H + D+ ++ ++ ++S DKTV
Sbjct: 303 FNPAGSVVASGSHDREIFLWNVHGDCKNFMVLKGHKNAVLDLHWTTDGTQIVSASPDKTV 482
Query: 80 RVWDVD 85
R WDV+
Sbjct: 483 RAWDVE 500
Score = 25.8 bits (55), Expect = 3.9
Identities = 10/34 (29%), Positives = 21/34 (61%)
Frame = +3
Query: 51 LEEHSLLITDVRFSPSMPRLATSSYDKTVRVWDV 84
L H I ++F+P+ +A+ S+D+ + +W+V
Sbjct: 267 LSGHQSAIYTMKFNPAGSVVASGSHDREIFLWNV 368
>TC206061 weakly similar to UP|Q9FNN2 (Q9FNN2) WD-repeat protein-like,
partial (36%)
Length = 657
Score = 47.8 bits (112), Expect = 1e-06
Identities = 27/82 (32%), Positives = 41/82 (49%), Gaps = 3/82 (3%)
Frame = +3
Query: 6 LNSVRASTNKVVCSHFSSDGKLLASGGHDKKVVLWYTD---SLKQKATLEEHSLLITDVR 62
L + A ++V FS +GK LAS +D+ ++W D L K L H ++ V
Sbjct: 189 LQILEAHDDEVWYVQFSHNGKYLASASNDRSAIIWEVDMNGELSVKHKLSGHQKPVSSVS 368
Query: 63 FSPSMPRLATSSYDKTVRVWDV 84
+SP+ L T ++ VR WDV
Sbjct: 369 WSPNDQELLTCGVEEAVRRWDV 434
>TC220739 weakly similar to UP|PRP4_HUMAN (O43172) U4/U6 small nuclear
ribonucleoprotein Prp4 (U4/U6 snRNP 60 kDa protein) (WD
splicing factor Prp4) (hPrp4), partial (34%)
Length = 1206
Score = 46.6 bits (109), Expect = 2e-06
Identities = 26/63 (41%), Positives = 34/63 (53%), Gaps = 1/63 (1%)
Frame = +1
Query: 21 FSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPR-LATSSYDKTV 79
FS +G LA+GG D +W K T+ HS LI+ V+F P L T+SYD T
Sbjct: 595 FSPNGYHLATGGEDNTCRIWDLRKKKSFYTIPAHSNLISQVKFEPHEGYFLVTASYDMTA 774
Query: 80 RVW 82
+VW
Sbjct: 775 KVW 783
Score = 46.2 bits (108), Expect = 3e-06
Identities = 29/81 (35%), Positives = 42/81 (51%)
Frame = +1
Query: 3 FSELNSVRASTNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVR 62
FSE+ R + CS FS DGK LA+ LW +K+ + + H+ TDV
Sbjct: 55 FSEIGDDRPLSG---CS-FSRDGKWLATCSLTGASKLWSMPKIKKHSIFKGHTERATDVA 222
Query: 63 FSPSMPRLATSSYDKTVRVWD 83
+SP LAT+S D+T + W+
Sbjct: 223 YSPVHDHLATASADRTAKYWN 285
Score = 43.5 bits (101), Expect = 2e-05
Identities = 23/64 (35%), Positives = 32/64 (49%)
Frame = +1
Query: 21 FSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSYDKTVR 80
F +DG L AS G D +W + + LE H + + FSP+ LAT D T R
Sbjct: 469 FHNDGSLAASCGLDSLARVWDLRTGRSILALEGHVKPVLGISFSPNGYHLATGGEDNTCR 648
Query: 81 VWDV 84
+WD+
Sbjct: 649 IWDL 660
Score = 39.3 bits (90), Expect = 3e-04
Identities = 21/58 (36%), Positives = 31/58 (53%)
Frame = +1
Query: 28 LASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSYDKTVRVWDVD 85
LA+ D+ W SL + T E H + + F PS L T+S+DKT R+WD++
Sbjct: 244 LATASADRTAKYWNQGSLLK--TFEGHLDRLARIAFHPSGKYLGTASFDKTWRLWDIE 411
Score = 33.1 bits (74), Expect = 0.025
Identities = 15/59 (25%), Positives = 26/59 (43%)
Frame = +1
Query: 24 DGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSYDKTVRVW 82
+G L + +D +W K TL H +T V + T S+D+T+++W
Sbjct: 733 EGYFLVTASYDMTAKVWSGRDFKPVKTLSGHEAKVTSVDVLGDGGSIVTVSHDRTIKLW 909
>TC222379 homologue to UP|Q9HGV7 (Q9HGV7) Gbeta like protein, partial (84%)
Length = 950
Score = 46.6 bits (109), Expect = 2e-06
Identities = 25/74 (33%), Positives = 43/74 (57%), Gaps = 3/74 (4%)
Frame = +1
Query: 14 NKVVCSHFSSDGKLLASGGHDKKVVLWYT-DSLKQKATLEEHSLLITDVRFSPSM--PRL 70
N V+ FS+D + + SG D+ + LW T K T + H+ ++ VRFSP+ P +
Sbjct: 163 NDVLSVSFSADNRQIVSGSRDRTIKLWNTLGDCKYTITDKGHTEWVSCVRFSPNPQNPVI 342
Query: 71 ATSSYDKTVRVWDV 84
++ +DK V+VW++
Sbjct: 343 VSAGWDKFVKVWEL 384
Score = 34.3 bits (77), Expect = 0.011
Identities = 17/62 (27%), Positives = 30/62 (47%)
Frame = +1
Query: 22 SSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSYDKTVRV 81
SSDG S DK + LW + + H+ + V FS ++ + S D+T+++
Sbjct: 61 SSDGAYALSSSWDKTLRLWELSTGQTTRRFVGHNNDVLSVSFSADNRQIVSGSRDRTIKL 240
Query: 82 WD 83
W+
Sbjct: 241 WN 246
Score = 30.4 bits (67), Expect = 0.16
Identities = 13/37 (35%), Positives = 23/37 (62%)
Frame = +1
Query: 48 KATLEEHSLLITDVRFSPSMPRLATSSYDKTVRVWDV 84
K +L HS +++D S +SS+DKT+R+W++
Sbjct: 13 KRSLHGHSHIVSDCVISSDGAYALSSSWDKTLRLWEL 123
Score = 28.9 bits (63), Expect = 0.46
Identities = 17/46 (36%), Positives = 24/46 (51%), Gaps = 4/46 (8%)
Frame = +1
Query: 22 SSDGKLLASGGHDKKVVLWYTDSLKQKATL----EEHSLLITDVRF 63
S DG L ASGG D +LW + K +L E H+L+ + R+
Sbjct: 448 SPDGSLCASGGKDGTTMLWDLNESKHLYSLTAGDEIHALVFSPNRY 585
>TC208265 similar to UP|Q6S7B0 (Q6S7B0) TAF5, partial (36%)
Length = 1048
Score = 45.8 bits (107), Expect = 4e-06
Identities = 22/67 (32%), Positives = 34/67 (49%)
Frame = +2
Query: 21 FSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSYDKTVR 80
FS G + S D + LW T + H+ + DV+FSP A+SS+D+T R
Sbjct: 11 FSPVGDFILSSSADSTIRLWSTKLNANLVCYKGHNYPVWDVQFSPVGHYFASSSHDRTAR 190
Query: 81 VWDVDNV 87
+W +D +
Sbjct: 191IWSMDRI 211
Score = 43.9 bits (102), Expect = 1e-05
Identities = 22/66 (33%), Positives = 34/66 (51%)
Frame = +2
Query: 21 FSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSYDKTVR 80
FS G AS HD+ +W D ++ + H + V++ + +AT S DKTVR
Sbjct: 137 FSPVGHYFASSSHDRTARIWSMDRIQPLRIMAGHLSDVDCVQWHANCNYIATGSSDKTVR 316
Query: 81 VWDVDN 86
+WDV +
Sbjct: 317 LWDVQS 334
Score = 42.0 bits (97), Expect = 5e-05
Identities = 21/64 (32%), Positives = 35/64 (53%)
Frame = +2
Query: 22 SSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSYDKTVRV 81
S DG+ +ASG D +++W S + L H+ + + FS +A+ S D TV++
Sbjct: 392 SPDGRYMASGDEDGTIMMWDLSSGRCLTPLIGHTSCVWSLAFSSEGSVIASGSADCTVKL 571
Query: 82 WDVD 85
WDV+
Sbjct: 572 WDVN 583
Score = 30.8 bits (68), Expect = 0.12
Identities = 17/55 (30%), Positives = 28/55 (50%)
Frame = +2
Query: 21 FSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSY 75
FSS+G ++ASG D V LW ++ + + EE +R ++P +T Y
Sbjct: 515 FSSEGSVIASGSADCTVKLWDVNTSTKVSRAEEKGGSANRLRSLKTLPTKSTPVY 679
>TC208379 similar to UP|Q9FN19 (Q9FN19) Arabidopsis thaliana genomic DNA,
chromosome 5, TAC clone:K8K14 (AT5g67320/K8K14_4),
partial (42%)
Length = 1062
Score = 45.4 bits (106), Expect = 5e-06
Identities = 27/81 (33%), Positives = 40/81 (49%), Gaps = 9/81 (11%)
Frame = +2
Query: 14 NKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPR---- 69
++V C + G LLAS D +W K EHS I +R+SP+ P
Sbjct: 287 SEVNCIKWDPTGSLLASCSDDMTAKIWSMKQDKYLHEFREHSKEIYTIRWSPTGPGTNNP 466
Query: 70 -----LATSSYDKTVRVWDVD 85
LA++S+D TV++WDV+
Sbjct: 467 NKNLVLASASFDSTVKLWDVE 529
Score = 34.3 bits (77), Expect = 0.011
Identities = 18/58 (31%), Positives = 30/58 (51%)
Frame = +2
Query: 27 LLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSYDKTVRVWDV 84
+LAS D V LW + K +L H + V FSP+ +A+ S D+++ +W +
Sbjct: 479 VLASASFDSTVKLWDVELGKLLYSLNGHRDRVYSVAFSPNGEYIASGSPDRSMLIWSL 652
Score = 27.7 bits (60), Expect = 1.0
Identities = 14/42 (33%), Positives = 25/42 (59%)
Frame = +2
Query: 6 LNSVRASTNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQ 47
L S+ ++V FS +G+ +ASG D+ +++W SLK+
Sbjct: 542 LYSLNGHRDRVYSVAFSPNGEYIASGSPDRSMLIW---SLKE 658
>TC211342 similar to UP|Q9LVF2 (Q9LVF2) Arabidopsis thaliana genomic DNA,
chromosome 3, P1 clone: MIL23, partial (29%)
Length = 862
Score = 44.3 bits (103), Expect = 1e-05
Identities = 24/84 (28%), Positives = 39/84 (45%)
Frame = +1
Query: 2 TFSELNSVRASTNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDV 61
TF S+ V+C SSDG L+ +G DK + +W D ++ H+ + V
Sbjct: 313 TFKFFLSLYGHKLPVLCMDISSDGDLIVTGSADKNIKIWGLDFGDCHKSIFAHADSVMAV 492
Query: 62 RFSPSMPRLATSSYDKTVRVWDVD 85
+F P + + D+ V+ WD D
Sbjct: 493 QFVPKTHYVFSVGKDRLVKYWDAD 564
Score = 35.8 bits (81), Expect = 0.004
Identities = 22/76 (28%), Positives = 34/76 (43%)
Frame = +1
Query: 8 SVRASTNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSM 67
S+ A + V+ F + S G D+ V W D + TLE H I + S
Sbjct: 457 SIFAHADSVMAVQFVPKTHYVFSVGKDRLVKYWDADKFELLLTLEGHHADIWCLAVSNRG 636
Query: 68 PRLATSSYDKTVRVWD 83
+ T S+D+++R WD
Sbjct: 637 DFIVTGSHDRSIRRWD 684
Score = 29.6 bits (65), Expect = 0.27
Identities = 23/88 (26%), Positives = 35/88 (39%), Gaps = 18/88 (20%)
Frame = +1
Query: 16 VVCSHFSSDGKLLASGGHDKKVVLW-------------YTDSLKQKATLEEHSLLITD-- 60
V ++ G LLASG D V+LW + D ++ T+ S + +
Sbjct: 49 VTTLRYNKAGSLLASGSRDNDVILWDVVGETGLFRLRGHRDQAAKQLTVSNVSTMKMNDD 228
Query: 61 ---VRFSPSMPRLATSSYDKTVRVWDVD 85
V SP +A + D TV+V D
Sbjct: 229 ALVVAISPDAKYIAVALLDSTVKVHFAD 312
>TC220018 similar to UP|Q9FN19 (Q9FN19) Arabidopsis thaliana genomic DNA,
chromosome 5, TAC clone:K8K14 (AT5g67320/K8K14_4),
partial (42%)
Length = 1050
Score = 43.9 bits (102), Expect = 1e-05
Identities = 30/83 (36%), Positives = 43/83 (51%), Gaps = 12/83 (14%)
Frame = +1
Query: 15 KVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKA---TLEEHSLLITDVRFSPSMPR-- 69
+V C + G LLAS D +W S+KQ L EHS I +R+SP+ P
Sbjct: 286 EVNCVKWDPTGSLLASCSDDITAKIW---SMKQDTYLHDLREHSKEIYTIRWSPTGPGTN 456
Query: 70 -------LATSSYDKTVRVWDVD 85
LA++S+D TV++WDV+
Sbjct: 457 NPNHKLVLASASFDSTVKLWDVE 525
Score = 35.4 bits (80), Expect = 0.005
Identities = 18/58 (31%), Positives = 30/58 (51%)
Frame = +1
Query: 27 LLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSYDKTVRVWDV 84
+LAS D V LW + K +L+ H + V FSP+ L + S D+++ +W +
Sbjct: 475 VLASASFDSTVKLWDVELGKLIYSLDGHRHPVYSVAFSPNGDYLVSGSLDRSMHIWSL 648
>TC206060 weakly similar to UP|Q9FNN2 (Q9FNN2) WD-repeat protein-like,
partial (62%)
Length = 1510
Score = 43.1 bits (100), Expect = 2e-05
Identities = 25/82 (30%), Positives = 39/82 (47%), Gaps = 3/82 (3%)
Frame = +3
Query: 6 LNSVRASTNKVVCSHFSSDGKLLASGGHDKKVVLW---YTDSLKQKATLEEHSLLITDVR 62
L + A ++V FS +GK LAS D+ ++W L K L H ++ V
Sbjct: 402 LQILEAHDDEVWYVQFSHNGKYLASASKDQTAIIWEVGINGRLSVKHRLSGHQKPVSSVS 581
Query: 63 FSPSMPRLATSSYDKTVRVWDV 84
+SP+ L T ++ +R WDV
Sbjct: 582 WSPNDQELLTCGVEEAIRRWDV 647
Score = 41.6 bits (96), Expect = 7e-05
Identities = 21/59 (35%), Positives = 36/59 (60%), Gaps = 1/59 (1%)
Frame = +3
Query: 28 LASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPR-LATSSYDKTVRVWDVD 85
+ASG D +V +W+ S + L HS + V ++P+ P LA++S D+T+RVW ++
Sbjct: 1113 IASGSEDSQVYIWHRSSGELIEALTGHSGSVNCVSWNPANPHMLASASDDRTIRVWGLN 1289
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.315 0.128 0.369
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,882,780
Number of Sequences: 63676
Number of extensions: 42418
Number of successful extensions: 455
Number of sequences better than 10.0: 153
Number of HSP's better than 10.0 without gapping: 351
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 436
length of query: 88
length of database: 12,639,632
effective HSP length: 64
effective length of query: 24
effective length of database: 8,564,368
effective search space: 205544832
effective search space used: 205544832
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 52 (24.6 bits)
Medicago: description of AC146722.4


